Research Briefing |
Featured
-
-
Article
| Open Access Tracking single-cell evolution using clock-like chromatin accessibility loci
Tracking single-cell evolution using clock-like chromatin accessibility loci
EpiTrace infers cell mitotic age and evolution from single-cell ATAC-seq data.
- Yu Xiao
- , Wan Jin
- & Yi Zhang
-
Article |
 Discovery of tumor-reactive T cell receptors by massively parallel library synthesis and screening
Discovery of tumor-reactive T cell receptors by massively parallel library synthesis and screening
Tumor-specific T cell receptors (TCRs) are identified by functional screening of synthetic TCR libraries.
- Ziva Moravec
- , Yue Zhao
- & Wouter Scheper
-
-
Article |
 Genome engineering with Cas9 and AAV repair templates generates frequent concatemeric insertions of viral vectors
Genome engineering with Cas9 and AAV repair templates generates frequent concatemeric insertions of viral vectors
AAV vectors form difficult-to-detect concatemers at Cas9 target sites.
- Fabian P. Suchy
- , Daiki Karigane
- & Hiromitsu Nakauchi
-
Research Briefing |
 Capturing and modeling cellular niches from dissociated single-cell and spatial data
Capturing and modeling cellular niches from dissociated single-cell and spatial data
Cells interact with their local environment to enact global tissue function. By harnessing gene–gene covariation in cellular neighborhoods from spatial transcriptomics data, the covariance environment (COVET) niche representation and the environmental variational inference (ENVI) data integration method model phenotype–microenvironment interplay and reconstruct the spatial context of dissociated single-cell RNA sequencing datasets.
-
Brief Communication
| Open Access De novo and somatic structural variant discovery with SVision-pro
De novo and somatic structural variant discovery with SVision-pro
SVision-pro enables pairwise genome comparison and improves de novo and somatic structural variant calling.
- Songbo Wang
- , Jiadong Lin
- & Kai Ye
-
Article |
 Near-cognate tRNAs increase the efficiency and precision of pseudouridine-mediated readthrough of premature termination codons
Near-cognate tRNAs increase the efficiency and precision of pseudouridine-mediated readthrough of premature termination codons
Disease-related premature stop codons are efficiently corrected by harnessing near-cognate tRNAs.
- Nan Luo
- , Qiang Huang
- & Chengqi Yi
-
Article |
 All-atom RNA structure determination from cryo-EM maps
All-atom RNA structure determination from cryo-EM maps
RNA structures are built from cryogenic electron microscopy maps using deep learning and backbone tracing.
- Tao Li
- , Jiahua He
- & Sheng-You Huang
-
Article |
 An oncolytic virus–T cell chimera for cancer immunotherapy
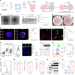
An oncolytic virus–T cell chimera for cancer immunotherapy
Oncolytic adenovirus is delivered on T cells and engineered to express a checkpoint inhibitor.
- Yuxuan Chen
- , Xiaohong Chen
- & Yuan Ping
-
Brief Communication
| Open Access Efficient prime editing in two-cell mouse embryos using PEmbryo
Efficient prime editing in two-cell mouse embryos using PEmbryo
Prime editing is used to efficiently edit early-stage mouse embryos with few on-target byproducts.
- Rebecca P. Kim-Yip
- , Ryan McNulty
- & Britt Adamson
-
News & Views |
 A new mass analyzer shakes up the proteomics field
A new mass analyzer shakes up the proteomics field
A new mass analyzer enables protein identification at high speed and depth.
- Bernhard Kuster
- , Johanna Tüshaus
- & Florian P. Bayer
-
Article
| Open Access Ultra-fast label-free quantification and comprehensive proteome coverage with narrow-window data-independent acquisition
Ultra-fast label-free quantification and comprehensive proteome coverage with narrow-window data-independent acquisition
A new mass spectrometry strategy combines the advantages of data-dependent and data-independent acquisition.
- Ulises H. Guzman
- , Ana Martinez-Val
- & Jesper V. Olsen
-
Article
| Open Access Activation of recombinases at specific DNA loci by zinc-finger domain insertions
Activation of recombinases at specific DNA loci by zinc-finger domain insertions
Recombinases fused with zinc-finger domains are active only when bound to their DNA targets.
- Liliya Mukhametzyanova
- , Lukas Theo Schmitt
- & Frank Buchholz
-
Research Briefing |
 Computer-based design of sensors that can monitor endogenous Ras activity
Computer-based design of sensors that can monitor endogenous Ras activity
Many questions on the activity of the Ras proto-oncogene are unanswered due to the lack of tools for detecting active Ras in living cells. Here, we used protein design and structure prediction algorithms to develop biosensors that detect the activity and environment of endogenous Ras.
-
Article
| Open Access Computationally designed sensors detect endogenous Ras activity and signaling effectors at subcellular resolution
Computationally designed sensors detect endogenous Ras activity and signaling effectors at subcellular resolution
Computationally designed sensors map the activity and signaling environment of Ras.
- Jason Z. Zhang
- , William H. Nguyen
- & David Baker
-
Article
| Open Access Mosaic integration and knowledge transfer of single-cell multimodal data with MIDAS
Mosaic integration and knowledge transfer of single-cell multimodal data with MIDAS
Single-cell, multiomic datasets are integrated using dimensionality reduction, imputation and batch correction.
- Zhen He
- , Shuofeng Hu
- & Xiaomin Ying
-
Article |
 Decoder-seq enhances mRNA capture efficiency in spatial RNA sequencing
Decoder-seq enhances mRNA capture efficiency in spatial RNA sequencing
A spatial transcriptomics method achieves high gene detection sensitivity and flexible resolution.
- Jiao Cao
- , Zhong Zheng
- & Chaoyong Yang
-
Article
| Open Access In vivo human T cell engineering with enveloped delivery vehicles
In vivo human T cell engineering with enveloped delivery vehicles
Cell-specific molecular delivery with enveloped delivery vehicles enables genome editing ex vivo and in vivo.
- Jennifer R. Hamilton
- , Evelyn Chen
- & Jennifer A. Doudna
-
Research Briefing |
 Generating expression profiles of single living cells from Raman microscopy
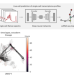
Generating expression profiles of single living cells from Raman microscopy
Expression profiles of single live cells are generated from Raman microscopy using deep learning, enabling us to track expression dynamics along cell reprogramming or differentiation.
-
Article |
 Engineered serum markers for non-invasive monitoring of gene expression in the brain
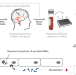
Engineered serum markers for non-invasive monitoring of gene expression in the brain
Gene expression in the mouse brain is measured with a simple blood draw.
- Sangsin Lee
- , Shirin Nouraein
- & Jerzy O. Szablowski
-
Article |
 Prime editing using CRISPR-Cas12a and circular RNAs in human cells
Prime editing using CRISPR-Cas12a and circular RNAs in human cells
Prime editing with CRISPR-Cas12a broadens targeting scope and enables multiplexing.
- Ronghong Liang
- , Zixin He
- & Caixia Gao
-
Brief Communication |
 Delivery of a sebum modulator by an engineered skin microbe in mice
Delivery of a sebum modulator by an engineered skin microbe in mice
An engineered skin microbe produces a therapeutic molecule that reduces sebum in mice.
- Nastassia Knödlseder
- , María-José Fábrega
- & Marc Güell
-
Article
| Open Access Engineered virus-like particles for transient delivery of prime editor ribonucleoprotein complexes in vivo
Engineered virus-like particles for transient delivery of prime editor ribonucleoprotein complexes in vivo
Delivery of prime editors in vivo is improved using virus-like particles.
- Meirui An
- , Aditya Raguram
- & David R. Liu
-
Research Briefing |
 Generation of programmable splicing factors using RNA-binding proteins that activate exon inclusion
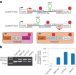
Generation of programmable splicing factors using RNA-binding proteins that activate exon inclusion
We developed luciferase-based splicing reporters to evaluate 718 RNA-binding proteins (RBPs) for the ability to activate exon inclusion. Our screen detected RBPs with no previously characterized roles in splicing and utilized RBP domains displaying potent exon activation to develop programmable splicing modulators with improved strength, generalizability and size over previous versions.
-
Article
| Open Access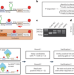 Large-scale evaluation of the ability of RNA-binding proteins to activate exon inclusion
Large-scale evaluation of the ability of RNA-binding proteins to activate exon inclusion
Alternative splicing is controlled with engineered fusion splicing factors.
- Jonathan C. Schmok
- , Manya Jain
- & Gene W. Yeo
-
Brief Communication |
 Inferring super-resolution tissue architecture by integrating spatial transcriptomics with histology
Inferring super-resolution tissue architecture by integrating spatial transcriptomics with histology
iStar predicts gene expression with near-single-cell resolution from histology images.
- Daiwei Zhang
- , Amelia Schroeder
- & Mingyao Li
-
Article
| Open Access Ultrafast bisulfite sequencing detection of 5-methylcytosine in DNA and RNA
Ultrafast bisulfite sequencing detection of 5-methylcytosine in DNA and RNA
Accelerated bisulfite sequencing detects 5-methylcytosine in low-input DNA and RNA samples
- Qing Dai
- , Chang Ye
- & Chuan He
-
Brief Communication
| Open Access A monomeric StayGold fluorescent protein
A monomeric StayGold fluorescent protein
The bright, photostable fluorescent protein StayGold is converted into a monomer.
- Esther Ivorra-Molla
- , Dipayan Akhuli
- & Mohan K. Balasubramanian
-
Article
| Open Access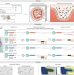 Spatial host–microbiome sequencing reveals niches in the mouse gut
Spatial host–microbiome sequencing reveals niches in the mouse gut
Spatial host–microbiome sequencing simultaneously profiles microbes and host transcriptomes from mouse colons.
- Britta Lötstedt
- , Martin Stražar
- & Sanja Vickovic
-
News & Views |
 Brain imaging turned inside out
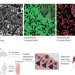
Brain imaging turned inside out
Neural connectivity in brain tissue is imaged by labeling the extracellular space.
- Shahrzad Askari
- & Thomas Misgeld
-
Research Briefing |
 Compressed Perturb-seq enables highly efficient genetic screens
Compressed Perturb-seq enables highly efficient genetic screens
Using an experimental and computational framework inspired by compressed sensing, we greatly reduced the number of measurements needed to run Perturb-seq. Our compressed Perturb-seq strategy relies on collecting measurements comprising random linear combinations of genetic perturbations, followed by deconvolving the perturbation effects on the transcriptome using sparsity-exploiting algorithms.
-
News & Views |
 Imaging glycosylated RNAs at the subcellular scale
Imaging glycosylated RNAs at the subcellular scale
A recently discovered RNA species on the cell surface is studied by proximity ligation.
- Petar Hristov
- & Ryan A. Flynn
-
Article
| Open Access Scalable genetic screening for regulatory circuits using compressed Perturb-seq
Scalable genetic screening for regulatory circuits using compressed Perturb-seq
Compressed Perturb-seq incorporates compressed sensing to genetic screening for scalable discovery of genetic interactions.
- Douglas Yao
- , Loic Binan
- & Brian Cleary
-
Article
| Open Access Systematic discovery of neoepitope–HLA pairs for neoantigens shared among patients and tumor types
Systematic discovery of neoepitope–HLA pairs for neoantigens shared among patients and tumor types
A large resource of shared tumor neoepitopes aims to accelerate cancer immunotherapy.
- Hem R. Gurung
- , Amy J. Heidersbach
- & Christopher M. Rose
-
Perspective |
 The chemistry of next-generation sequencing
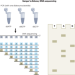
The chemistry of next-generation sequencing
A historical perspective on how next-generation sequencing chemistry was developed.
- Raphaël Rodriguez
- & Yamuna Krishnan
-
Article
| Open Access Single-cell lineage capture across genomic modalities with CellTag-multi reveals fate-specific gene regulatory changes
Single-cell lineage capture across genomic modalities with CellTag-multi reveals fate-specific gene regulatory changes
Lineage tracing using both transcriptomics and chromatin accessibility provides mechanistic insights into cell fate.
- Kunal Jindal
- , Mohd Tayyab Adil
- & Samantha A. Morris
-
Research Briefing |
 Fast and accurate identification of plasmids and viruses in sequencing data using geNomad
Fast and accurate identification of plasmids and viruses in sequencing data using geNomad
The detection of mobile genetic elements is crucial for exploring the ecology and evolution of microbial communities, and it has diverse implications in biotechnology and public health. geNomad is a computational framework that enables researchers to precisely identifiy and annotate plasmids and viruses in sequencing data on a large scale.
-
Correspondence |
A new chance for genome editing in Europe
- Hervé Vanderschuren
- , Patience Chatukuta
- & Devang Mehta
-
Patents |
 The global patent landscape of mRNA for diagnosis and therapy
The global patent landscape of mRNA for diagnosis and therapy
The COVID-19 pandemic brought mRNA vaccines to market in a short period, pointing the entire drug development field in the direction of mRNA treatment.
- Mengru Lyu
- , Jiyuan Chen
- & Yuan Gao
-
Brief Communication
| Open Access Spatial multimodal analysis of transcriptomes and metabolomes in tissues
Spatial multimodal analysis of transcriptomes and metabolomes in tissues
Metabolites and RNA in a tissue section are profiled simultaneously.
- Marco Vicari
- , Reza Mirzazadeh
- & Joakim Lundeberg
-
Article
| Open Access Imaging brain tissue architecture across millimeter to nanometer scales
Imaging brain tissue architecture across millimeter to nanometer scales
Mapping fixed brain samples with extracellular labeling and optical microscopy reveals synaptic connections.
- Julia M. Michalska
- , Julia Lyudchik
- & Johann G. Danzl
-
Article |
 Joint single-cell profiling resolves 5mC and 5hmC and reveals their distinct gene regulatory effects
Joint single-cell profiling resolves 5mC and 5hmC and reveals their distinct gene regulatory effects
Simultaneous single-cell profiling of 5hmC and 5mC shows their unique regulatory roles.
- Emily B. Fabyanic
- , Peng Hu
- & Hao Wu
-
Correspondence |
The digital and analog worlds of protein engineering
- Lada Nuzhna
- & Tess van Stekelenburg
-
Research Briefing |
 Benchmarking of single-cell ATAC sequencing tools
Benchmarking of single-cell ATAC sequencing tools
A systematic examination of eight different single-cell assay for transposase-accessible chromatin by sequencing (scATAC-seq) technologies revealed marked differences in the complexity of sequencing libraries and the specificity of DNA tagmentation that they achieve. Our pipeline for universal mapping of scATAC-seq data (PUMATAC) allowed a fair benchmarking of existing methods and enables the seamless integration of future datasets and technologies.
-
Brief Communication
| Open Access Near-infrared co-illumination of fluorescent proteins reduces photobleaching and phototoxicity
Near-infrared co-illumination of fluorescent proteins reduces photobleaching and phototoxicity
A dual illumination method reduces photobleaching for green and yellow fluorescent proteins.
- Lucie Ludvikova
- , Emma Simon
- & Agathe Espagne
-
Article |
 High-efficiency transgene integration by homology-directed repair in human primary cells using DNA-PKcs inhibition
High-efficiency transgene integration by homology-directed repair in human primary cells using DNA-PKcs inhibition
A small molecule enhances targeted gene integration at therapeutically relevant loci in human primary cells.
- Sridhar Selvaraj
- , William N. Feist
- & Matthew H. Porteus
-
Article |
 PhyloVelo enhances transcriptomic velocity field mapping using monotonically expressed genes
PhyloVelo enhances transcriptomic velocity field mapping using monotonically expressed genes
A refined velocity model improves cell fate mapping with lineage-traced scRNA-seq data.
- Kun Wang
- , Liangzhen Hou
- & Zheng Hu
-
Matters Arising |
 Reply to: Methodological concerns and lack of evidence for single-synapse RNA-seq
Reply to: Methodological concerns and lack of evidence for single-synapse RNA-seq
- Muchun Niu
- & Chenghang Zong
-
Research Briefing |
 Breaking boundaries in whole-body imaging and disease understanding with wildDISCO
Breaking boundaries in whole-body imaging and disease understanding with wildDISCO
Wild three-dimensional imaging of solvent-cleared organs (wildDISCO) uses conventional antibodies for whole-body staining in mice, creating comprehensive biological atlases of nerves, blood vessels and lymphatics. It uncovers pathological changes, such as tertiary lymphoid structures in cancer, and it enables the precise tracking of therapeutic molecules and cells, enhancing our understanding of disease pathology and treatment.
Browse broader subjects
Browse narrower subjects
- Analytical biochemistry
- Behavioural methods
- Bioinformatics
- Biological models
- Biophysical methods
- Cytological techniques
- Electrophysiology
- Epigenetics analysis
- Experimental organisms
- Gene delivery
- Gene expression analysis
- Genetic engineering
- Genetic techniques
- Genomic analysis
- High-throughput screening
- Imaging
- Immunological techniques
- Isolation, separation and purification
- Lab-on-a-chip
- Mass spectrometry
- Metabolomics
- Microbiology techniques
- Microscopy
- Molecular engineering
- Nanobiotechnology
- Optogenetics
- Proteomic analysis
- Sensors and probes
- Sequencing
- Software
- Optical spectroscopy
- Structure determination

 Decoding cell replicational age from single-cell ATAC-seq data
Decoding cell replicational age from single-cell ATAC-seq data