Article
|
Open Access
Featured
-
-
Article
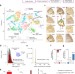
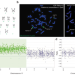
| Open Access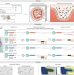 Spatial host–microbiome sequencing reveals niches in the mouse gut
Spatial host–microbiome sequencing reveals niches in the mouse gut
Spatial host–microbiome sequencing simultaneously profiles microbes and host transcriptomes from mouse colons.
- Britta Lötstedt
- , Martin Stražar
- & Sanja Vickovic
-
Research Briefing |
 Compressed Perturb-seq enables highly efficient genetic screens
Compressed Perturb-seq enables highly efficient genetic screens
Using an experimental and computational framework inspired by compressed sensing, we greatly reduced the number of measurements needed to run Perturb-seq. Our compressed Perturb-seq strategy relies on collecting measurements comprising random linear combinations of genetic perturbations, followed by deconvolving the perturbation effects on the transcriptome using sparsity-exploiting algorithms.
-
Brief Communication
| Open Access Spatial multimodal analysis of transcriptomes and metabolomes in tissues
Spatial multimodal analysis of transcriptomes and metabolomes in tissues
Metabolites and RNA in a tissue section are profiled simultaneously.
- Marco Vicari
- , Reza Mirzazadeh
- & Joakim Lundeberg
-
News & Views |
 Subtle cell states resolved in single-cell data
Subtle cell states resolved in single-cell data
SEACells identifies rare cell states in large datasets, enabling atlas-scale studies.
- Caleb Lareau
-
Article
| Open Access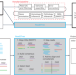 Inference of phylogenetic trees directly from raw sequencing reads using Read2Tree
Inference of phylogenetic trees directly from raw sequencing reads using Read2Tree
Phylogenetic trees are generated from sequencing reads without genome assembly or annotation.
- David Dylus
- , Adrian Altenhoff
- & Christophe Dessimoz
-
Brief Communication
| Open Access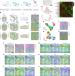 High-plex protein and whole transcriptome co-mapping at cellular resolution with spatial CITE-seq
High-plex protein and whole transcriptome co-mapping at cellular resolution with spatial CITE-seq
Co-indexing of transcriptomes and epitopes is extended to the spatial dimension with large protein panels.
- Yang Liu
- , Marcello DiStasio
- & Rong Fan
-
Brief Communication
| Open Access Solid-phase capture and profiling of open chromatin by spatial ATAC
Solid-phase capture and profiling of open chromatin by spatial ATAC
Chromatin accessibility profiles are spatially resolved in tissue sections.
- Enric Llorens-Bobadilla
- , Margherita Zamboni
- & Patrik L. Ståhl
-
Article |
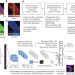 Control-independent mosaic single nucleotide variant detection with DeepMosaic
Control-independent mosaic single nucleotide variant detection with DeepMosaic
A deep learning model detects low frequency mosaic variants with improved accuracy.
- Xiaoxu Yang
- , Xin Xu
- & Joseph G. Gleeson
-
Research Briefing |
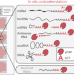 Spatial total RNA-sequencing maps coding, noncoding and viral RNAs in tissues
Spatial total RNA-sequencing maps coding, noncoding and viral RNAs in tissues
Spatial total RNA-sequencing (STRS) combines in situ polyadenylation with existing spatial transcriptomics technologies to enable a broader view of the transcriptome in tissues. We use STRS to spatially map coding, noncoding and nonhost RNAs in models of skeletal muscle regeneration and viral myocarditis.
-
Article |
 A comparison of experimental assays and analytical methods for genome-wide identification of active enhancers
A comparison of experimental assays and analytical methods for genome-wide identification of active enhancers
Experimental and computational methods to detect enhancer RNAs are benchmarked.
- Li Yao
- , Jin Liang
- & Haiyuan Yu
-
Correspondence |
 Ready-to-use public infrastructure for global SARS-CoV-2 monitoring
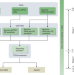
Ready-to-use public infrastructure for global SARS-CoV-2 monitoring
- Wolfgang Maier
- , Simon Bray
- & Anton Nekrutenko
-
Article |
 Scalable, multimodal profiling of chromatin accessibility, gene expression and protein levels in single cells
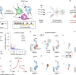
Scalable, multimodal profiling of chromatin accessibility, gene expression and protein levels in single cells
Chromatin accessibility, gene expression and protein levels are measured in the same single cell.
- Eleni P. Mimitou
- , Caleb A. Lareau
- & Peter Smibert
-
Letter
| Open Access Fully phased human genome assembly without parental data using single-cell strand sequencing and long reads
Fully phased human genome assembly without parental data using single-cell strand sequencing and long reads
Assembly of haplotype-resolved human genomes is achieved by combining short and long reads.
- David Porubsky
- , Peter Ebert
- & Tobias Marschall
-
Analysis |
 Systematic comparison of single-cell and single-nucleus RNA-sequencing methods
Systematic comparison of single-cell and single-nucleus RNA-sequencing methods
Seven methods for single-cell RNA sequencing are benchmarked on cell lines, primary cells and mouse cortex.
- Jiarui Ding
- , Xian Adiconis
- & Joshua Z. Levin
-
Letter |
 Accurate detection of mosaic variants in sequencing data without matched controls
Accurate detection of mosaic variants in sequencing data without matched controls
MosaicForecast detects mosaic single-nucleotide variants and indels in human samples.
- Yanmei Dou
- , Minseok Kwon
- & Peter J. Park
-
Letter |
 Single-cell multiomic analysis identifies regulatory programs in mixed-phenotype acute leukemia
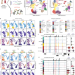
Single-cell multiomic analysis identifies regulatory programs in mixed-phenotype acute leukemia
Analyzing DNA accessibility, transcriptome and protein expression in single cells uncovers cancer-regulatory programs.
- Jeffrey M. Granja
- , Sandy Klemm
- & William J. Greenleaf
-
Letter |
 Genome-wide screening for functional long noncoding RNAs in human cells by Cas9 targeting of splice sites
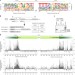
Genome-wide screening for functional long noncoding RNAs in human cells by Cas9 targeting of splice sites
Essential long noncoding RNAs are identified using Cas9 targeting of splice sites.
- Ying Liu
- , Zhongzheng Cao
- & Wensheng Wei
-
Letter |
 Single-cell isoform RNA sequencing characterizes isoforms in thousands of cerebellar cells
Single-cell isoform RNA sequencing characterizes isoforms in thousands of cerebellar cells
Full-length spliced RNA isoforms are identified in thousands of single cerebellar cells.
- Ishaan Gupta
- , Paul G Collier
- & Hagen U Tilgner
-
News & Views |
 Profiling DNA–transcription factor interactions
Profiling DNA–transcription factor interactions
Global transcription factor activity is measured by screening cell extracts with an oligonucleotide library.
- Cheen Euong Ang
- & Marius Wernig
-
News & Views |
 An archive written in DNA
An archive written in DNA
New research sets a world record in the volume of data stored in and retrieved from DNA.
- Reinhard Heckel
-
News Feature |
 DNA makes an appearance
DNA makes an appearance
How much can genome variation reveal about the appearance of an individual? Laura DeFrancesco investigates the field of molecular photofitting.
- Laura DeFrancesco
-
Article |
 GRID-seq reveals the global RNA–chromatin interactome
GRID-seq reveals the global RNA–chromatin interactome
The RNAs bound to the genome and their binding sites are detected with GRID-seq.
- Xiao Li
- , Bing Zhou
- & Xiang-Dong Fu
-
News & Views |
 Core promoters across the genome
Core promoters across the genome
Promoter strength is measured genome-wide with two high-throughput reporter assays.
- Nevena Cvetesic
- & Boris Lenhard
-
Article |
 Genome-wide mapping of mutations at single-nucleotide resolution for protein, metabolic and genome engineering
Genome-wide mapping of mutations at single-nucleotide resolution for protein, metabolic and genome engineering
An approach called CREATE enables multiplex genome engineering, protein engineering and mapping of mutations in bacterial and yeast cells.
- Andrew D Garst
- , Marcelo C Bassalo
- & Ryan T Gill
-
-
Letter |
 Single-cell 5hmC sequencing reveals chromosome-wide cell-to-cell variability and enables lineage reconstruction
Single-cell 5hmC sequencing reveals chromosome-wide cell-to-cell variability and enables lineage reconstruction
A genome-wide, strand-specific sequencing method to detect 5-hydroxymethylcytosine marks in single cells is developed.
- Dylan Mooijman
- , Siddharth S Dey
- & Alexander van Oudenaarden
-
Letter |
 High-throughput determination of RNA structure by proximity ligation
High-throughput determination of RNA structure by proximity ligation
RNA structure is revealed by a high-throughput method that relies on proximity ligation and deep sequencing.
- Vijay Ramani
- , Ruolan Qiu
- & Jay Shendure
-
-
-
News & Views |
 Mapping the precision of genome editing
Mapping the precision of genome editing
Genome-wide, unbiased methods provide a comprehensive picture of off-target cleavage by engineered nucleases.
- Richard Gabriel
- , Christof von Kalle
- & Manfred Schmidt
-
Article |
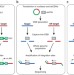
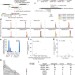 GUIDE-seq enables genome-wide profiling of off-target cleavage by CRISPR-Cas nucleases
GUIDE-seq enables genome-wide profiling of off-target cleavage by CRISPR-Cas nucleases
An unbiased approach for the genome-wide detection of off-target cleavage by CRISPR-Cas9 RNA–guided nucleases reveals wide variability in the off-target activity of different guide RNAs.
- Shengdar Q Tsai
- , Zongli Zheng
- & J Keith Joung
-
-
Correspondence |
Reply to Just the facts, please
-
-
Correspondence |
Reply to Rational drug repositioning by medical genetics
- Philippe Sanseau
- , Pankaj Agarwal
- & Vincent Mooser
-
Review Article |
 Pharmacogenomics in clinical practice and drug development
Pharmacogenomics in clinical practice and drug development
- Andrew R Harper
- & Eric J Topol
-
Letter |
 Associative transcriptomics of traits in the polyploid crop species Brassica napus
Associative transcriptomics of traits in the polyploid crop species Brassica napus
Sequencing a genome and identifying genetic markers lays the groundwork for genome-wide association studies, but can be difficult to achieve for polyploid species. Harper et al. present an approach for performing association studies using genetic maps and markers generated from transcriptome sequencing data alone and apply it to the polyploid crop Brassica napus.
- Andrea L Harper
- , Martin Trick
- & Ian Bancroft
-
Correspondence |
 Use of genome-wide association studies for drug repositioning
Use of genome-wide association studies for drug repositioning
- Philippe Sanseau
- , Pankaj Agarwal
- & Vincent Mooser
-
Correspondence |
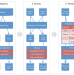 Detecting and annotating genetic variations using the HugeSeq pipeline
Detecting and annotating genetic variations using the HugeSeq pipeline
- Hugo Y K Lam
- , Cuiping Pan
- & Michael Snyder
-
Resource |
 Family-wide chemical profiling and structural analysis of PARP and tankyrase inhibitors
Family-wide chemical profiling and structural analysis of PARP and tankyrase inhibitors
PARP inhibitors have recently entered phase 3 clinical trials as cancer therapeutics, but the specificity of many of these compounds is unknown. Wahlberg et al. used biochemical approaches to show that most PARP inhibitors target multiple PARP family members.
- Elisabet Wahlberg
- , Tobias Karlberg
- & Johan Weigelt
-
Correspondence |
 Recurrent copy number variations in human induced pluripotent stem cells
Recurrent copy number variations in human induced pluripotent stem cells
- Kristen Martins-Taylor
- , Benjamin S Nisler
- & Ren-He Xu
-
Letter |
 Haplotype-resolved genome sequencing of a Gujarati Indian individual
Haplotype-resolved genome sequencing of a Gujarati Indian individual
Sequencing a human genome using next-generation methods does not distinguish between the two copies of each chromosome. Kitzman et al. determine a haplotype-resolved genome sequence by efficiently constructing and sequencing long-insert clones that cover the diploid genome with a low likelihood of overlap.
- Jacob O Kitzman
- , Alexandra P MacKenzie
- & Jay Shendure
-
Editorial |
Making a mark
High-throughput technologies are enabling epigenetic modifications to be mapped on a genome-wide scale, but whether such knowledge can be rapidly translated into biomedical applications remains unclear.
-
Resource |
 High-resolution DNA analysis of human embryonic stem cell lines reveals culture-induced copy number changes and loss of heterozygosity
High-resolution DNA analysis of human embryonic stem cell lines reveals culture-induced copy number changes and loss of heterozygosity
Cultured human embryonic stem cells often acquire chromosomal abnormalities that could be detrimental in certain applications. Närvä et al. report the highest-resolution genetic analysis of these cells to date and identify genes whose expression is altered by culture-induced genetic changes.
- Elisa Närvä
- , Reija Autio
- & Riitta Lahesmaa
-
News & Views |
 ChIPs and regulatory bits
ChIPs and regulatory bits
Machine learning reveals combinatorial patterns of transcription factor binding that drive gene expression.
- Xin He
- & Saurabh Sinha
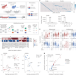
 Bidirectional epigenetic editing reveals hierarchies in gene regulation
Bidirectional epigenetic editing reveals hierarchies in gene regulation Strata rolls out drug-matching service, offers free tumor profiling
Strata rolls out drug-matching service, offers free tumor profiling NCI-MATCH pairs tumor mutations with matching drugs
NCI-MATCH pairs tumor mutations with matching drugs Roche spends $1 billion on Foundation Medicine's tumor profiling
Roche spends $1 billion on Foundation Medicine's tumor profiling Rational drug repositioning by medical genetics
Rational drug repositioning by medical genetics