Research Briefing |
Featured
-
-
Article
| Open Access Tracking single-cell evolution using clock-like chromatin accessibility loci
Tracking single-cell evolution using clock-like chromatin accessibility loci
EpiTrace infers cell mitotic age and evolution from single-cell ATAC-seq data.
- Yu Xiao
- , Wan Jin
- & Yi Zhang
-
Article
| Open Access Mosaic integration and knowledge transfer of single-cell multimodal data with MIDAS
Mosaic integration and knowledge transfer of single-cell multimodal data with MIDAS
Single-cell, multiomic datasets are integrated using dimensionality reduction, imputation and batch correction.
- Zhen He
- , Shuofeng Hu
- & Xiaomin Ying
-
Research Briefing |
 Fast and accurate identification of plasmids and viruses in sequencing data using geNomad
Fast and accurate identification of plasmids and viruses in sequencing data using geNomad
The detection of mobile genetic elements is crucial for exploring the ecology and evolution of microbial communities, and it has diverse implications in biotechnology and public health. geNomad is a computational framework that enables researchers to precisely identifiy and annotate plasmids and viruses in sequencing data on a large scale.
-
Research Briefing |
 Benchmarking of single-cell ATAC sequencing tools
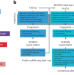
Benchmarking of single-cell ATAC sequencing tools
A systematic examination of eight different single-cell assay for transposase-accessible chromatin by sequencing (scATAC-seq) technologies revealed marked differences in the complexity of sequencing libraries and the specificity of DNA tagmentation that they achieve. Our pipeline for universal mapping of scATAC-seq data (PUMATAC) allowed a fair benchmarking of existing methods and enables the seamless integration of future datasets and technologies.
-
-
Correspondence |
 The scverse project provides a computational ecosystem for single-cell omics data analysis
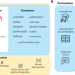
The scverse project provides a computational ecosystem for single-cell omics data analysis
- Isaac Virshup
- , Danila Bredikhin
- & Fabian J. Theis
-
Article |
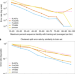 Using deep learning to annotate the protein universe
Using deep learning to annotate the protein universe
A deep learning model predicts protein functional annotations for unaligned amino acid sequences.
- Maxwell L. Bileschi
- , David Belanger
- & Lucy J. Colwell
-
Article
| Open Access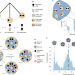 Fluctuating methylation clocks for cell lineage tracing at high temporal resolution in human tissues
Fluctuating methylation clocks for cell lineage tracing at high temporal resolution in human tissues
Lineage tracing of human stem cells is achieved by measuring fluctuating DNA methylation.
- Calum Gabbutt
- , Ryan O. Schenck
- & Darryl Shibata
-
Article |
 Identifying phenotype-associated subpopulations by integrating bulk and single-cell sequencing data
Identifying phenotype-associated subpopulations by integrating bulk and single-cell sequencing data
Bulk and single cell measurements are integrated to identify phenotype-associated subpopulations of cells.
- Duanchen Sun
- , Xiangnan Guan
- & Zheng Xia
-
Review Article |
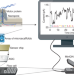 Nanopore sequencing technology, bioinformatics and applications
Nanopore sequencing technology, bioinformatics and applications
Au and colleagues outline the field of nanopore sequencing.
- Yunhao Wang
- , Yue Zhao
- & Kin Fai Au
-
Review Article |
 Identification of tumor antigens with immunopeptidomics
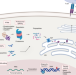
Identification of tumor antigens with immunopeptidomics
Chong et al. review how the integration of mass spectrometry with proteogenomic approaches can identify noncanonical antigens.
- Chloe Chong
- , George Coukos
- & Michal Bassani-Sternberg
-
Article |
 Performance assessment of DNA sequencing platforms in the ABRF Next-Generation Sequencing Study
Performance assessment of DNA sequencing platforms in the ABRF Next-Generation Sequencing Study
Next-generation sequencing platforms are benchmarked using human, bacterial and metagenomics reference materials.
- Jonathan Foox
- , Scott W. Tighe
- & Christopher E. Mason
-
Resource |
 A pan-cancer landscape of somatic mutations in non-unique regions of the human genome
A pan-cancer landscape of somatic mutations in non-unique regions of the human genome
Cancer mutations in non-unique sequences are identified by machine learning on short-read data.
- Maxime Tarabichi
- , Jonas Demeulemeester
- & Tomasz Konopka
-
Article |
 Single-cell CUT&Tag profiles histone modifications and transcription factors in complex tissues
Single-cell CUT&Tag profiles histone modifications and transcription factors in complex tissues
An improved method for single-cell analysis of histone modifications is applied to the mouse brain.
- Marek Bartosovic
- , Mukund Kabbe
- & Gonçalo Castelo-Branco
-
Matters Arising |
Reply to: UMI or not UMI, that is the question for scRNA-seq zero-inflation
- Valentine Svensson
-
Matters Arising |
 Reply to: EEG-based model and antidepressant response
Reply to: EEG-based model and antidepressant response
- Wei Wu
- , Diego A. Pizzagall
- & Amit Etkin
-
Letter
| Open Access Fully phased human genome assembly without parental data using single-cell strand sequencing and long reads
Fully phased human genome assembly without parental data using single-cell strand sequencing and long reads
Assembly of haplotype-resolved human genomes is achieved by combining short and long reads.
- David Porubsky
- , Peter Ebert
- & Tobias Marschall
-
-
Article |
 Massively parallel single-cell mitochondrial DNA genotyping and chromatin profiling
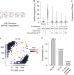
Massively parallel single-cell mitochondrial DNA genotyping and chromatin profiling
Combining droplet-based ATAC-seq and mitochondrial DNA sequencing reveals clonal variation in human cells and tissues.
- Caleb A. Lareau
- , Leif S. Ludwig
- & Vijay G. Sankaran
-
Resource |
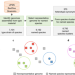 A complete domain-to-species taxonomy for Bacteria and Archaea
A complete domain-to-species taxonomy for Bacteria and Archaea
A full species classification is built for all publicly available bacterial and archaeal genomes.
- Donovan H. Parks
- , Maria Chuvochina
- & Philip Hugenholtz
-
Analysis |
 Systematic comparison of single-cell and single-nucleus RNA-sequencing methods
Systematic comparison of single-cell and single-nucleus RNA-sequencing methods
Seven methods for single-cell RNA sequencing are benchmarked on cell lines, primary cells and mouse cortex.
- Jiarui Ding
- , Xian Adiconis
- & Joshua Z. Levin
-
Article |
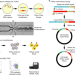 Massively parallel interrogation and mining of natively paired human TCRαβ repertoires
Massively parallel interrogation and mining of natively paired human TCRαβ repertoires
T cell receptors are identified in large-scale screening of T cell repertoires cloned from primary human T cells.
- Matthew J. Spindler
- , Ayla L. Nelson
- & David S. Johnson
-
-
Letter |
 A DNA-of-things storage architecture to create materials with embedded memory
A DNA-of-things storage architecture to create materials with embedded memory
A DNA-based method for embedding data in materials enables the conversion of everyday objects into data storage devices.
- Julian Koch
- , Silvan Gantenbein
- & Robert N. Grass
-
News & Views |
 Exploring a world of a thousand dimensions
Exploring a world of a thousand dimensions
PHATE enhances the visualization of high-dimensional data.
- Catalina A. Vallejos
-
News & Views |
 Profiling DNA–transcription factor interactions
Profiling DNA–transcription factor interactions
Global transcription factor activity is measured by screening cell extracts with an oligonucleotide library.
- Cheen Euong Ang
- & Marius Wernig
-
Correspondence |
 Antigen receptor repertoire profiling from RNA-seq data
Antigen receptor repertoire profiling from RNA-seq data
- Dmitriy A Bolotin
- , Stanislav Poslavsky
- & Dmitriy M Chudakov
-
Perspective |
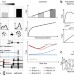 Challenges in long-term imaging and quantification of single-cell dynamics
Challenges in long-term imaging and quantification of single-cell dynamics
The experimental and computational tools that enable continuous imaging of single cells for days and weeks have advanced rapidly in recent years, and solutions to current limitations are on the horizon.
- Stavroula Skylaki
- , Oliver Hilsenbeck
- & Timm Schroeder
-
Analysis |
 A multicenter study benchmarks software tools for label-free proteome quantification
A multicenter study benchmarks software tools for label-free proteome quantification
LFQbench, a software tool to assess the quality of label-free quantitative proteomics analyses, enables developers to benchmark and improve analytic methods.
- Pedro Navarro
- , Jörg Kuharev
- & Stefan Tenzer
-
News & Views |
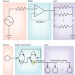 Lightening the load in synthetic biology
Lightening the load in synthetic biology
A new biological device known as a 'load driver' improves the performance of synthetic circuits by insulating genetic parts from each other.
- Eric Klavins
-
Commentary |
 In silico research in the era of cloud computing
In silico research in the era of cloud computing
Snapshots of computer systems that are stored and shared 'in the cloud' could make computational analyses more reproducible.
- Joel T Dudley
- & Atul J Butte
-
Letter |
 Monoclonal antibodies isolated without screening by analyzing the variable-gene repertoire of plasma cells
Monoclonal antibodies isolated without screening by analyzing the variable-gene repertoire of plasma cells
Isolation of antigen-specific antibodies or antibody fragments, whether through B-cell immortalization or recombinant libraries, generally requires laborious screening. Reddy et al. circumvent this step using high-throughput sequencing of plasma cells and bioinformatic analysis of the variable-gene repertoire.
- Sai T Reddy
- , Xin Ge
- & George Georgiou

 Decoding cell replicational age from single-cell ATAC-seq data
Decoding cell replicational age from single-cell ATAC-seq data Leveling up citizen science
Leveling up citizen science Droplet scRNA-seq is not zero-inflated
Droplet scRNA-seq is not zero-inflated