Featured
-
-
Article
| Open Access Systematic discovery of neoepitope–HLA pairs for neoantigens shared among patients and tumor types
Systematic discovery of neoepitope–HLA pairs for neoantigens shared among patients and tumor types
A large resource of shared tumor neoepitopes aims to accelerate cancer immunotherapy.
- Hem R. Gurung
- , Amy J. Heidersbach
- & Christopher M. Rose
-
Brief Communication
| Open Access Spatial multimodal analysis of transcriptomes and metabolomes in tissues
Spatial multimodal analysis of transcriptomes and metabolomes in tissues
Metabolites and RNA in a tissue section are profiled simultaneously.
- Marco Vicari
- , Reza Mirzazadeh
- & Joakim Lundeberg
-
Article |
 Post-translational modifications reshape the antigenic landscape of the MHC I immunopeptidome in tumors
Post-translational modifications reshape the antigenic landscape of the MHC I immunopeptidome in tumors
A computational pipeline identifies tumor antigen post-translational modifications guiding immune responses.
- Assaf Kacen
- , Aaron Javitt
- & Yifat Merbl
-
Research Briefing |
 Framework for multiplicative scaling of single-cell proteomics
Framework for multiplicative scaling of single-cell proteomics
Many biomedical questions demand scalable, deep, and accurate proteome analysis of small samples, including single cells. A scalable framework of multiplexed data-independent acquisition for mass spectrometry enables time saving by parallel analysis of both peptide ions and protein samples, thereby realizing multiplicative gains in throughput.
-
Article |
 Increasing the throughput of sensitive proteomics by plexDIA
Increasing the throughput of sensitive proteomics by plexDIA
Proteomics of small sample sizes using data-independent acquisition methods achieves higher throughput with multiplexing.
- Jason Derks
- , Andrew Leduc
- & Nikolai Slavov
-
Brief Communication |
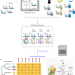 Enhancing untargeted metabolomics using metadata-based source annotation
Enhancing untargeted metabolomics using metadata-based source annotation
Metabolomics is improved by using a reference library of both known and unknown molecules.
- Julia M. Gauglitz
- , Kiana A. West
- & Pieter C. Dorrestein
-
Brief Communication |
 Auto-deconvolution and molecular networking of gas chromatography–mass spectrometry data
Auto-deconvolution and molecular networking of gas chromatography–mass spectrometry data
A machine learning workflow enables auto-deconvolution of gas chromatography–mass spectrometry data.
- Alexander A. Aksenov
- , Ivan Laponogov
- & Kirill Veselkov
-
Letter |
 Detection of SARS-CoV-2 in nasal swabs using MALDI-MS
Detection of SARS-CoV-2 in nasal swabs using MALDI-MS
SARS-CoV-2 is reliably detected in nasal swab samples using mass spectrometry and machine learning analysis.
- Fabiane M. Nachtigall
- , Alfredo Pereira
- & Leonardo S. Santos
-
Brief Communication |
 A lipidome atlas in MS-DIAL 4
A lipidome atlas in MS-DIAL 4
Mass spectral fragmentations of lipids across 117 lipid subclasses are presented in a lipidome atlas.
- Hiroshi Tsugawa
- , Kazutaka Ikeda
- & Makoto Arita
-
Resource |
 Global landscape of cell envelope protein complexes in Escherichia coli
Global landscape of cell envelope protein complexes in Escherichia coli
A survey of cell envelope proteins and their interacting partners in E. coli by mass spectrometry reveals insights into essential bacterial processes.
- Mohan Babu
- , Cedoljub Bundalovic-Torma
- & Andrew Emili
-
Analysis |
 A multicenter study benchmarks software tools for label-free proteome quantification
A multicenter study benchmarks software tools for label-free proteome quantification
LFQbench, a software tool to assess the quality of label-free quantitative proteomics analyses, enables developers to benchmark and improve analytic methods.
- Pedro Navarro
- , Jörg Kuharev
- & Stefan Tenzer
-
Resource |
 Mitochondrial protein functions elucidated by multi-omic mass spectrometry profiling
Mitochondrial protein functions elucidated by multi-omic mass spectrometry profiling
Proteomics, lipidomics and metabolomics of single gene deletion yeast strains sheds light on mitochondrial protein biology.
- Jonathan A Stefely
- , Nicholas W Kwiecien
- & Joshua J Coon
-
Perspective |
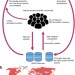 Sharing and community curation of mass spectrometry data with Global Natural Products Social Molecular Networking
Sharing and community curation of mass spectrometry data with Global Natural Products Social Molecular Networking
GNPS is an open-access community-curated analysis platform for sharing natural product mass spectrometry data that enables continuous, automatic reanalysis of deposited 'living' data sets.
- Mingxun Wang
- , Jeremy J Carver
- & Nuno Bandeira
-
News Feature |
 Technology comes to typing
Technology comes to typing
As mass spectrometry makes inroads into pathogen identification in the clinical laboratory, deep sequencing—even nanopore sequencing—is waiting in the wings. Jeffrey L. Fox investigates.
- Jeffrey L Fox
-
Correspondence |
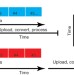 Metabolomic data streaming for biology-dependent data acquisition
Metabolomic data streaming for biology-dependent data acquisition
- Duane Rinehart
- , Caroline H Johnson
- & Gary Siuzdak
-
-
Correspondence |
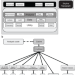 A cross-platform toolkit for mass spectrometry and proteomics
A cross-platform toolkit for mass spectrometry and proteomics
- Matthew C Chambers
- , Brendan Maclean
- & Parag Mallick
-
News & Views |
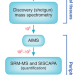 Streamlining biomarker discovery
Streamlining biomarker discovery
Integrating targeted mass spectrometry into proteomic pipelines speeds the validation of disease biomarkers.
- Martin Latterich
- & Jan E Schnitzer
-
Correspondence |
 ProHits: integrated software for mass spectrometry–based interaction proteomics
ProHits: integrated software for mass spectrometry–based interaction proteomics
- Guomin Liu
- , Jianping Zhang
- & Anne-Claude Gingras
-
Perspective |
 Options and considerations when selecting a quantitative proteomics strategy
Options and considerations when selecting a quantitative proteomics strategy
- Bruno Domon
- & Ruedi Aebersold
-
News & Views |
 Enriching quantitative proteomics with SIN
Enriching quantitative proteomics with SIN
A new metric called the normalized spectral index (SIN) provides a simple way to quantify and compare label-free proteomics data.
- Mihaela E Sardiu
- & Michael P Washburn

 Ultra-fast label-free quantification and comprehensive proteome coverage with narrow-window data-independent acquisition
Ultra-fast label-free quantification and comprehensive proteome coverage with narrow-window data-independent acquisition