Featured
-
-
Article |
 Decoder-seq enhances mRNA capture efficiency in spatial RNA sequencing
Decoder-seq enhances mRNA capture efficiency in spatial RNA sequencing
A spatial transcriptomics method achieves high gene detection sensitivity and flexible resolution.
- Jiao Cao
- , Zhong Zheng
- & Chaoyong Yang
-
Article |
 PhyloVelo enhances transcriptomic velocity field mapping using monotonically expressed genes
PhyloVelo enhances transcriptomic velocity field mapping using monotonically expressed genes
A refined velocity model improves cell fate mapping with lineage-traced scRNA-seq data.
- Kun Wang
- , Liangzhen Hou
- & Zheng Hu
-
Brief Communication |
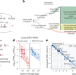 High-throughput RNA isoform sequencing using programmed cDNA concatenation
High-throughput RNA isoform sequencing using programmed cDNA concatenation
Programmable concatenation of cDNA molecules increases the throughput of PacBio sequencing about 15-fold.
- Aziz M. Al’Khafaji
- , Jonathan T. Smith
- & Nir Hacohen
-
Research Briefing |
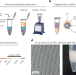 The next generation of single-cell sequencing methods can be microfluidics-free
The next generation of single-cell sequencing methods can be microfluidics-free
Sequencing individual cells in a sample enables scientists to infer the unique characteristics of important subsets. Single-cell sequencing methods that rely on microfluidics for cell barcoding are limited in speed, scale and flexibility. We developed a technique that uses particle-templated emulsification instead of microfluidics and can process millions of cells within minutes.
-
Article
| Open Access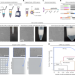 Microfluidics-free single-cell genomics with templated emulsification
Microfluidics-free single-cell genomics with templated emulsification
A microfluidics-free, scalable single-cell RNA-sequencing method produces high-quality transcriptomes.
- Iain C. Clark
- , Kristina M. Fontanez
- & Adam R. Abate
-
Brief Communication
| Open Access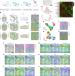 High-plex protein and whole transcriptome co-mapping at cellular resolution with spatial CITE-seq
High-plex protein and whole transcriptome co-mapping at cellular resolution with spatial CITE-seq
Co-indexing of transcriptomes and epitopes is extended to the spatial dimension with large protein panels.
- Yang Liu
- , Marcello DiStasio
- & Rong Fan
-
Article |
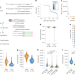 Droplet-based transcriptome profiling of individual synapses
Droplet-based transcriptome profiling of individual synapses
High-throughput profiling of the transcriptomes of individual synapses shows molecular heterogeneity.
- Muchun Niu
- , Wenjian Cao
- & Chenghang Zong
-
Article
| Open Access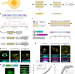 Recording of cellular physiological histories along optically readable self-assembling protein chains
Recording of cellular physiological histories along optically readable self-assembling protein chains
A history of cellular events is recorded in self-assembling protein chains.
- Changyang Linghu
- , Bobae An
- & Edward S. Boyden
-
Research Briefing |
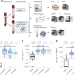 Measuring SARS-CoV-2 T cell immunity with a scalable qPCR-based assay
Measuring SARS-CoV-2 T cell immunity with a scalable qPCR-based assay
We developed two quantitative PCR-based assays to detect SARS-CoV-2-specific T cell immunity: qTACT and dqTACT. The assays quantify CXCL10 mRNA, after incubation of whole blood with viral peptides, as a proxy of an antigen-specific T cell response, and will allow population-level monitoring of cellular immunity to SARS-CoV-2.
-
Article |
 Cell2location maps fine-grained cell types in spatial transcriptomics
Cell2location maps fine-grained cell types in spatial transcriptomics
A Bayesian model maps the location of cell types in tissues with higher sensitivity.
- Vitalii Kleshchevnikov
- , Artem Shmatko
- & Omer Ali Bayraktar
-
Article |
 Single-cell profiling of proteins and chromatin accessibility using PHAGE-ATAC
Single-cell profiling of proteins and chromatin accessibility using PHAGE-ATAC
Protein epitopes and chromatin accessibility are measured in single cells using phage display.
- Evgenij Fiskin
- , Caleb A. Lareau
- & Aviv Regev
-
Brief Communication |
 Multiplexed direct detection of barcoded protein reporters on a nanopore array
Multiplexed direct detection of barcoded protein reporters on a nanopore array
Reporter proteins are easily multiplexed using protein barcodes measured with nanopores.
- Nicolas Cardozo
- , Karen Zhang
- & Jeff Nivala
-
Analysis |
 Bayesian inference of gene expression states from single-cell RNA-seq data
Bayesian inference of gene expression states from single-cell RNA-seq data
A Bayesian procedure overcomes challenges in single-cell RNA-seq data normalization.
- Jérémie Breda
- , Mihaela Zavolan
- & Erik van Nimwegen
-
Article |
 Spatial tissue profiling by imaging-free molecular tomography
Spatial tissue profiling by imaging-free molecular tomography
A refined spatial sampling technique transforms standard RNA sequencing into a spatial transcriptomics method.
- Halima Hannah Schede
- , Christian G. Schneider
- & Gioele La Manno
-
Matters Arising |
Reply to: UMI or not UMI, that is the question for scRNA-seq zero-inflation
- Valentine Svensson
-
Letter |
 RNA timestamps identify the age of single molecules in RNA sequencing
RNA timestamps identify the age of single molecules in RNA sequencing
The age of individual RNA molecules at 1-h resolution is inferred by measuring A-to-I editing.
- Samuel G. Rodriques
- , Linlin M. Chen
- & Fei Chen
-
Article |
 Sci-fate characterizes the dynamics of gene expression in single cells
Sci-fate characterizes the dynamics of gene expression in single cells
Gene expression dynamics in single cells are tracked by labeling newly synthesized RNA in indexed cells.
- Junyue Cao
- , Wei Zhou
- & Jay Shendure
-
-
Article |
 Integrating microarray-based spatial transcriptomics and single-cell RNA-seq reveals tissue architecture in pancreatic ductal adenocarcinomas
Integrating microarray-based spatial transcriptomics and single-cell RNA-seq reveals tissue architecture in pancreatic ductal adenocarcinomas
Combining single-cell RNA-seq data and microarray-based spatial transcriptomics maps the location of different cell types and cell states in pancreatic tumors.
- Reuben Moncada
- , Dalia Barkley
- & Itai Yanai
-
News & Views |
 From single-cell RNA-seq to transcriptional regulation
From single-cell RNA-seq to transcriptional regulation
Analysis of gene expression and chromatin accessibility in single cells provides insight into development and disease.
- Gioele La Manno
-
Article |
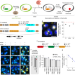 In situ readout of DNA barcodes and single base edits facilitated by in vitro transcription
In situ readout of DNA barcodes and single base edits facilitated by in vitro transcription
The spatial location and sequence of DNA barcodes are detected with high sensitivity in fixed tissues.
- Amjad Askary
- , Luis Sanchez-Guardado
- & Michael B. Elowitz
-
Article |
 Human 5′ UTR design and variant effect prediction from a massively parallel translation assay
Human 5′ UTR design and variant effect prediction from a massively parallel translation assay
Ribosome loading is predictably controlled through design of 5′ UTR sequences.
- Paul J. Sample
- , Ban Wang
- & Georg Seelig
-
Review Article |
 Characterization of noncoding regulatory DNA in the human genome
Characterization of noncoding regulatory DNA in the human genome
Genome-wide mapping of regulatory elements will improve our understanding of how genetic variation in the noncoding genome affects disease phenotypes.
- Ran Elkon
- & Reuven Agami
-
Article |
 Genome-wide assessment of sequence-intrinsic enhancer responsiveness at single-base-pair resolution
Genome-wide assessment of sequence-intrinsic enhancer responsiveness at single-base-pair resolution
STAP-seq quantifies the ability of genomic DNA sequences to convert enhancer activities into transcription initiation events.
- Cosmas D Arnold
- , Muhammad A Zabidi
- & Alexander Stark
-
News & Views |
 Tracking translation one mRNA at a time
Tracking translation one mRNA at a time
Imaging of single mRNA molecules undergoing translation provides insights into the regulation of protein synthesis.
- Jiahui Wu
- & Samie R Jaffrey
-
Letter |
 Integrated genome and transcriptome sequencing of the same cell
Integrated genome and transcriptome sequencing of the same cell
Method for combined genome and transcriptome sequencing from the same single cell shows that copy number variations influence cell-to-cell variability in gene expression levels.
- Siddharth S Dey
- , Lennart Kester
- & Alexander van Oudenaarden
-
News & Views |
 How deep is enough in single-cell RNA-seq?
How deep is enough in single-cell RNA-seq?
Guidelines for determining sequencing depth facilitate transcriptome profiling of single cells in heterogeneous populations.
- Aaron M Streets
- & Yanyi Huang
-
Analysis |
 Detecting and correcting systematic variation in large-scale RNA sequencing data
Detecting and correcting systematic variation in large-scale RNA sequencing data
Li et al. identify the top-performing methods to improve cross-site differential gene expression analysis with RNA-seq.
- Sheng Li
- , Paweł P Łabaj
- & Christopher E Mason
-
Article |
 A comprehensive assessment of RNA-seq accuracy, reproducibility and information content by the Sequencing Quality Control Consortium
A comprehensive assessment of RNA-seq accuracy, reproducibility and information content by the Sequencing Quality Control Consortium
The Sequencing Quality Control (SEQC) consortium shows that junction discovery and differential gene expression profiling with RNA-seq can be robust but transcript-level and absolute measurements remain challenging.
- Zhenqiang Su
- , Paweł P Łabaj
- & Leming Shi
-
Patents |
 DREAMing of a patent-free human genome for clinical sequencing
DREAMing of a patent-free human genome for clinical sequencing
Can methylation be the key to challenging the interpretation of existing gene patent claims? And can a novel PCR method be used to enable the sequencing of hundreds of genes?
- Kevin J McKernan
- , Jessica Spangler
- & Vasisht Tadigotla
-
Letter |
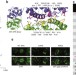 Engineered ascorbate peroxidase as a genetically encoded reporter for electron microscopy
Engineered ascorbate peroxidase as a genetically encoded reporter for electron microscopy
Martell et al. engineer APEX, a protein tag for electron microscopy that does not require light activation, enabling the imaging of subcellular protein localization in large or thick sections.
- Jeffrey D Martell
- , Thomas J Deerinck
- & Alice Y Ting
-
Letter |
 RNA processing enables predictable programming of gene expression
RNA processing enables predictable programming of gene expression
Qi et al. control the expression levels of transgenes using bacterial CRISPR RNA-processing technology.
- Lei Qi
- , Rachel E Haurwitz
- & Adam P Arkin
-
Article |
 Single-cell dissection of transcriptional heterogeneity in human colon tumors
Single-cell dissection of transcriptional heterogeneity in human colon tumors
Not all cells in a tumor are alike, but our ability to characterize cancer heterogeneity in detail has been limited. Dalerba et al. use high-throughput single-cell expression analysis to define clinically relevant subpopulations in normal and cancerous colon tissue.
- Piero Dalerba
- , Tomer Kalisky
- & Stephen R Quake
-
Letter |
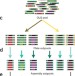 Scalable gene synthesis by selective amplification of DNA pools from high-fidelity microchips
Scalable gene synthesis by selective amplification of DNA pools from high-fidelity microchips
Long DNA molecules, such as those encoding genes, can be assembled from short oligonucleotides created on a microarray. Kosuri et al. improve the fidelity and scalability of this process, enabling synthesis of 40 antibody fragments having repetitive regions and other challenging sequence features.
- Sriram Kosuri
- , Nikolai Eroshenko
- & George M Church
-
Letter |
 Programmable in situ amplification for multiplexed imaging of mRNA expression
Programmable in situ amplification for multiplexed imaging of mRNA expression
The simultaneous detection of multiple mRNA species in thick tissues or whole-mount embryos has remained technically challenging. Choi et al. present a method based on the triggered polymerization of RNA stem-loop structures that allows the distribution of up to five mRNAs in intact zebrafish embryos to be imaged at the same time.
- Harry M T Choi
- , Joann Y Chang
- & Niles A Pierce
-
Article |
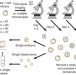 Non-invasive imaging of human embryos before embryonic genome activation predicts development to the blastocyst stage
Non-invasive imaging of human embryos before embryonic genome activation predicts development to the blastocyst stage
The grading systems used by in vitro fertilization clinics cannot determine reliably whether a given embryo will lead to a successful pregnancy. Wong et al. address one part of this problem by showing that development of an embryo to the blastocyst stage can be predicted with high confidence at day 2 post fertilization.
- Connie C Wong
- , Kevin E Loewke
- & Renee A Reijo Pera
-
Editorial |
MAQC-II: analyze that!
The MAQC consortium's latest study suggests that human error in handling DNA microarray data analysis software could delay the technology's wider adoption in the clinic.
-
News & Views |
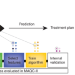 Microarrays in the clinic
Microarrays in the clinic
The MicroArray Quality Control (MAQC) consortium has evaluated methods for making clinically useful predictions from large-scale gene expression data.
- Guy W Tillinghast
-
Article |
 The MicroArray Quality Control (MAQC)-II study of common practices for the development and validation of microarray-based predictive models
The MicroArray Quality Control (MAQC)-II study of common practices for the development and validation of microarray-based predictive models
The Microarray Quality Control consortium pitted 36 teams against each other to evaluate methods for creating genomic classifiers, computational tools for interpreting gene expression profiles. The performance of the classifiers on blinded validation data—and metadata on the analytic methods—reveal the challenges facing the field.
- Leming Shi
- , Gregory Campbell
- & Russell D Wolfinger
-
Feature |
 Proteomics retrenches
Proteomics retrenches
Improvements in technology are making proteomics research less descriptive and more analytic, but the field has yet to deliver on its aspirations.
- Peter Mitchell

 Capturing and modeling cellular niches from dissociated single-cell and spatial data
Capturing and modeling cellular niches from dissociated single-cell and spatial data Droplet scRNA-seq is not zero-inflated
Droplet scRNA-seq is not zero-inflated