Featured
-
-
Brief Communication |
 Simultaneous nanopore profiling of mRNA m6A and pseudouridine reveals translation coordination
Simultaneous nanopore profiling of mRNA m6A and pseudouridine reveals translation coordination
A machine learning pipeline for nanopore sequencing analyzes m6A and Ψ simultaneously.
- Sihao Huang
- , Adam C. Wylder
- & Tao Pan
-
Article
| Open Access KARR-seq reveals cellular higher-order RNA structures and RNA–RNA interactions
KARR-seq reveals cellular higher-order RNA structures and RNA–RNA interactions
RNA interactions and structures are determined with a chemical labeling and crosslinking method.
- Tong Wu
- , Anthony Youzhi Cheng
- & Chuan He
-
Article |
 Decoder-seq enhances mRNA capture efficiency in spatial RNA sequencing
Decoder-seq enhances mRNA capture efficiency in spatial RNA sequencing
A spatial transcriptomics method achieves high gene detection sensitivity and flexible resolution.
- Jiao Cao
- , Zhong Zheng
- & Chaoyong Yang
-
Research Briefing |
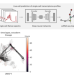 Generating expression profiles of single living cells from Raman microscopy
Generating expression profiles of single living cells from Raman microscopy
Expression profiles of single live cells are generated from Raman microscopy using deep learning, enabling us to track expression dynamics along cell reprogramming or differentiation.
-
Article |
 Prediction of single-cell RNA expression profiles in live cells by Raman microscopy with Raman2RNA
Prediction of single-cell RNA expression profiles in live cells by Raman microscopy with Raman2RNA
Transcriptional profiles of single living cells are generated from Raman imaging data.
- Koseki J. Kobayashi-Kirschvink
- , Charles S. Comiter
- & Aviv Regev
-
Article
| Open Access Ultrafast bisulfite sequencing detection of 5-methylcytosine in DNA and RNA
Ultrafast bisulfite sequencing detection of 5-methylcytosine in DNA and RNA
Accelerated bisulfite sequencing detects 5-methylcytosine in low-input DNA and RNA samples
- Qing Dai
- , Chang Ye
- & Chuan He
-
News & Views |
 Spatial methods for microbiome–host interactions
Spatial methods for microbiome–host interactions
Spatial transcriptomics offers a glimpse of how microbiomes interact with their hosts.
- Ioannis Ntekas
- & Iwijn De Vlaminck
-
Article
| Open Access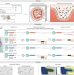 Spatial host–microbiome sequencing reveals niches in the mouse gut
Spatial host–microbiome sequencing reveals niches in the mouse gut
Spatial host–microbiome sequencing simultaneously profiles microbes and host transcriptomes from mouse colons.
- Britta Lötstedt
- , Martin Stražar
- & Sanja Vickovic
-
Perspective |
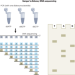 The chemistry of next-generation sequencing
The chemistry of next-generation sequencing
A historical perspective on how next-generation sequencing chemistry was developed.
- Raphaël Rodriguez
- & Yamuna Krishnan
-
News & Views |
 A genome-wide view of disordered proteins
A genome-wide view of disordered proteins
Transcription factors containing disordered regions can now be mapped across the genome, aiding functional studies.
- Benjamin J. Leslie
- , Benjamin Lang
- & M. Madan Babu
-
Article
| Open Access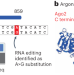 Detection of transcriptome-wide microRNA–target interactions in single cells with agoTRIBE
Detection of transcriptome-wide microRNA–target interactions in single cells with agoTRIBE
Functional miRNA targets are identified transcriptome-wide in single cells.
- Vaishnovi Sekar
- , Emilio Mármol-Sánchez
- & Marc R. Friedländer
-
Research Briefing |
 Electroosmotic flow across nanopores for single-molecule protein sequencing
Electroosmotic flow across nanopores for single-molecule protein sequencing
By using fixed charges to engineer a strong electroosmotic flow, we achieve the unidirectional transport of natural polypeptides across nanopores. Our approach enables native proteins to be transported enzymatically and non-enzymatically in the absence of denaturant and electrophoretic tags, with potential applications for protein sequencing.
-
Article
| Open Access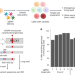 Direct measurement of engineered cancer mutations and their transcriptional phenotypes in single cells
Direct measurement of engineered cancer mutations and their transcriptional phenotypes in single cells
Engineered cancer mutations are linked with phenotypes in a multiplexed single-cell technology.
- Heon Seok Kim
- , Susan M. Grimes
- & Hanlee P. Ji
-
Article
| Open Access Single-nucleotide variant calling in single-cell sequencing data with Monopogen
Single-nucleotide variant calling in single-cell sequencing data with Monopogen
Monopogen identifies single-nucleotide variants in single-cell sequencing data.
- Jinzhuang Dou
- , Yukun Tan
- & Ken Chen
-
Matters Arising |
 Reply to: Methodological concerns and lack of evidence for single-synapse RNA-seq
Reply to: Methodological concerns and lack of evidence for single-synapse RNA-seq
- Muchun Niu
- & Chenghang Zong
-
Brief Communication |
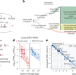 High-throughput RNA isoform sequencing using programmed cDNA concatenation
High-throughput RNA isoform sequencing using programmed cDNA concatenation
Programmable concatenation of cDNA molecules increases the throughput of PacBio sequencing about 15-fold.
- Aziz M. Al’Khafaji
- , Jonathan T. Smith
- & Nir Hacohen
-
News & Views |
 Integration of multi-modal single-cell data
Integration of multi-modal single-cell data
Single-cell data from RNA-seq, chromatin accessibility, DNA methylation and other modalities can be readily integrated using two new methods.
- Michelle Y. Y. Lee
- & Mingyao Li
-
Article
| Open Access Sequencing by avidity enables high accuracy with low reagent consumption
Sequencing by avidity enables high accuracy with low reagent consumption
A sequencing chemistry that separates nucleotide identification from nucleotide incorporation achieves high accuracy.
- Sinan Arslan
- , Francisco J. Garcia
- & Michael Previte
-
News & Views |
 Speed reading the epigenome and genome
Speed reading the epigenome and genome
Dual sequencing of the epigenome and genome could have broad implications in oncology.
- James M. George
- & Arul M. Chinnaiyan
-
Research Briefing |
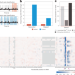 Exploring tRNAs and their modifications and crosstalk using Nano-tRNAseq
Exploring tRNAs and their modifications and crosstalk using Nano-tRNAseq
Nano-tRNAseq is a nanopore-based, cost-effective and high-throughput approach to quantify transfer RNA (tRNA) abundances and modifications simultaneously, providing a framework to study the ‘tRNAome’ at single-molecule resolution. We envision that Nano-tRNAseq will enable us to study the role of tRNA molecules and their modifications in a wide variety of contexts.
-
Article
| Open Access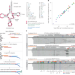 Quantitative analysis of tRNA abundance and modifications by nanopore RNA sequencing
Quantitative analysis of tRNA abundance and modifications by nanopore RNA sequencing
tRNA abundance and chemical modifications are measured simultaneously with nanopores.
- Morghan C. Lucas
- , Leszek P. Pryszcz
- & Eva Maria Novoa
-
Article
| Open Access A relay velocity model infers cell-dependent RNA velocity
A relay velocity model infers cell-dependent RNA velocity
cellDancer enables RNA velocity estimation with cell-specific kinetics.
- Shengyu Li
- , Pengzhi Zhang
- & Guangyu Wang
-
Research Briefing |
 Measuring the impact of chromatin context on transcription factor binding affinities
Measuring the impact of chromatin context on transcription factor binding affinities
We have designed a method, binding affinities to native chromatin by sequencing (BANC-seq), to determine the transcription factor concentrations required for binding to regulatory elements across the genome. Our study shows that chromatin context and DNA accessibility are key regulators of transcription factor binding.
-
Research Briefing |
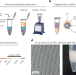 The next generation of single-cell sequencing methods can be microfluidics-free
The next generation of single-cell sequencing methods can be microfluidics-free
Sequencing individual cells in a sample enables scientists to infer the unique characteristics of important subsets. Single-cell sequencing methods that rely on microfluidics for cell barcoding are limited in speed, scale and flexibility. We developed a technique that uses particle-templated emulsification instead of microfluidics and can process millions of cells within minutes.
-
Article
| Open Access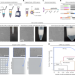 Microfluidics-free single-cell genomics with templated emulsification
Microfluidics-free single-cell genomics with templated emulsification
A microfluidics-free, scalable single-cell RNA-sequencing method produces high-quality transcriptomes.
- Iain C. Clark
- , Kristina M. Fontanez
- & Adam R. Abate
-
Article |
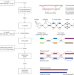 Telomere-to-telomere assembly of diploid chromosomes with Verkko
Telomere-to-telomere assembly of diploid chromosomes with Verkko
Integration of long and ultra-long reads results in improved phased, diploid assemblies.
- Mikko Rautiainen
- , Sergey Nurk
- & Sergey Koren
-
Article |
 Single-cell mapping of combinatorial target antigens for CAR switches using logic gates
Single-cell mapping of combinatorial target antigens for CAR switches using logic gates
CAR-T targets are identified using logic gates and single-cell expression data.
- Joonha Kwon
- , Junho Kang
- & Jung Kyoon Choi
-
Article |
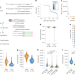 Droplet-based transcriptome profiling of individual synapses
Droplet-based transcriptome profiling of individual synapses
High-throughput profiling of the transcriptomes of individual synapses shows molecular heterogeneity.
- Muchun Niu
- , Wenjian Cao
- & Chenghang Zong
-
Research Briefing |
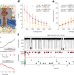 Protein identification using a digestion-free nanopore approach
Protein identification using a digestion-free nanopore approach
Sequencing of proteins is a technically difficult task that typically requires digestion into short peptides before detection and identification. We developed a digestion-free method to chemically unfold and ‘scan’ full-length proteins through a nanopore, producing electrical fingerprints unique to individual protein molecules that are useful in their identification.
-
Article |
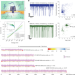 Unidirectional single-file transport of full-length proteins through a nanopore
Unidirectional single-file transport of full-length proteins through a nanopore
Full-length, unfolded proteins are slowly translocated through nanopores without enzymes and fingerprinted.
- Luning Yu
- , Xinqi Kang
- & Meni Wanunu
-
Research Briefing |
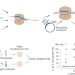 Single-cell recording of cellular RNAs in bacteria
Single-cell recording of cellular RNAs in bacteria
The collection of RNAs in a cell reflects its identity and behavior. Although probing these RNAs in the present is commonplace, peering into the past remains challenging. We developed TIGER (transcribed RNAs inferred by genetically encoded records) to record selected RNAs at the single-cell level, enabling us to connect the past with the present.
-
Article
| Open Access Dynamic, adaptive sampling during nanopore sequencing using Bayesian experimental design
Dynamic, adaptive sampling during nanopore sequencing using Bayesian experimental design
Nanopore selective sequencing with real-time decision updates mitigates coverage bias.
- Lukas Weilguny
- , Nicola De Maio
- & Nick Goldman
-
Brief Communication |
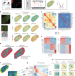 Integration of whole transcriptome spatial profiling with protein markers
Integration of whole transcriptome spatial profiling with protein markers
Visium spatial transcriptomics is combined with markers for more than 30 proteins.
- Nir Ben-Chetrit
- , Xiang Niu
- & Dan A. Landau
-
News & Views |
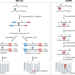 Mapping mRNA modifications for functional studies
Mapping mRNA modifications for functional studies
New methods for mapping the mRNA modifications N6-methyladenosine and pseudouridine enable large-scale functional analyses.
- Joshua D. Jones
- , Daniel E. Eyler
- & Kristin S. Koutmou
-
Article |
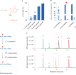 Absolute quantification of single-base m6A methylation in the mammalian transcriptome using GLORI
Absolute quantification of single-base m6A methylation in the mammalian transcriptome using GLORI
The m6A modification is mapped transcriptome-wide at single-base resolution in mammalian cells.
- Cong Liu
- , Hanxiao Sun
- & Jing Wang
-
Brief Communication
| Open Access Modeling intercellular communication in tissues using spatial graphs of cells
Modeling intercellular communication in tissues using spatial graphs of cells
How cells in a tissue communicate is modeled with a graph neural network.
- David S. Fischer
- , Anna C. Schaar
- & Fabian J. Theis
-
Article
| Open Access Quantitative sequencing using BID-seq uncovers abundant pseudouridines in mammalian mRNA at base resolution
Quantitative sequencing using BID-seq uncovers abundant pseudouridines in mammalian mRNA at base resolution
Pseudouridine sites in mRNA are detected at base resolution and functionally investigated.
- Qing Dai
- , Li-Sheng Zhang
- & Chuan He
-
Review Article |
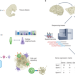 The expanding vistas of spatial transcriptomics
The expanding vistas of spatial transcriptomics
Spatial transcriptomics workflows, metrics and limitations are reviewed and discussed.
- Luyi Tian
- , Fei Chen
- & Evan Z. Macosko
-
Article
| Open Access Mostly natural sequencing-by-synthesis for scRNA-seq using Ultima sequencing
Mostly natural sequencing-by-synthesis for scRNA-seq using Ultima sequencing
A mostly natural sequencing-by-synthesis method is effective for scRNA-seq, particularly in large-scale studies.
- Sean K. Simmons
- , Gila Lithwick-Yanai
- & Joshua Z. Levin
-
Article |
 Comparison and imputation-aided integration of five commercial platforms for targeted DNA methylome analysis
Comparison and imputation-aided integration of five commercial platforms for targeted DNA methylome analysis
Differences in CpG coverage of targeted bisulfite sequencing methods can be overcome using imputation.
- Miljana Tanić
- , Ismail Moghul
- & Stephan Beck
-
Brief Communication
| Open Access Fast and highly sensitive full-length single-cell RNA sequencing using FLASH-seq
Fast and highly sensitive full-length single-cell RNA sequencing using FLASH-seq
FLASH-seq speeds up high-sensitivity scRNA-seq while sparing resources.
- Vincent Hahaut
- , Dinko Pavlinic
- & Simone Picelli
-
Article |
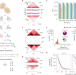 Identifying synergistic high-order 3D chromatin conformations from genome-scale nanopore concatemer sequencing
Identifying synergistic high-order 3D chromatin conformations from genome-scale nanopore concatemer sequencing
High-order chromatin contacts are identified using a combination of 3C, nanopore sequencing and robust statistical analysis.
- Aditya S. Deshpande
- , Netha Ulahannan
- & Marcin Imieliński
-
Article
| Open Access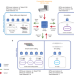 Accelerated identification of disease-causing variants with ultra-rapid nanopore genome sequencing
Accelerated identification of disease-causing variants with ultra-rapid nanopore genome sequencing
A streamlined sequencing process enables identification of disease-causing variants in the clinic within 8 hours.
- Sneha D. Goenka
- , John E. Gorzynski
- & Euan A. Ashley
-
Article |
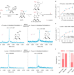 m6A RNA modifications are measured at single-base resolution across the mammalian transcriptome
m6A RNA modifications are measured at single-base resolution across the mammalian transcriptome
m6A-SAC-seq uses chemical labeling to quantify m6A at single-base resolution in the mammalian transcriptome.
- Lulu Hu
- , Shun Liu
- & Chuan He
-
Article |
 Efficient discovery of SARS-CoV-2-neutralizing antibodies via B cell receptor sequencing and ligand blocking
Efficient discovery of SARS-CoV-2-neutralizing antibodies via B cell receptor sequencing and ligand blocking
B cell receptor sequencing with ligand blocking speeds up neutralizing antibody discovery.
- Andrea R. Shiakolas
- , Kevin J. Kramer
- & Ivelin S. Georgiev
-
Brief Communication |
 Mitochondrial variant enrichment from high-throughput single-cell RNA sequencing resolves clonal populations
Mitochondrial variant enrichment from high-throughput single-cell RNA sequencing resolves clonal populations
Clonal dynamics are inferred from mitochondrial variants in primary human cells.
- Tyler E. Miller
- , Caleb A. Lareau
- & Peter van Galen
-
Brief Communication
| Open Access Cell types of origin of the cell-free transcriptome
Cell types of origin of the cell-free transcriptome
Cell types affected by various diseases are inferred from cell-free RNA.
- Sevahn K. Vorperian
- , Mira N. Moufarrej
- & Stephen R. Quake
-
Article |
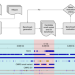 Curated variation benchmarks for challenging medically relevant autosomal genes
Curated variation benchmarks for challenging medically relevant autosomal genes
Variant detection in problematic genes is facilitated with a curated benchmark.
- Justin Wagner
- , Nathan D. Olson
- & Fritz J. Sedlazeck
-
News & Views |
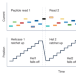 Back and forth with nanopore peptide sequencing
Back and forth with nanopore peptide sequencing
Tagging peptides with DNA allows repeated reads via a helicase in a nanopore with reduced error rates.
- Meni Wanunu

 Mapping medically relevant RNA isoform diversity in the aged human frontal cortex with deep long-read RNA-seq
Mapping medically relevant RNA isoform diversity in the aged human frontal cortex with deep long-read RNA-seq