Featured
-
-
Brief Communication
| Open Access De novo and somatic structural variant discovery with SVision-pro
De novo and somatic structural variant discovery with SVision-pro
SVision-pro enables pairwise genome comparison and improves de novo and somatic structural variant calling.
- Songbo Wang
- , Jiadong Lin
- & Kai Ye
-
Brief Communication
| Open Access Disentanglement of single-cell data with biolord
Disentanglement of single-cell data with biolord
Biolord models the contributions of diverse biological processes to single-cell expression profiles.
- Zoe Piran
- , Niv Cohen
- & Mor Nitzan
-
Article
| Open Access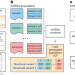 Detection of mosaic and population-level structural variants with Sniffles2
Detection of mosaic and population-level structural variants with Sniffles2
Sniffles2 detects mosaic structural variation from bulk long-read sequencing data.
- Moritz Smolka
- , Luis F. Paulin
- & Fritz J. Sedlazeck
-
Research Briefing |
 Fast and accurate identification of plasmids and viruses in sequencing data using geNomad
Fast and accurate identification of plasmids and viruses in sequencing data using geNomad
The detection of mobile genetic elements is crucial for exploring the ecology and evolution of microbial communities, and it has diverse implications in biotechnology and public health. geNomad is a computational framework that enables researchers to precisely identifiy and annotate plasmids and viruses in sequencing data on a large scale.
-
Article
| Open Access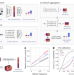 Identification of mobile genetic elements with geNomad
Identification of mobile genetic elements with geNomad
geNomad identifies mobile genetic elements in sequencing data.
- Antonio Pedro Camargo
- , Simon Roux
- & Nikos C. Kyrpides
-
Correspondence |
The digital and analog worlds of protein engineering
- Lada Nuzhna
- & Tess van Stekelenburg
-
Article
| Open Access Single-nucleotide variant calling in single-cell sequencing data with Monopogen
Single-nucleotide variant calling in single-cell sequencing data with Monopogen
Monopogen identifies single-nucleotide variants in single-cell sequencing data.
- Jinzhuang Dou
- , Yukun Tan
- & Ken Chen
-
-
Research Briefing |
 Detecting somatic mutations in single-cell data sets
Detecting somatic mutations in single-cell data sets
We present an algorithm, SComatic, that can be used to directly detect somatic mutations in single-cell data sets without using a reference sample. This method opens the possibility of studying clonal relationships among cells, mutational processes at single-cell resolution, and the impact of somatic mutations on cell function in development and disease.
-
Research Briefing |
 Combining reference genomes into a pangenome graph improves accuracy and reduces bias
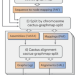
Combining reference genomes into a pangenome graph improves accuracy and reduces bias
Minigraph-Cactus, a method to efficiently combine multiple reference genome assemblies into a pangenome reference graph, can be used to improve accuracy of read mapping and variant calling compared with a single reference in downstream applications.
-
Brief Communication |
 scDesign3 generates realistic in silico data for multimodal single-cell and spatial omics
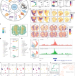
scDesign3 generates realistic in silico data for multimodal single-cell and spatial omics
The challenge of simulating multiomic single-cell data is addressed by a probabilistic model.
- Dongyuan Song
- , Qingyang Wang
- & Jingyi Jessica Li
-
Brief Communication
| Open Access Fast and accurate protein structure search with Foldseek
Fast and accurate protein structure search with Foldseek
Foldseek speeds up protein structural search by four to five orders of magnitude.
- Michel van Kempen
- , Stephanie S. Kim
- & Martin Steinegger
-
Correspondence |
 The scverse project provides a computational ecosystem for single-cell omics data analysis
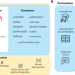
The scverse project provides a computational ecosystem for single-cell omics data analysis
- Isaac Virshup
- , Danila Bredikhin
- & Fabian J. Theis
-
Article
| Open Access A relay velocity model infers cell-dependent RNA velocity
A relay velocity model infers cell-dependent RNA velocity
cellDancer enables RNA velocity estimation with cell-specific kinetics.
- Shengyu Li
- , Pengzhi Zhang
- & Guangyu Wang
-
Brief Communication
| Open Access High-resolution alignment of single-cell and spatial transcriptomes with CytoSPACE
High-resolution alignment of single-cell and spatial transcriptomes with CytoSPACE
CytoSPACE maps individual cells from a reference single-cell RNA sequencing atlas to spatial transcriptomics data.
- Milad R. Vahid
- , Erin L. Brown
- & Aaron M. Newman
-
Article
| Open Access TACCO unifies annotation transfer and decomposition of cell identities for single-cell and spatial omics
TACCO unifies annotation transfer and decomposition of cell identities for single-cell and spatial omics
Annotation transfer from reference to new datasets is improved with a probabilistic approach.
- Simon Mages
- , Noa Moriel
- & Mor Nitzan
-
Brief Communication
| Open Access Accurate isoform discovery with IsoQuant using long reads
Accurate isoform discovery with IsoQuant using long reads
IsoQuant predicts novel isoforms from long-read RNA sequencing.
- Andrey D. Prjibelski
- , Alla Mikheenko
- & Hagen U. Tilgner
-
Article
| Open Access Real-time denoising enables high-sensitivity fluorescence time-lapse imaging beyond the shot-noise limit
Real-time denoising enables high-sensitivity fluorescence time-lapse imaging beyond the shot-noise limit
DeepCAD-RT denoises fluorescence time-lapse images in real time.
- Xinyang Li
- , Yixin Li
- & Qionghai Dai
-
Research Briefing |
 Normalizing cancer RNA-seq data for library size, tumor purity and batch effects
Normalizing cancer RNA-seq data for library size, tumor purity and batch effects
Accurate identification and effective removal of unwanted variation is essential to derive meaningful biological results from large and complex RNA-seq studies. Technical replicates together with negative and positive control genes are key tools for carrying out this task. We show how to proceed when technical replicates are unavailable.
-
Article |
 DeepConsensus improves the accuracy of sequences with a gap-aware sequence transformer
DeepConsensus improves the accuracy of sequences with a gap-aware sequence transformer
Deep learning reduces errors in sequences from PacBio HiFi reads.
- Gunjan Baid
- , Daniel E. Cook
- & Andrew Carroll
-
Article
| Open Access Multi-omics single-cell data integration and regulatory inference with graph-linked embedding
Multi-omics single-cell data integration and regulatory inference with graph-linked embedding
Different single-cell data modalities are integrated at atlas-scale by modeling regulatory interactions.
- Zhi-Jie Cao
- & Ge Gao
-
Research Briefing |
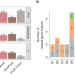 Expanding the spectrum of cancer targets by predicting individual gene fusions
Expanding the spectrum of cancer targets by predicting individual gene fusions
We developed EasyFuse, a computational machine learning pipeline that detects cancer-specific gene fusions with superior performance over existing tools. Individual gene fusions exhibit a high frequency of pre-established CD4+ and CD8+ T-cell responses and thus represent a previously untapped source of neo-antigens that can be exploited for personalized immunotherapies.
-
Brief Communication |
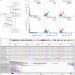 Haplotype-resolved assembly of diploid genomes without parental data
Haplotype-resolved assembly of diploid genomes without parental data
Haplotype-resolved genome assemblies are generated by combining HiFi reads with Hi-C long-range interactions.
- Haoyu Cheng
- , Erich D. Jarvis
- & Heng Li
-
Article |
 CoSpar identifies early cell fate biases from single-cell transcriptomic and lineage information
CoSpar identifies early cell fate biases from single-cell transcriptomic and lineage information
A computational algorithm integrates lineage tracing with single-cell RNA sequencing and improves early cell fate prediction.
- Shou-Wen Wang
- , Michael J. Herriges
- & Allon M. Klein
-
Correspondence |
 A Python library for probabilistic analysis of single-cell omics data
A Python library for probabilistic analysis of single-cell omics data
- Adam Gayoso
- , Romain Lopez
- & Nir Yosef
-
Article
| Open Access A knowledge graph to interpret clinical proteomics data
A knowledge graph to interpret clinical proteomics data
A knowledge graph platform integrates proteomics with other omics data and biomedical databases.
- Alberto Santos
- , Ana R. Colaço
- & Matthias Mann
-
Article |
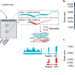 Partitioning RNAs by length improves transcriptome reconstruction from short-read RNA-seq data
Partitioning RNAs by length improves transcriptome reconstruction from short-read RNA-seq data
The quality of RNA sequencing transcriptomes is improved when mRNAs are separated by length.
- Francisca Rojas Ringeling
- , Shounak Chakraborty
- & Stefan Canzar
-
Article |
 Fast and accurate metagenotyping of the human gut microbiome with GT-Pro
Fast and accurate metagenotyping of the human gut microbiome with GT-Pro
Alignment-free SNP calling from metagenomes reduces computational time by two orders of magnitude.
- Zhou Jason Shi
- , Boris Dimitrov
- & Katherine S. Pollard
-
Article |
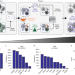 Whole-cell segmentation of tissue images with human-level performance using large-scale data annotation and deep learning
Whole-cell segmentation of tissue images with human-level performance using large-scale data annotation and deep learning
Deep learning algorithms perform as well as humans in identifying cells in tissue images.
- Noah F. Greenwald
- , Geneva Miller
- & David Van Valen
-
Correspondence |
 A Python-based programming language for high-performance computational genomics
A Python-based programming language for high-performance computational genomics
- Ariya Shajii
- , Ibrahim Numanagić
- & Bonnie Berger
-
Article |
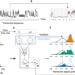 Identification of differential RNA modifications from nanopore direct RNA sequencing with xPore
Identification of differential RNA modifications from nanopore direct RNA sequencing with xPore
m6A RNA modifications are quantified in cancer patient samples and cell lines using nanopore sequencing.
- Ploy N. Pratanwanich
- , Fei Yao
- & Jonathan Göke
-
Article |
 Quantitative profiling of pseudouridylation dynamics in native RNAs with nanopore sequencing
Quantitative profiling of pseudouridylation dynamics in native RNAs with nanopore sequencing
Nanopore sequencing detects pseudouridine and 2′-O-methylation modifications in cellular RNAs.
- Oguzhan Begik
- , Morghan C. Lucas
- & Eva Maria Novoa
-
Article |
 Gene signature extraction and cell identity recognition at the single-cell level with Cell-ID
Gene signature extraction and cell identity recognition at the single-cell level with Cell-ID
Cell-ID facilitates the analysis of cell-type heterogeneity and cell identity across multiple samples at the single-cell level.
- Akira Cortal
- , Loredana Martignetti
- & Antonio Rausell
-
Letter |
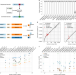 Modular, efficient and constant-memory single-cell RNA-seq preprocessing
Modular, efficient and constant-memory single-cell RNA-seq preprocessing
A preprocessing workflow for single-cell RNA-seq data achieves near-optimal speed.
- Páll Melsted
- , A. Sina Booeshaghi
- & Lior Pachter
-
Analysis |
 A unified haplotype-based method for accurate and comprehensive variant calling
A unified haplotype-based method for accurate and comprehensive variant calling
Octopus detects germline and somatic variants with high sensitivity and accuracy.
- Daniel P. Cooke
- , David C. Wedge
- & Gerton Lunter
-
Article
| Open Access Efficient hybrid de novo assembly of human genomes with WENGAN
Efficient hybrid de novo assembly of human genomes with WENGAN
The genome assembler WENGAN produces high-quality human genome sequences at low computational cost.
- Alex Di Genova
- , Elena Buena-Atienza
- & Marie-France Sagot
-
Correspondence |
 Increased cyber-biosecurity for DNA synthesis
Increased cyber-biosecurity for DNA synthesis
- Rami Puzis
- , Dor Farbiash
- & Dov Greenbaum
-
Article |
 A long-term study of AAV gene therapy in dogs with hemophilia A identifies clonal expansions of transduced liver cells
A long-term study of AAV gene therapy in dogs with hemophilia A identifies clonal expansions of transduced liver cells
AAV therapy in dogs leads to clonal expansions of transduced cells.
- Giang N. Nguyen
- , John K. Everett
- & Denise E. Sabatino
-
Article |
 Determination of isoform-specific RNA structure with nanopore long reads
Determination of isoform-specific RNA structure with nanopore long reads
Using long reads to probe RNA structure reveals differences between transcript isoforms.
- Jong Ghut Ashley Aw
- , Shaun W. Lim
- & Yue Wan
-
Correspondence |
 Visualizing and interpreting cancer genomics data via the Xena platform
Visualizing and interpreting cancer genomics data via the Xena platform
- Mary J. Goldman
- , Brian Craft
- & David Haussler
-
Review Article |
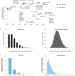 Design and analysis of CRISPR–Cas experiments
Design and analysis of CRISPR–Cas experiments
Hanna and Doench review the computational methods and tools that have become indispensable for planning and analyzing CRISPR experiments.
- Ruth E. Hanna
- & John G. Doench
-
Correspondence |
 MEMOTE for standardized genome-scale metabolic model testing
MEMOTE for standardized genome-scale metabolic model testing
- Christian Lieven
- , Moritz E. Beber
- & Cheng Zhang
-
Resource |
 A community effort to create standards for evaluating tumor subclonal reconstruction
A community effort to create standards for evaluating tumor subclonal reconstruction
Methods for reconstructing tumor evolution are benchmarked in the DREAM Somatic Mutation Calling Tumour Heterogeneity Challenge.
- Adriana Salcedo
- , Maxime Tarabichi
- & Paul C. Boutros
-
Correspondence |
 Reproducible, interactive, scalable and extensible microbiome data science using QIIME 2
Reproducible, interactive, scalable and extensible microbiome data science using QIIME 2
- Evan Bolyen
- , Jai Ram Rideout
- & J. Gregory Caporaso
-
Article |
 Taxonomic assignment of uncultivated prokaryotic virus genomes is enabled by gene-sharing networks
Taxonomic assignment of uncultivated prokaryotic virus genomes is enabled by gene-sharing networks
Classification of archaeal and bacterial viruses can be automated with an algorithm that identifies relationships on the basis of shared gene content.
- Ho Bin Jang
- , Benjamin Bolduc
- & Matthew B. Sullivan
-
Analysis |
 Efficient integration of heterogeneous single-cell transcriptomes using Scanorama
Efficient integration of heterogeneous single-cell transcriptomes using Scanorama
Scanorama integrates single-cell RNA-seq datasets from different tissues, different labs, different experiments or different technologies.
- Brian Hie
- , Bryan Bryson
- & Bonnie Berger
-
Article |
 A comparison of single-cell trajectory inference methods
A comparison of single-cell trajectory inference methods
The authors comprehensively benchmark the accuracy, scalability, stability and usability of 45 single-cell trajectory inference methods.
- Wouter Saelens
- , Robrecht Cannoodt
- & Yvan Saeys
-
Article |
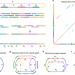 Assembly of long, error-prone reads using repeat graphs
Assembly of long, error-prone reads using repeat graphs
Flye improves the speed and accuracy of genome assembly by using repeat graphs to resolve repeat regions.
- Mikhail Kolmogorov
- , Jeffrey Yuan
- & Pavel A. Pevzner
-
Correspondence |
 The International Cancer Genome Consortium Data Portal
The International Cancer Genome Consortium Data Portal
- Junjun Zhang
- , Rosita Bajari
- & Vincent Ferretti

 Gene trajectory inference for single-cell data by optimal transport metrics
Gene trajectory inference for single-cell data by optimal transport metrics