Featured
-
-
Brief Communication
| Open Access Single-cell m6A mapping in vivo using picoMeRIP–seq
Single-cell m6A mapping in vivo using picoMeRIP–seq
A picogram-scale m6A mapping method is applied to single mouse oocytes and embryos in vivo.
- Yanjiao Li
- , Yunhao Wang
- & John Arne Dahl
-
Article
| Open Access scChIX-seq infers dynamic relationships between histone modifications in single cells
scChIX-seq infers dynamic relationships between histone modifications in single cells
Analysis of two histone marks in single cells reveals interplay between modifications.
- Jake Yeung
- , Maria Florescu
- & Alexander van Oudenaarden
-
Article
| Open Access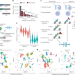 Multimodal chromatin profiling using nanobody-based single-cell CUT&Tag
Multimodal chromatin profiling using nanobody-based single-cell CUT&Tag
Three epigenetic features are mapped at single-cell resolution using Tn5-nanobody fusion proteins
- Marek Bartosovic
- & Gonçalo Castelo-Branco
-
Research Briefing |
 Profiling chromatin–protein crosstalk at locus resolution in single cells
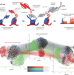
Profiling chromatin–protein crosstalk at locus resolution in single cells
Development is orchestrated in part by myriad chromatin-associated proteins that decorate the genome and physically coordinate in ways that are often unclear. An epigenomics technique called MulTI-Tag maps the genomic locations of different chromatin proteins in the same individual cells, helping to unveil their interactions.
-
Article
| Open Access Multifactorial profiling of epigenetic landscapes at single-cell resolution using MulTI-Tag
Multifactorial profiling of epigenetic landscapes at single-cell resolution using MulTI-Tag
A scalable barcoding method measures multiple chromatin-associated proteins in single cells.
- Michael P. Meers
- , Geneva Llagas
- & Steven Henikoff
-
Article |
 Comparison and imputation-aided integration of five commercial platforms for targeted DNA methylome analysis
Comparison and imputation-aided integration of five commercial platforms for targeted DNA methylome analysis
Differences in CpG coverage of targeted bisulfite sequencing methods can be overcome using imputation.
- Miljana Tanić
- , Ismail Moghul
- & Stephan Beck
-
Article |
 Chromatin Velocity reveals epigenetic dynamics by single-cell profiling of heterochromatin and euchromatin
Chromatin Velocity reveals epigenetic dynamics by single-cell profiling of heterochromatin and euchromatin
Single-cell mapping of heterochromatin and euchromatin using chromatin velocity defines trajectories of epigenetic modifications.
- Martina Tedesco
- , Francesca Giannese
- & Giovanni Tonon
-
Article |
 Measurement of histone replacement dynamics with genetically encoded exchange timers in yeast
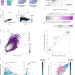
Measurement of histone replacement dynamics with genetically encoded exchange timers in yeast
The dynamics of histone exchange are measured with genetically encoded timers.
- Gilad Yaakov
- , Felix Jonas
- & Naama Barkai
-
Article |
 Extended-representation bisulfite sequencing of gene regulatory elements in multiplexed samples and single cells
Extended-representation bisulfite sequencing of gene regulatory elements in multiplexed samples and single cells
DNA methylation is efficiently profiled in non-coding regulatory sequences in single cells.
- Sarah J. Shareef
- , Samantha M. Bevill
- & Bradley E. Bernstein
-
Article |
 Profiling the genetic determinants of chromatin accessibility with scalable single-cell CRISPR screens
Profiling the genetic determinants of chromatin accessibility with scalable single-cell CRISPR screens
The effects of gene knockouts on chromatin accessibility are measured with single-cell CRISPR screens.
- Noa Liscovitch-Brauer
- , Antonino Montalbano
- & Neville E. Sanjana
-
Article |
 Single-cell CUT&Tag profiles histone modifications and transcription factors in complex tissues
Single-cell CUT&Tag profiles histone modifications and transcription factors in complex tissues
An improved method for single-cell analysis of histone modifications is applied to the mouse brain.
- Marek Bartosovic
- , Mukund Kabbe
- & Gonçalo Castelo-Branco
-
Letter |
 Single-cell CUT&Tag analysis of chromatin modifications in differentiation and tumor progression
Single-cell CUT&Tag analysis of chromatin modifications in differentiation and tumor progression
An improved method for single-cell analysis of histone modifications is applied to stem cell differentiation and cancer.
- Steven J. Wu
- , Scott N. Furlan
- & Anoop P. Patel
-
Article |
 GPSeq reveals the radial organization of chromatin in the cell nucleus
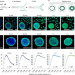
GPSeq reveals the radial organization of chromatin in the cell nucleus
The location of genetic and epigenetic elements in mammalian nuclei is measured by gradual DNA fragmentation.
- Gabriele Girelli
- , Joaquin Custodio
- & Magda Bienko
-
Letter |
 Bisulfite-free direct detection of 5-methylcytosine and 5-hydroxymethylcytosine at base resolution
Bisulfite-free direct detection of 5-methylcytosine and 5-hydroxymethylcytosine at base resolution
A new method using mild chemical reactions and enzymatic oxidation allows nondestructive sequencing of 5-methylcytosine and 5-hydroxymethylcytosine with base-level resolution.
- Yibin Liu
- , Paulina Siejka-Zielińska
- & Chun-Xiao Song
-
Letter |
 Targeted DNA demethylation in vivo using dCas9–peptide repeat and scFv–TET1 catalytic domain fusions
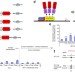
Targeted DNA demethylation in vivo using dCas9–peptide repeat and scFv–TET1 catalytic domain fusions
DNA methyl groups are selectively removed at target loci using inactive Cas9 fused to a SunTag-based peptide repeat that recruits the enzymatic activity.
- Sumiyo Morita
- , Hirofumi Noguchi
- & Izuho Hatada
-
Analysis
| Open Access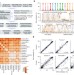 Quantitative comparison of DNA methylation assays for biomarker development and clinical applications
Quantitative comparison of DNA methylation assays for biomarker development and clinical applications
A multicenter comparison of DNA methylation assays identifies robust methods for translational research and clinical diagnostics.
- Christoph Bock
- , Florian Halbritter
- & Kun Zhang
-
Analysis |
 Reduced local mutation density in regulatory DNA of cancer genomes is linked to DNA repair
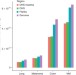
Reduced local mutation density in regulatory DNA of cancer genomes is linked to DNA repair
Polak et al. analyze somatic mutations and chromatin accessibility data to gain insight into mutational processes in cancer genomes.
- Paz Polak
- , Michael S Lawrence
- & Shamil R Sunyaev
-
Letter |
 Locus-specific editing of histone modifications at endogenous enhancers
Locus-specific editing of histone modifications at endogenous enhancers
The function of specific enhancers is studied using TAL effectors fused to a histone demethylase.
- Eric M Mendenhall
- , Kaylyn E Williamson
- & Bradley E Bernstein
-
Article |
 Genome-wide mapping of methylated adenine residues in pathogenic Escherichia coli using single-molecule real-time sequencing
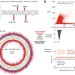
Genome-wide mapping of methylated adenine residues in pathogenic Escherichia coli using single-molecule real-time sequencing
Fang et al. generate the first genome-wide, strand-specific, single base–pair resolution map of m6A modifications in Escherichia coli.
- Gang Fang
- , Diana Munera
- & Eric E Schadt
-
News & Views |
 New lysine methyltransferase drug targets in cancer
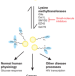
New lysine methyltransferase drug targets in cancer
Recent preclinical studies suggest that inhibitors of histone methyltransferases represent promising drug candidates for cancer therapy.
- Tobias Wagner
- & Manfred Jung
-
-
Letter |
 Selective chemical labeling reveals the genome-wide distribution of 5-hydroxymethylcytosine
Selective chemical labeling reveals the genome-wide distribution of 5-hydroxymethylcytosine
Song et al. present the first method for global analysis of 5-hydroxymethylcytosine, a recently identified epigenetic modification in mammalian cells. They use a bacteriophage-derived enzyme to tag the hydroxymethyl group with an azide-modified glucose residue that can be used for affinity purification and sequencing of modified genomic DNA fragments.
- Chun-Xiao Song
- , Keith E Szulwach
- & Chuan He
-
News & Views |
Taking the measure of the methylome
Two comparative studies from the International Human Epigenome Project find high concordance between different methods for measuring genomic methylation.
- Stephan Beck
-
-
Analysis |
 Comparison of sequencing-based methods to profile DNA methylation and identification of monoallelic epigenetic modifications
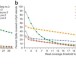
Comparison of sequencing-based methods to profile DNA methylation and identification of monoallelic epigenetic modifications
Methods for profiling DNA methylation differ in the physical principles used to detect modified cytosines. Harris et al. compare the performances of four sequencing-based technologies for genome-wide analysis of DNA methylation and combine two methods to enable detection of allelic differences in epigenetic marks.
- R Alan Harris
- , Ting Wang
- & Joseph F Costello
-
Analysis |
 Quantitative comparison of genome-wide DNA methylation mapping technologies
Quantitative comparison of genome-wide DNA methylation mapping technologies
Comparison of the methylation patterns of cells in different developmental or disease states can help to elucidate both normal and pathological regulatory mechanisms. Bock et al. evaluate the ability of three sequencing-based methods and one microarray-based technology to detect differentially methylated regions on a genome-wide scale.
- Christoph Bock
- , Eleni M Tomazou
- & Alexander Meissner
-
Analysis |
 Discovery and characterization of chromatin states for systematic annotation of the human genome
Discovery and characterization of chromatin states for systematic annotation of the human genome
Which of the possible combinations of epigenetic marks have biological significance is a major question in epigenetics. Analyzing data from human T-cells, Ernst and Kellis discover 51 distinct, recurring combinations of histone modifications that can be correlated with the functional annotations of the underlying DNA sequences.
- Jason Ernst
- & Manolis Kellis
-

 Joint single-cell profiling resolves 5mC and 5hmC and reveals their distinct gene regulatory effects
Joint single-cell profiling resolves 5mC and 5hmC and reveals their distinct gene regulatory effects Epigenetic modifications and human disease
Epigenetic modifications and human disease Detecting methylated bases in real time
Detecting methylated bases in real time