Featured
-
-
Article
| Open Access Mapping medically relevant RNA isoform diversity in the aged human frontal cortex with deep long-read RNA-seq
Mapping medically relevant RNA isoform diversity in the aged human frontal cortex with deep long-read RNA-seq
The landscape of RNA isoforms in human cortex is revealed by deep long-read RNA sequencing.
- Bernardo Aguzzoli Heberle
- , J. Anthony Brandon
- & Mark T. W. Ebbert
-
Article |
 Development of multiplexed orthogonal base editor (MOBE) systems
Development of multiplexed orthogonal base editor (MOBE) systems
Multiplexed orthogonal base editing systems introduce different point mutations at two loci simultaneously.
- Quinn T. Cowan
- , Sifeng Gu
- & Alexis C. Komor
-
Research Briefing |
 Advancing programmable gene expression in plants using CRISPRi-based Boolean gates
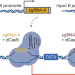
Advancing programmable gene expression in plants using CRISPRi-based Boolean gates
To advance the toolset for controlling plant gene expression, we developed a CRISPR interference-based platform for the construction of synthetic Boolean logic gates that is functional in multiple plant species. These genetic circuits are programmable and reversible in nature, which will enable spatiotemporal control of plant responses to dynamic cues.
-
Article |
 CRISPRi-based circuits to control gene expression in plants
CRISPRi-based circuits to control gene expression in plants
Programmable and reversible CRISPRi-based genetic circuits function in a variety of plants.
- Muhammad Adil Khan
- , Gabrielle Herring
- & Ryan Lister
-
News & Views |
 Decoding gene regulation with CRISPR perturbations
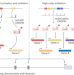
Decoding gene regulation with CRISPR perturbations
Two CRISPR tools for combinatorial genetic perturbations reveal gene regulatory networks.
- Stefan Oberlin
- & Michael T. McManus
-
Article
| Open Access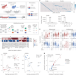 Bidirectional epigenetic editing reveals hierarchies in gene regulation
Bidirectional epigenetic editing reveals hierarchies in gene regulation
CRISPRai simultaneously activates and represses two genes in single cells.
- Naomi M. Pacalin
- , Zachary Steinhart
- & Howard Y. Chang
-
Article
| Open Access Engineered CRISPR-Cas12a for higher-order combinatorial chromatin perturbations
Engineered CRISPR-Cas12a for higher-order combinatorial chromatin perturbations
An engineered Cas12a enables higher-order combinatorial functional genomic screens using CRISPR interference.
- C. C.-S. Hsiung
- , C. M. Wilson
- & L. A. Gilbert
-
News & Views |
 A milestone method to make natural killer T cells
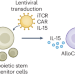
A milestone method to make natural killer T cells
A differentiation protocol to produce off-the-shelf natural killer T cells may enable clinical application.
- Leonid S. Metelitsa
-
Article |
 Protein-adaptive differential scanning fluorimetry using conformationally responsive dyes
Protein-adaptive differential scanning fluorimetry using conformationally responsive dyes
A library of fluorogenic dyes enables the measurement of protein unfolding for a large range of proteins.
- Taiasean Wu
- , Joshua C. Yu
- & Jason E. Gestwicki
-
Article
| Open Access Generation of allogeneic CAR-NKT cells from hematopoietic stem and progenitor cells using a clinically guided culture method
Generation of allogeneic CAR-NKT cells from hematopoietic stem and progenitor cells using a clinically guided culture method
Off-the-shelf CAR-NKT cells for cancer immunotherapy are advanced toward clinical translation.
- Yan-Ruide Li
- , Yang Zhou
- & Lili Yang
-
Article
| Open Access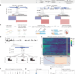 Linking CRISPR–Cas9 double-strand break profiles to gene editing precision with BreakTag
Linking CRISPR–Cas9 double-strand break profiles to gene editing precision with BreakTag
The genome-wide landscape of Cas-induced double-strand breaks and end structures is profiled at nucleotide resolution.
- Gabriel M. C. Longo
- , Sergi Sayols
- & Vassilis Roukos
-
Research Briefing |
 Decoding cell replicational age from single-cell ATAC-seq data
Decoding cell replicational age from single-cell ATAC-seq data
The replicational age of single cells provides a temporal reference for tracking cell fate transition trajectories. The computational framework EpiTrace measures cell age using single-cell ATAC-seq data, specifically by considering chromatin accessibility at clock-like genomic loci, enabling the reconstruction of the history of developmental and pathological processes.
-
Article
| Open Access Tracking single-cell evolution using clock-like chromatin accessibility loci
Tracking single-cell evolution using clock-like chromatin accessibility loci
EpiTrace infers cell mitotic age and evolution from single-cell ATAC-seq data.
- Yu Xiao
- , Wan Jin
- & Yi Zhang
-
Brief Communication
| Open Access Identification of clinically relevant T cell receptors for personalized T cell therapy using combinatorial algorithms
Identification of clinically relevant T cell receptors for personalized T cell therapy using combinatorial algorithms
Tumor-reactive T cell receptors are predicted based on single-cell RNA sequencing and single-cell T cell receptor sequencing.
- Rémy Pétremand
- , Johanna Chiffelle
- & Alexandre Harari
-
Article
| Open Access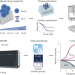 Decrypting the molecular basis of cellular drug phenotypes by dose-resolved expression proteomics
Decrypting the molecular basis of cellular drug phenotypes by dose-resolved expression proteomics
Proteome-wide measurements of dose–response curves for 144 drugs reveal drug mechanisms of action.
- Stephan Eckert
- , Nicola Berner
- & Bernhard Kuster
-
-
Correspondence |
Response to “The perpetual motion machine of AI-generated data and the distraction of ChatGPT as a ‘scientist’”
- William Stafford Noble
-
News & Views |
 Designing drugs with reversible activity
Designing drugs with reversible activity
A strategy for creating drugs that can be quickly neutralized is demonstrated for anticoagulants.
- Dario Neri
-
Article
| Open Access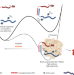 Development of supramolecular anticoagulants with on-demand reversibility
Development of supramolecular anticoagulants with on-demand reversibility
A supramolecular drug design yields anticoagulants that are rapidly reversible.
- Millicent Dockerill
- , Daniel J. Ford
- & Nicolas Winssinger
-
Article |
 Analysis and benchmarking of small and large genomic variants across tandem repeats
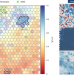
Analysis and benchmarking of small and large genomic variants across tandem repeats
A truth-set catalog of human tandem repeats allows benchmarking of variant analysis tools.
- Adam C. English
- , Egor Dolzhenko
- & Fritz J. Sedlazeck
-
Research Briefing |
 Assessing the laboratory performance of AI-generated enzymes
Assessing the laboratory performance of AI-generated enzymes
A set of 20 computational metrics was evaluated to determine whether they could predict the functionality of synthetic enzyme sequences produced by generative protein models, resulting in the development of a computational filter, COMPSS, that increased experimental success rates by 50–150%, tested in over 500 natural and AI-generated enzymes.
-
Article |
 Discovery of tumor-reactive T cell receptors by massively parallel library synthesis and screening
Discovery of tumor-reactive T cell receptors by massively parallel library synthesis and screening
Tumor-specific T cell receptors (TCRs) are identified by functional screening of synthetic TCR libraries.
- Ziva Moravec
- , Yue Zhao
- & Wouter Scheper
-
Article
| Open Access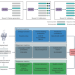 Computational scoring and experimental evaluation of enzymes generated by neural networks
Computational scoring and experimental evaluation of enzymes generated by neural networks
Metrics that predict the success of folding and the activity of designed protein sequences are developed and experimentally validated.
- Sean R. Johnson
- , Xiaozhi Fu
- & Kevin K. Yang
-
Article |
 Therapeutic application of circular RNA aptamers in a mouse model of psoriasis
Therapeutic application of circular RNA aptamers in a mouse model of psoriasis
A circular RNA aptamer shows therapeutic efficacy in vivo in a psoriasis model.
- Si-Kun Guo
- , Chu-Xiao Liu
- & Ling-Ling Chen
-
-
Research Briefing |
 Video game unleashes millions of citizen scientists on microbiome research
Video game unleashes millions of citizen scientists on microbiome research
Borderlands Science is a casual mini-game released within a mass-market video game that crowdsources the alignment of one million RNA sequences from the human microbiome. In 3 years, 4 million participants generated over 135 million puzzle solutions that were used to build a reference alignment and improve microbial phylogeny.
-
Patents |
 Mapping the global landscape for induced pluripotent stem cells from patents and clinical trials
Mapping the global landscape for induced pluripotent stem cells from patents and clinical trials
Although the clinical application of induced pluripotent stem cells is still in its infancy, the development of iPSC technologies reflected by an increasing number of patents provides hope that they will realize their therapeutic potential.
- Liyang Lyu
- , Ye Feng
- & Yuanjia Hu
-
Article
| Open Access Improving microbial phylogeny with citizen science within a mass-market video game
Improving microbial phylogeny with citizen science within a mass-market video game
Gamification of the multiple sequence alignment problem improves microbial phylogeny estimates.
- Roman Sarrazin-Gendron
- , Parham Ghasemloo Gheidari
- & Jérôme Waldispühl
-
Article
| Open Access Inferring gene regulatory networks from single-cell multiome data using atlas-scale external data
Inferring gene regulatory networks from single-cell multiome data using atlas-scale external data
Accuracy of gene regulatory network inference is increased by combining multiome single-cell and atlas-scale bulk data.
- Qiuyue Yuan
- & Zhana Duren
-
Article |
 Genome engineering with Cas9 and AAV repair templates generates frequent concatemeric insertions of viral vectors
Genome engineering with Cas9 and AAV repair templates generates frequent concatemeric insertions of viral vectors
AAV vectors form difficult-to-detect concatemers at Cas9 target sites.
- Fabian P. Suchy
- , Daiki Karigane
- & Hiromitsu Nakauchi
-
Article |
 Gene trajectory inference for single-cell data by optimal transport metrics
Gene trajectory inference for single-cell data by optimal transport metrics
Gene dynamics of concurrent biological processes are unraveled by considering gene trajectories instead of cell trajectories.
- Rihao Qu
- , Xiuyuan Cheng
- & Yuval Kluger
-
Research Briefing |
 Capturing and modeling cellular niches from dissociated single-cell and spatial data
Capturing and modeling cellular niches from dissociated single-cell and spatial data
Cells interact with their local environment to enact global tissue function. By harnessing gene–gene covariation in cellular neighborhoods from spatial transcriptomics data, the covariance environment (COVET) niche representation and the environmental variational inference (ENVI) data integration method model phenotype–microenvironment interplay and reconstruct the spatial context of dissociated single-cell RNA sequencing datasets.
-
Research Briefing |
 Genetically encoding colors and images into bioengineered microbial materials
Genetically encoding colors and images into bioengineered microbial materials
Using synthetic biology, we engineered a cellulose-producing bacterium that can produce eumelanin and respond to light, so that it is possible to grow a microbial leather material that is colored black or contains projected black patterns.
-
Article
| Open Access Self-pigmenting textiles grown from cellulose-producing bacteria with engineered tyrosinase expression
Self-pigmenting textiles grown from cellulose-producing bacteria with engineered tyrosinase expression
Genetic engineering of material-producing bacteria creates pigmented products.
- Kenneth T. Walker
- , Ivy S. Li
- & Tom Ellis
-
Article
| Open Access The covariance environment defines cellular niches for spatial inference
The covariance environment defines cellular niches for spatial inference
Using gene–gene covariance structure to represent cellular neighborhoods improves the integration of single-cell RNA sequencing and spatial transcriptomics data.
- Doron Haviv
- , Ján Remšík
- & Dana Pe’er
-
Brief Communication
| Open Access De novo and somatic structural variant discovery with SVision-pro
De novo and somatic structural variant discovery with SVision-pro
SVision-pro enables pairwise genome comparison and improves de novo and somatic structural variant calling.
- Songbo Wang
- , Jiadong Lin
- & Kai Ye
-
Article |
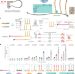 Branched chemically modified poly(A) tails enhance the translation capacity of mRNA
Branched chemically modified poly(A) tails enhance the translation capacity of mRNA
mRNA with engineered poly(A) tails produces prolonged higher levels of protein.
- Hongyu Chen
- , Dangliang Liu
- & Xiao Wang
-
Article
| Open Access Starfysh integrates spatial transcriptomic and histologic data to reveal heterogeneous tumor–immune hubs
Starfysh integrates spatial transcriptomic and histologic data to reveal heterogeneous tumor–immune hubs
Reference-free integrative analysis of spatial transcriptomics data is achieved with archetype analysis.
- Siyu He
- , Yinuo Jin
- & Elham Azizi
-
Correspondence |
 Making genome editing a success story in Africa
Making genome editing a success story in Africa
- Hussein M. Abkallo
- , Patrick Arbuthnot
- & Charles Wondji
-
News & Views |
 A retrotransposon for site-specific gene transfer
A retrotransposon for site-specific gene transfer
An engineered retrotransposon achieves targeted gene transfer into the human genome.
- Fred Dyda
- & Alison B. Hickman
-
Research Briefing |
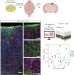 Glia-enriched cortical organoids implanted in mice capture astrocyte diversity
Glia-enriched cortical organoids implanted in mice capture astrocyte diversity
The underrepresentation of functional glial cells is a major challenge in brain organoid models. We developed an astroglia-enriched cortical organoid model that allows efficient generation of functional astrocytes and enables the formation of astroglial morphological subclasses with layer-specific gene expression profiles upon transplantation into the mouse brain.
-
Article
| Open Access High-throughput evaluation of genetic variants with prime editing sensor libraries
High-throughput evaluation of genetic variants with prime editing sensor libraries
Prime editing sensor libraries evaluate diverse genetic variants in their endogenous genomic contexts.
- Samuel I. Gould
- , Alexandra N. Wuest
- & Francisco J. Sánchez Rivera
-
Article
| Open Access A small-molecule TNIK inhibitor targets fibrosis in preclinical and clinical models
A small-molecule TNIK inhibitor targets fibrosis in preclinical and clinical models
An AI-generated small-molecule inhibitor treats fibrosis in vivo and in phase I clinical trials.
- Feng Ren
- , Alex Aliper
- & Alex Zhavoronkov
-
Article
| Open Access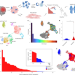 Prediction of tumor-reactive T cell receptors from scRNA-seq data for personalized T cell therapy
Prediction of tumor-reactive T cell receptors from scRNA-seq data for personalized T cell therapy
Tumor-reactive T cell receptors are rapidly identified from lymphocyte scRNA-seq data.
- C. L. Tan
- , K. Lindner
- & E. W. Green
-
Article |
 Near-cognate tRNAs increase the efficiency and precision of pseudouridine-mediated readthrough of premature termination codons
Near-cognate tRNAs increase the efficiency and precision of pseudouridine-mediated readthrough of premature termination codons
Disease-related premature stop codons are efficiently corrected by harnessing near-cognate tRNAs.
- Nan Luo
- , Qiang Huang
- & Chengqi Yi
-
Research Briefing |
 Imbalanced single-cell data integration leads to loss of biological information
Imbalanced single-cell data integration leads to loss of biological information
The Iniquitate pipeline assessed the impacts of cell-type imbalance on single-cell RNA sequencing integration through perturbations to dataset balance. The results indicated that cell-type imbalance not only leads to loss of biological signal in the integrated space, but also can change the interpretation of downstream analyses after integration.
-
Article |
 Characterizing the impacts of dataset imbalance on single-cell data integration
Characterizing the impacts of dataset imbalance on single-cell data integration
Iniquitate quantifies the effects of cell type imbalance on single-cell RNA sequencing data integration.
- Hassaan Maan
- , Lin Zhang
- & Bo Wang
-
Article |
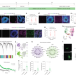 Morphological diversification and functional maturation of human astrocytes in glia-enriched cortical organoid transplanted in mouse brain
Morphological diversification and functional maturation of human astrocytes in glia-enriched cortical organoid transplanted in mouse brain
Transplantation of glia-enriched organoids into the mouse brain enables the study of astrocyte subtypes.
- Meiyan Wang
- , Lei Zhang
- & Fred H. Gage
-
Perspective |
 RNA interference in the era of nucleic acid therapeutics
RNA interference in the era of nucleic acid therapeutics
With six approved drugs, siRNA is now an established therapeutic modality poised for expansion.
- Vasant Jadhav
- , Akshay Vaishnaw
- & Martin A. Maier
Browse narrower subjects
- Biochemistry
- Biological techniques
- Biophysics
- Biotechnology
- Cancer
- Cell biology
- Chemical biology
- Computational biology and bioinformatics
- Developmental biology
- Drug discovery
- Ecology
- Evolution
- Genetics
- Immunology
- Microbiology
- Molecular biology
- Neuroscience
- Physiology
- Plant sciences
- Psychology
- Stem cells
- Structural biology
- Systems biology
- Zoology

 Mapping variant effects on anti-tumor hallmarks of primary human T cells with base-editing screens
Mapping variant effects on anti-tumor hallmarks of primary human T cells with base-editing screens