News & Views |
Featured
-
-
Article
| Open Access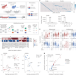 Bidirectional epigenetic editing reveals hierarchies in gene regulation
Bidirectional epigenetic editing reveals hierarchies in gene regulation
CRISPRai simultaneously activates and represses two genes in single cells.
- Naomi M. Pacalin
- , Zachary Steinhart
- & Howard Y. Chang
-
Article |
 Protein-adaptive differential scanning fluorimetry using conformationally responsive dyes
Protein-adaptive differential scanning fluorimetry using conformationally responsive dyes
A library of fluorogenic dyes enables the measurement of protein unfolding for a large range of proteins.
- Taiasean Wu
- , Joshua C. Yu
- & Jason E. Gestwicki
-
Article |
 Discovery of tumor-reactive T cell receptors by massively parallel library synthesis and screening
Discovery of tumor-reactive T cell receptors by massively parallel library synthesis and screening
Tumor-specific T cell receptors (TCRs) are identified by functional screening of synthetic TCR libraries.
- Ziva Moravec
- , Yue Zhao
- & Wouter Scheper
-
Research Briefing |
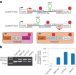 Generation of programmable splicing factors using RNA-binding proteins that activate exon inclusion
Generation of programmable splicing factors using RNA-binding proteins that activate exon inclusion
We developed luciferase-based splicing reporters to evaluate 718 RNA-binding proteins (RBPs) for the ability to activate exon inclusion. Our screen detected RBPs with no previously characterized roles in splicing and utilized RBP domains displaying potent exon activation to develop programmable splicing modulators with improved strength, generalizability and size over previous versions.
-
Article
| Open Access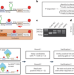 Large-scale evaluation of the ability of RNA-binding proteins to activate exon inclusion
Large-scale evaluation of the ability of RNA-binding proteins to activate exon inclusion
Alternative splicing is controlled with engineered fusion splicing factors.
- Jonathan C. Schmok
- , Manya Jain
- & Gene W. Yeo
-
Research Briefing |
 Compressed Perturb-seq enables highly efficient genetic screens
Compressed Perturb-seq enables highly efficient genetic screens
Using an experimental and computational framework inspired by compressed sensing, we greatly reduced the number of measurements needed to run Perturb-seq. Our compressed Perturb-seq strategy relies on collecting measurements comprising random linear combinations of genetic perturbations, followed by deconvolving the perturbation effects on the transcriptome using sparsity-exploiting algorithms.
-
Article
| Open Access Scalable genetic screening for regulatory circuits using compressed Perturb-seq
Scalable genetic screening for regulatory circuits using compressed Perturb-seq
Compressed Perturb-seq incorporates compressed sensing to genetic screening for scalable discovery of genetic interactions.
- Douglas Yao
- , Loic Binan
- & Brian Cleary
-
Article |
 Prediction of on-target and off-target activity of CRISPR–Cas13d guide RNAs using deep learning
Prediction of on-target and off-target activity of CRISPR–Cas13d guide RNAs using deep learning
A machine learning model predicts on-target and off-target activity of Cas13d in human cells.
- Hans-Hermann Wessels
- , Andrew Stirn
- & Neville E. Sanjana
-
Brief Communication
| Open Access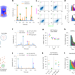 High-throughput measurement of the content and properties of nano-sized bioparticles with single-particle profiler
High-throughput measurement of the content and properties of nano-sized bioparticles with single-particle profiler
Lipid nanoparticles, viruses, exosomes and liposomes are characterized by analysis of fluorescence fluctuations.
- Taras Sych
- , Jan Schlegel
- & Erdinc Sezgin
-
Article |
 RIP-PEN-seq identifies a class of kink-turn RNAs as splicing regulators
RIP-PEN-seq identifies a class of kink-turn RNAs as splicing regulators
Immunoprecipitation coupled with paired-end sequencing of non-coding RNAs identifies a class of splicing regulators.
- Bin Li
- , Shurong Liu
- & Jianhua Yang
-
Article
| Open Access Prediction of prime editing insertion efficiencies using sequence features and DNA repair determinants
Prediction of prime editing insertion efficiencies using sequence features and DNA repair determinants
Prime editing efficiency is affected by insertion sequence features, secondary structure and DNA repair context.
- Jonas Koeppel
- , Juliane Weller
- & Leopold Parts
-
Article |
 Massively parallel knock-in engineering of human T cells
Massively parallel knock-in engineering of human T cells
Unbiased selection of CAR-T cells is achieved by massively parallel knock-in engineering.
- Xiaoyun Dai
- , Jonathan J. Park
- & Sidi Chen
-
Article |
 Predicting prime editing efficiency and product purity by deep learning
Predicting prime editing efficiency and product purity by deep learning
The design of prime editing guide RNAs is optimized by deep learning.
- Nicolas Mathis
- , Ahmed Allam
- & Gerald Schwank
-
Article
| Open Access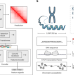 Cell-type-specific prediction of 3D chromatin organization enables high-throughput in silico genetic screening
Cell-type-specific prediction of 3D chromatin organization enables high-throughput in silico genetic screening
Deep learning predicts cell-type-specific 3D chromatin contacts from DNA sequence, chromatin accessibility and CTCF binding.
- Jimin Tan
- , Nina Shenker-Tauris
- & Aristotelis Tsirigos
-
Article |
 A proteome-wide atlas of drug mechanism of action
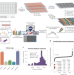
A proteome-wide atlas of drug mechanism of action
A proteome-wide atlas of the effect of 875 compounds reveals mechanisms of action and off-target effects.
- Dylan C. Mitchell
- , Miljan Kuljanin
- & Steven P. Gygi
-
Article
| Open Access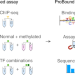 Prediction of protein–ligand binding affinity from sequencing data with interpretable machine learning
Prediction of protein–ligand binding affinity from sequencing data with interpretable machine learning
Protein–ligand binding affinity is predicted quantitatively from sequencing data.
- H. Tomas Rube
- , Chaitanya Rastogi
- & Harmen J. Bussemaker
-
Article |
 Efficient discovery of SARS-CoV-2-neutralizing antibodies via B cell receptor sequencing and ligand blocking
Efficient discovery of SARS-CoV-2-neutralizing antibodies via B cell receptor sequencing and ligand blocking
B cell receptor sequencing with ligand blocking speeds up neutralizing antibody discovery.
- Andrea R. Shiakolas
- , Kevin J. Kramer
- & Ivelin S. Georgiev
-
Article |
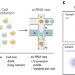 Massively parallel phenotyping of coding variants in cancer with Perturb-seq
Massively parallel phenotyping of coding variants in cancer with Perturb-seq
The functional impact of somatic mutations in cancer genes is determined by pooled Perturb-seq.
- Oana Ursu
- , James T. Neal
- & Jesse S. Boehm
-
Article |
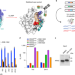 Conformation-locking antibodies for the discovery and characterization of KRAS inhibitors
Conformation-locking antibodies for the discovery and characterization of KRAS inhibitors
Antibodies that lock KRAS mutants in inactive conformations facilitate drug discovery.
- Christopher W. Davies
- , Angela J. Oh
- & Marie Evangelista
-
News & Views |
 Improving CRISPR tools by elucidating DNA repair
Improving CRISPR tools by elucidating DNA repair
Genome-editing tools are optimized by identifying genes that affect DNA repair.
- Jung Min Lim
- & Hyongbum Henry Kim
-
Article |
 Functional single-cell genomics of human cytomegalovirus infection
Functional single-cell genomics of human cytomegalovirus infection
Pooled and single-cell CRISPR screens provide a detailed map of human cytomegalovirus infection.
- Marco Y. Hein
- & Jonathan S. Weissman
-
Article |
 Genome-wide interrogation of gene functions through base editor screens empowered by barcoded sgRNAs
Genome-wide interrogation of gene functions through base editor screens empowered by barcoded sgRNAs
CRISPR genetic screens with base editors avoid the confounding effects of DNA breaks.
- Ping Xu
- , Zhiheng Liu
- & Wensheng Wei
-
Article |
 Profiling the genetic determinants of chromatin accessibility with scalable single-cell CRISPR screens
Profiling the genetic determinants of chromatin accessibility with scalable single-cell CRISPR screens
The effects of gene knockouts on chromatin accessibility are measured with single-cell CRISPR screens.
- Noa Liscovitch-Brauer
- , Antonino Montalbano
- & Neville E. Sanjana
-
Article |
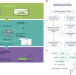 Biological activity-based modeling identifies antiviral leads against SARS-CoV-2
Biological activity-based modeling identifies antiviral leads against SARS-CoV-2
Activity profiles generated from quantitative high-throughput screening improve drug candidate prediction.
- Ruili Huang
- , Miao Xu
- & Wei Zheng
-
Article |
 Optimization of AsCas12a for combinatorial genetic screens in human cells
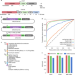
Optimization of AsCas12a for combinatorial genetic screens in human cells
Improved Cas12a variants and sgRNA design rules enhance genome-wide screens.
- Peter C. DeWeirdt
- , Kendall R. Sanson
- & John G. Doench
-
Letter |
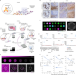 Multiplex digital spatial profiling of proteins and RNA in fixed tissue
Multiplex digital spatial profiling of proteins and RNA in fixed tissue
A turnkey system allows for spatial profiling of proteins and RNA in fixed tissues, providing a window on cellular heterogeneity.
- Christopher R. Merritt
- , Giang T. Ong
- & Joseph M. Beechem
-
Article |
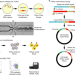 Massively parallel interrogation and mining of natively paired human TCRαβ repertoires
Massively parallel interrogation and mining of natively paired human TCRαβ repertoires
T cell receptors are identified in large-scale screening of T cell repertoires cloned from primary human T cells.
- Matthew J. Spindler
- , Ayla L. Nelson
- & David S. Johnson
-
Letter |
 Massively parallel Cas13 screens reveal principles for guide RNA design
Massively parallel Cas13 screens reveal principles for guide RNA design
Design of optimal guide RNAs for Cas13d-mediated RNA knockdown is enabled by computational modeling of large-scale screening data.
- Hans-Hermann Wessels
- , Alejandro Méndez-Mancilla
- & Neville E. Sanjana
-
Matters Arising |
Reply to: Fitness effects of CRISPR/Cas9-targeting of long noncoding RNA genes
- Ying Liu
- , Zhiheng Liu
- & Wensheng Wei
-
Article |
 Titrating gene expression using libraries of systematically attenuated CRISPR guide RNAs
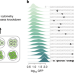
Titrating gene expression using libraries of systematically attenuated CRISPR guide RNAs
Gene expression is precisely modulated with CRISPR interference and mismatched guide RNAs.
- Marco Jost
- , Daniel A. Santos
- & Jonathan S. Weissman
-
Article |
 Immuno-SABER enables highly multiplexed and amplified protein imaging in tissues
Immuno-SABER enables highly multiplexed and amplified protein imaging in tissues
A DNA-based amplification scheme enables highly multiplexed immunofluorescence imaging
- Sinem K. Saka
- , Yu Wang
- & Peng Yin
-
Article |
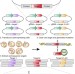 Predicting the mutations generated by repair of Cas9-induced double-strand breaks
Predicting the mutations generated by repair of Cas9-induced double-strand breaks
Analysis of more than a billion repair outcomes yields an algorithm that predicts Cas9-induced mutations.
- Felicity Allen
- , Luca Crepaldi
- & Leopold Parts
-
Letter |
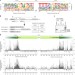 Genome-wide screening for functional long noncoding RNAs in human cells by Cas9 targeting of splice sites
Genome-wide screening for functional long noncoding RNAs in human cells by Cas9 targeting of splice sites
Essential long noncoding RNAs are identified using Cas9 targeting of splice sites.
- Ying Liu
- , Zhongzheng Cao
- & Wensheng Wei
-
Resource |
 Evaluation of 244,000 synthetic sequences reveals design principles to optimize translation in Escherichia coli
Evaluation of 244,000 synthetic sequences reveals design principles to optimize translation in Escherichia coli
Large and systematic mRNA library design disentangles the complex sequence determinants of translation efficiency in bacteria.
- Guillaume Cambray
- , Joao C Guimaraes
- & Adam Paul Arkin
-
Letter |
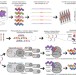 Encoding human serine phosphopeptides in bacteria for proteome-wide identification of phosphorylation-dependent interactions
Encoding human serine phosphopeptides in bacteria for proteome-wide identification of phosphorylation-dependent interactions
Screening with bacteria encoding human serine phosphopeptides reveals phosphoprotein interactions.
- Karl W Barber
- , Paul Muir
- & Jesse Rinehart
-
Correspondence |
The international MAQC Society launches to enhance reproducibility of high-throughput technologies
- Leming Shi
- , Rebecca Kusko
- & Weida Tong
-
Correspondence |
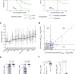 Alternative drug sensitivity metrics improve preclinical cancer pharmacogenomics
Alternative drug sensitivity metrics improve preclinical cancer pharmacogenomics
- Marc Hafner
- , Mario Niepel
- & Peter K Sorger
-
News & Views |
 Single-minded CRISPR screening
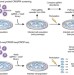
Single-minded CRISPR screening
Transcriptome-wide effects of gene knockouts are screened at the single-cell level by combining high-throughput, single-cell RNA-seq with lentiviral CRISPR libraries.
- Bryan R Lanning
- & Christopher R Vakoc
-
Article |
 Genome-scale deletion screening of human long non-coding RNAs using a paired-guide RNA CRISPR–Cas9 library
Genome-scale deletion screening of human long non-coding RNAs using a paired-guide RNA CRISPR–Cas9 library
Long non-coding RNAs are identified using a high-throughput paired-guide RNA genomic deletion screen.
- Shiyou Zhu
- , Wei Li
- & Wensheng Wei
-
Article |
 Large-scale detection of antigen-specific T cells using peptide-MHC-I multimers labeled with DNA barcodes
Large-scale detection of antigen-specific T cells using peptide-MHC-I multimers labeled with DNA barcodes
Detection of more than 1,000 different T-cell specificities in parallel facilitates identification of cancer and autoimmune antigens.
- Amalie Kai Bentzen
- , Andrea Marion Marquard
- & Sine Reker Hadrup
-
News & Views |
 Deep phenotyping predicts Huntington's genotype
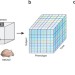
Deep phenotyping predicts Huntington's genotype
Comprehensive phenotyping of Huntington's disease in mice yields phenotypic signatures of disease-causing genetic variants.
- Douglas M Ruderfer
- & Joel T Dudley
-
Brief Communication |
 Systematic comparison of CRISPR/Cas9 and RNAi screens for essential genes
Systematic comparison of CRISPR/Cas9 and RNAi screens for essential genes
A side-by-side comparison of CRISPR/Cas9 and RNAi screens for essential genes reveals method-specific differences in performance.
- David W Morgens
- , Richard M Deans
- & Michael C Bassik
-
Brief Communication |
 CRISPR knockout screening outperforms shRNA and CRISPRi in identifying essential genes
CRISPR knockout screening outperforms shRNA and CRISPRi in identifying essential genes
CRISPR knockout screens outperform shRNA and CRISPR-interference screens in a side-by-side comparison.
- Bastiaan Evers
- , Katarzyna Jastrzebski
- & Rene Bernards
-
News & Views |
 Mapping regulatory elements
Mapping regulatory elements
Regulatory genomic elements can now be studied in their native context using two CRISPR-based high-throughput approaches.
- Yuexin Zhou
- & Wensheng Wei
-
Article |
 Optimized sgRNA design to maximize activity and minimize off-target effects of CRISPR-Cas9
Optimized sgRNA design to maximize activity and minimize off-target effects of CRISPR-Cas9
Genome-wide sgRNA libraries based on rules for on-target activity improve results of Cas9-based screens and facilitate a further refinement of on- and off-target prediction algorithms.
- John G Doench
- , Nicolo Fusi
- & David E Root
-
Article |
 Improving drug discovery with high-content phenotypic screens by systematic selection of reporter cell lines
Improving drug discovery with high-content phenotypic screens by systematic selection of reporter cell lines
A method to systematically identify optimal biomarkers improves high-content screening for drug candidates.
- Jungseog Kang
- , Chien-Hsiang Hsu
- & Lani F Wu
-
Article |
 Massively parallel high-order combinatorial genetics in human cells
Massively parallel high-order combinatorial genetics in human cells
Massively parallel genetic screening reveals synergies between miRNAs regulating cancer cell proliferation and drug resistance.
- Alan S L Wong
- , Gigi C G Choi
- & Timothy K Lu
-
Analysis
| Open Access Prediction of human population responses to toxic compounds by a collaborative competition
Prediction of human population responses to toxic compounds by a collaborative competition
Crowdsourcing analysis improves computational prediction of cytotoxic responses in humans.
- Federica Eduati
- , Lara M Mangravite
- & Julio Saez-Rodriguez
-
Analysis |
 High-throughput spatial mapping of single-cell RNA-seq data to tissue of origin
High-throughput spatial mapping of single-cell RNA-seq data to tissue of origin
Single cells profiled by RNA-seq are rapidly assigned to their location in a complextissue using data in gene expression atlases.
- Kaia Achim
- , Jean-Baptiste Pettit
- & John C Marioni

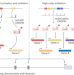 Decoding gene regulation with CRISPR perturbations
Decoding gene regulation with CRISPR perturbations