News & Views |
Featured
-
-
Article
| Open Access Ultra-fast label-free quantification and comprehensive proteome coverage with narrow-window data-independent acquisition
Ultra-fast label-free quantification and comprehensive proteome coverage with narrow-window data-independent acquisition
A new mass spectrometry strategy combines the advantages of data-dependent and data-independent acquisition.
- Ulises H. Guzman
- , Ana Martinez-Val
- & Jesper V. Olsen
-
Article
| Open Access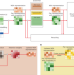 Protein structure prediction with in-cell photo-crosslinking mass spectrometry and deep learning
Protein structure prediction with in-cell photo-crosslinking mass spectrometry and deep learning
Limitations of Alphafold2 structure prediction are addressed by including experimentally determined distance constraints.
- Kolja Stahl
- , Andrea Graziadei
- & Juri Rappsilber
-
Article |
 Increasing the throughput of sensitive proteomics by plexDIA
Increasing the throughput of sensitive proteomics by plexDIA
Proteomics of small sample sizes using data-independent acquisition methods achieves higher throughput with multiplexing.
- Jason Derks
- , Andrew Leduc
- & Nikolai Slavov
-
Research Briefing |
 Spatial characterization of single tumor cells by proteomics
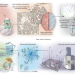
Spatial characterization of single tumor cells by proteomics
By quantifying thousands of proteins in tumor cells in an unbiased manner, Deep Visual Proteomics uncovers mechanisms that drive tumor evolution and reveals new therapeutic targets. The method incorporates advanced microscopy, artificial intelligence and ultra-high-sensitivity proteomics to characterize individual cell identities.
-
News & Views |
 Back and forth with nanopore peptide sequencing
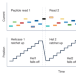
Back and forth with nanopore peptide sequencing
Tagging peptides with DNA allows repeated reads via a helicase in a nanopore with reduced error rates.
- Meni Wanunu
-
News & Views |
 Increasing proteomics throughput
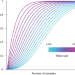
Increasing proteomics throughput
A new dimension for analyzing mass spectrometry data allows rapid quantification of up to 70% more peptides.
- Nikolai Slavov
-
Resource |
 Reconstructing kinase network topologies from phosphoproteomics data reveals cancer-associated rewiring
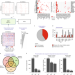
Reconstructing kinase network topologies from phosphoproteomics data reveals cancer-associated rewiring
Improved methods to reconstruct global kinase-signaling networks from phosphoproteomics data are applied in cancer cells.
- Maruan Hijazi
- , Ryan Smith
- & Pedro R. Cutillas
-
Letter |
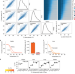 Identifying drug targets in tissues and whole blood with thermal-shift profiling
Identifying drug targets in tissues and whole blood with thermal-shift profiling
The targets of small-molecule drugs are detected in tissue and blood using thermal proteome assays.
- Jessica Perrin
- , Thilo Werner
- & Giovanna Bergamini
-
Resource |
 Co-regulation map of the human proteome enables identification of protein functions
Co-regulation map of the human proteome enables identification of protein functions
Human protein co-regulation map is derived from a large set of quantitative proteomics experiments.
- Georg Kustatscher
- , Piotr Grabowski
- & Juri Rappsilber
-
Analysis |
 Multi-omic measurements of heterogeneity in HeLa cells across laboratories
Multi-omic measurements of heterogeneity in HeLa cells across laboratories
Systems-wide analysis of HeLa cell lines from 13 labs identifies substantial molecular and phenotypic variability.
- Yansheng Liu
- , Yang Mi
- & Ruedi Aebersold
-
News & Views |
 Proteomics goes parallel
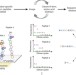
Proteomics goes parallel
Massively parallel sequencing of peptides could signal a new era of high-throughput proteomics.
- Ben C Collins
- & Ruedi Aebersold
-
Letter |
 Revealing the cellular degradome by mass spectrometry analysis of proteasome-cleaved peptides
Revealing the cellular degradome by mass spectrometry analysis of proteasome-cleaved peptides
Cleavage products of the proteasome are profiled by mass spectrometry.
- Hila Wolf-Levy
- , Aaron Javitt
- & Yifat Merbl
-
Brief Communication |
 Comprehensive identification of peptides in tandem mass spectra using an efficient open search engine
Comprehensive identification of peptides in tandem mass spectra using an efficient open search engine
Peptide identification in proteomics data is improved using an efficient open search engine.
- Hao Chi
- , Chao Liu
- & Si-Min He
-
News & Views |
 Making brain proteomics true to type
Making brain proteomics true to type
Two independent but related strategies use unnatural amino acid incorporation to enable in vivo cell-type-specific proteomics in the mouse brain.
- Rashaun S Wilson
- & Angus C Nairn
-
Resource |
 Global landscape of cell envelope protein complexes in Escherichia coli
Global landscape of cell envelope protein complexes in Escherichia coli
A survey of cell envelope proteins and their interacting partners in E. coli by mass spectrometry reveals insights into essential bacterial processes.
- Mohan Babu
- , Cedoljub Bundalovic-Torma
- & Andrew Emili
-
Letter |
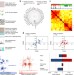 Detection of dysregulated protein-association networks by high-throughput proteomics predicts cancer vulnerabilities
Detection of dysregulated protein-association networks by high-throughput proteomics predicts cancer vulnerabilities
Multiplexed mass spectrometry analysis of breast cancer cell lines uncovers co-regulation of protein abundance that predicts the vulnerabilities of different cancers to drugs.
- John D Lapek Jr
- , Patricia Greninger
- & Wilhelm Haas
-
Correspondence |
 Firmiana: towards a one-stop proteomic cloud platform for data processing and analysis
Firmiana: towards a one-stop proteomic cloud platform for data processing and analysis
- Jinwen Feng
- , Chen Ding
- & Jun Qin
-
Analysis |
 A multicenter study benchmarks software tools for label-free proteome quantification
A multicenter study benchmarks software tools for label-free proteome quantification
LFQbench, a software tool to assess the quality of label-free quantitative proteomics analyses, enables developers to benchmark and improve analytic methods.
- Pedro Navarro
- , Jörg Kuharev
- & Stefan Tenzer
-
Resource |
 The quantitative and condition-dependent Escherichia coli proteome
The quantitative and condition-dependent Escherichia coli proteome
A large-scale quantification of Escherichia coli proteomes across 22 experimental conditions.
- Alexander Schmidt
- , Karl Kochanowski
- & Matthias Heinemann
-
News & Views |
 Illuminating the dark matter of shotgun proteomics
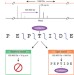
Illuminating the dark matter of shotgun proteomics
Many of the unassignable spectra in proteomics data represent peptides with post-translational modifications.
- Owen S Skinner
- & Neil L Kelleher
-
News & Views |
 Drafts of the human proteome
Drafts of the human proteome
Two studies catalog nearly all human proteins and their abundance throughout the body.
- Robert T Lawrence
- & Judit Villén
-
News & Views |
 SORTing out cellular proteomes in vivo
SORTing out cellular proteomes in vivo
Incorporation of a new unnatural amino acid during protein synthesis enables proteomic profiling of specific cell types in whole animals.
- Tao Peng
- & Howard C Hang
-
Correspondence |
Alternative splicing and protein interaction data sets
- David Talavera
- , David L Robertson
- & Simon C Lovell
-
Correspondence |
 ProtoNet: charting the expanding universe of protein sequences
ProtoNet: charting the expanding universe of protein sequences
- Nadav Rappoport
- , Nathan Linial
- & Michal Linial

 A new mass analyzer shakes up the proteomics field
A new mass analyzer shakes up the proteomics field