Research Briefing |
Featured
-
-
Brief Communication
| Open Access Near-infrared co-illumination of fluorescent proteins reduces photobleaching and phototoxicity
Near-infrared co-illumination of fluorescent proteins reduces photobleaching and phototoxicity
A dual illumination method reduces photobleaching for green and yellow fluorescent proteins.
- Lucie Ludvikova
- , Emma Simon
- & Agathe Espagne
-
Article |
 High-efficiency transgene integration by homology-directed repair in human primary cells using DNA-PKcs inhibition
High-efficiency transgene integration by homology-directed repair in human primary cells using DNA-PKcs inhibition
A small molecule enhances targeted gene integration at therapeutically relevant loci in human primary cells.
- Sridhar Selvaraj
- , William N. Feist
- & Matthew H. Porteus
-
Article |
 PhyloVelo enhances transcriptomic velocity field mapping using monotonically expressed genes
PhyloVelo enhances transcriptomic velocity field mapping using monotonically expressed genes
A refined velocity model improves cell fate mapping with lineage-traced scRNA-seq data.
- Kun Wang
- , Liangzhen Hou
- & Zheng Hu
-
Matters Arising |
 Reply to: Methodological concerns and lack of evidence for single-synapse RNA-seq
Reply to: Methodological concerns and lack of evidence for single-synapse RNA-seq
- Muchun Niu
- & Chenghang Zong
-
Research Briefing |
 Breaking boundaries in whole-body imaging and disease understanding with wildDISCO
Breaking boundaries in whole-body imaging and disease understanding with wildDISCO
Wild three-dimensional imaging of solvent-cleared organs (wildDISCO) uses conventional antibodies for whole-body staining in mice, creating comprehensive biological atlases of nerves, blood vessels and lymphatics. It uncovers pathological changes, such as tertiary lymphoid structures in cancer, and it enables the precise tracking of therapeutic molecules and cells, enhancing our understanding of disease pathology and treatment.
-
Article
| Open Access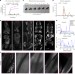 Whole-body cellular mapping in mouse using standard IgG antibodies
Whole-body cellular mapping in mouse using standard IgG antibodies
Whole mice are immunolabeled with standard antibodies using wildDISCO.
- Hongcheng Mai
- , Jie Luo
- & Ali Ertürk
-
Article |
 Prediction of on-target and off-target activity of CRISPR–Cas13d guide RNAs using deep learning
Prediction of on-target and off-target activity of CRISPR–Cas13d guide RNAs using deep learning
A machine learning model predicts on-target and off-target activity of Cas13d in human cells.
- Hans-Hermann Wessels
- , Andrew Stirn
- & Neville E. Sanjana
-
Research Briefing |
 High-throughput screening of TnpB proteins identifies hypercompact genome editors
High-throughput screening of TnpB proteins identifies hypercompact genome editors
TnpB proteins are hypercompact RNA-guided DNA endonucleases. By systematically mining and characterizing TnpB proteins from the IS605 family, we identified potent genome editors and established a framework for high-throughput annotation and screening of TnpB systems in prokaryotic genomes.
-
Article |
 Evolutionary mining and functional characterization of TnpB nucleases identify efficient miniature genome editors
Evolutionary mining and functional characterization of TnpB nucleases identify efficient miniature genome editors
Compact genome editors were discovered by mining a transposon library.
- Guanghai Xiang
- , Yuanqing Li
- & Haoyi Wang
-
Research Briefing |
 Efficient A-to-C base editing with high specificity
Efficient A-to-C base editing with high specificity
Generating A-to-C transversions in specific targets via base editing technology has been challenging. By fusing an evolved alkyladenine DNA glycosylase with an engineered adenine deaminase TadA-8e variant and nickase Cas9, we have developed A-to-C base editors that generate precise and efficient A-to-C transversions in cells and in mouse embryos, expanding the possible applications of base editing.
-
Brief Communication
| Open Access Single-cell m6A mapping in vivo using picoMeRIP–seq
Single-cell m6A mapping in vivo using picoMeRIP–seq
A picogram-scale m6A mapping method is applied to single mouse oocytes and embryos in vivo.
- Yanjiao Li
- , Yunhao Wang
- & John Arne Dahl
-
Article |
 Adenine transversion editors enable precise, efficient A•T-to-C•G base editing in mammalian cells and embryos
Adenine transversion editors enable precise, efficient A•T-to-C•G base editing in mammalian cells and embryos
A base editor for precise adenine transversions is demonstrated in mouse embryos.
- Liang Chen
- , Mengjia Hong
- & Dali Li
-
Brief Communication
| Open Access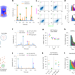 High-throughput measurement of the content and properties of nano-sized bioparticles with single-particle profiler
High-throughput measurement of the content and properties of nano-sized bioparticles with single-particle profiler
Lipid nanoparticles, viruses, exosomes and liposomes are characterized by analysis of fluorescence fluctuations.
- Taras Sych
- , Jan Schlegel
- & Erdinc Sezgin
-
Brief Communication |
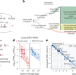 High-throughput RNA isoform sequencing using programmed cDNA concatenation
High-throughput RNA isoform sequencing using programmed cDNA concatenation
Programmable concatenation of cDNA molecules increases the throughput of PacBio sequencing about 15-fold.
- Aziz M. Al’Khafaji
- , Jonathan T. Smith
- & Nir Hacohen
-
Research Briefing |
 CRISPR-free, strand-selective mitochondrial DNA base editing using a nickase
CRISPR-free, strand-selective mitochondrial DNA base editing using a nickase
The fusion of a programmable transcription-activator-like effector (TALE) protein with a nickase, in conjunction with a deaminase, enables efficient and strand-selective DNA base editing. This approach has the potential to advance our understanding and treatment of diseases associated with mutations in the mitochondrial or nuclear genome.
-
Article |
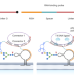 Spatial imaging of glycoRNA in single cells with ARPLA
Spatial imaging of glycoRNA in single cells with ARPLA
GlycoRNA imaging in cells is enabled with a proximity ligation assay.
- Yuan Ma
- , Weijie Guo
- & Yi Lu
-
Article
| Open Access Strand-selective base editing of human mitochondrial DNA using mitoBEs
Strand-selective base editing of human mitochondrial DNA using mitoBEs
Mitochondrial point mutations are corrected in human cells using a transcription activator-like effector (TALE)-based system.
- Zongyi Yi
- , Xiaoxue Zhang
- & Wensheng Wei
-
News & Views |
 Subtle cell states resolved in single-cell data
Subtle cell states resolved in single-cell data
SEACells identifies rare cell states in large datasets, enabling atlas-scale studies.
- Caleb Lareau
-
Article |
 Deep learning models to predict the editing efficiencies and outcomes of diverse base editors
Deep learning models to predict the editing efficiencies and outcomes of diverse base editors
The best base editor for specific applications is predicted with a deep learning model.
- Nahye Kim
- , Sungchul Choi
- & Hyongbum Henry Kim
-
Article |
 A prime editor mouse to model a broad spectrum of somatic mutations in vivo
A prime editor mouse to model a broad spectrum of somatic mutations in vivo
Prime-editing mouse models enable the study of specific cancer mutations in vivo.
- Zackery A. Ely
- , Nicolas Mathey-Andrews
- & Tyler Jacks
-
-
Article
| Open Access Efficient prime editing in mouse brain, liver and heart with dual AAVs
Efficient prime editing in mouse brain, liver and heart with dual AAVs
Prime editors are efficiently delivered to mouse organs with AAV vectors.
- Jessie R. Davis
- , Samagya Banskota
- & David R. Liu
-
Article |
 Efficient engineering of human and mouse primary cells using peptide-assisted genome editing
Efficient engineering of human and mouse primary cells using peptide-assisted genome editing
Peptide-assisted genome editing enables efficient single and multiplex editing in hematopoietic cells.
- Zhen Zhang
- , Amy E. Baxter
- & Junwei Shi
-
Article
| Open Access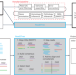 Inference of phylogenetic trees directly from raw sequencing reads using Read2Tree
Inference of phylogenetic trees directly from raw sequencing reads using Read2Tree
Phylogenetic trees are generated from sequencing reads without genome assembly or annotation.
- David Dylus
- , Adrian Altenhoff
- & Christophe Dessimoz
-
Research Briefing |
 Augmenting electron microscopy with barcoded gene reporters
Augmenting electron microscopy with barcoded gene reporters
Genetically encoded concentric barcodes function as multiplexed electron microscopy (EM) gene reporters (EMcapsulins) for cell culture and in vivo models. These barcodes can be used to visualize gene expression information as automatically labeled overlays on volume EM data.
-
Correspondence |
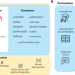 The scverse project provides a computational ecosystem for single-cell omics data analysis
The scverse project provides a computational ecosystem for single-cell omics data analysis
- Isaac Virshup
- , Danila Bredikhin
- & Fabian J. Theis
-
Article |
 RIP-PEN-seq identifies a class of kink-turn RNAs as splicing regulators
RIP-PEN-seq identifies a class of kink-turn RNAs as splicing regulators
Immunoprecipitation coupled with paired-end sequencing of non-coding RNAs identifies a class of splicing regulators.
- Bin Li
- , Shurong Liu
- & Jianhua Yang
-
Research Briefing |
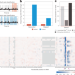 Exploring tRNAs and their modifications and crosstalk using Nano-tRNAseq
Exploring tRNAs and their modifications and crosstalk using Nano-tRNAseq
Nano-tRNAseq is a nanopore-based, cost-effective and high-throughput approach to quantify transfer RNA (tRNA) abundances and modifications simultaneously, providing a framework to study the ‘tRNAome’ at single-molecule resolution. We envision that Nano-tRNAseq will enable us to study the role of tRNA molecules and their modifications in a wide variety of contexts.
-
Research Briefing |
 Imaging the biological microcosmos with a tiny telescope
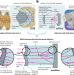
Imaging the biological microcosmos with a tiny telescope
We present the Schmidt objective, a novel astronomy-inspired concept for designing multi-immersion microscope objectives with high numerical aperture, long working distance and large field of view. The Schmidt objective uses a spherical mirror and a correction plate to focus light. It is well suited for high-resolution imaging deep inside cleared biological samples.
-
Article
| Open Access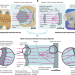 Reflective multi-immersion microscope objectives inspired by the Schmidt telescope
Reflective multi-immersion microscope objectives inspired by the Schmidt telescope
Imaging of large, cleared samples in diverse media is achieved using a mirror objective.
- Fabian F. Voigt
- , Anna Maria Reuss
- & Fritjof Helmchen
-
Research Briefing |
 Measuring the impact of chromatin context on transcription factor binding affinities
Measuring the impact of chromatin context on transcription factor binding affinities
We have designed a method, binding affinities to native chromatin by sequencing (BANC-seq), to determine the transcription factor concentrations required for binding to regulatory elements across the genome. Our study shows that chromatin context and DNA accessibility are key regulators of transcription factor binding.
-
Article
| Open Access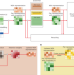 Protein structure prediction with in-cell photo-crosslinking mass spectrometry and deep learning
Protein structure prediction with in-cell photo-crosslinking mass spectrometry and deep learning
Limitations of Alphafold2 structure prediction are addressed by including experimentally determined distance constraints.
- Kolja Stahl
- , Andrea Graziadei
- & Juri Rappsilber
-
Research Briefing |
 Imaging large cell populations with fast, automated super-resolution
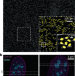
Imaging large cell populations with fast, automated super-resolution
Leveraging advances in hardware and probes, we have combined innovations in optics and algorithms to allow automated single-molecule-based super-resolution imaging at unprecedented throughput. This approach allows us to obtain nanoscale information from large cell populations, bridging the gap between imaging and indirect ensemble methods.
-
Article |
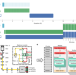 An integrated platform for high-throughput nanoscopy
An integrated platform for high-throughput nanoscopy
A fast data processing platform enables super-resolution microscopy with increased throughput.
- Andrew E. S. Barentine
- , Yu Lin
- & David Baddeley
-
Research Briefing |
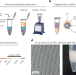 The next generation of single-cell sequencing methods can be microfluidics-free
The next generation of single-cell sequencing methods can be microfluidics-free
Sequencing individual cells in a sample enables scientists to infer the unique characteristics of important subsets. Single-cell sequencing methods that rely on microfluidics for cell barcoding are limited in speed, scale and flexibility. We developed a technique that uses particle-templated emulsification instead of microfluidics and can process millions of cells within minutes.
-
Brief Communication |
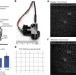 Trans-segmental imaging in the spinal cord of behaving mice
Trans-segmental imaging in the spinal cord of behaving mice
A wearable macroscope enables cellular imaging across multiple spinal segments in behaving mice.
- Pavel Shekhtmeyster
- , Daniela Duarte
- & Axel Nimmerjahn
-
Article
| Open Access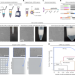 Microfluidics-free single-cell genomics with templated emulsification
Microfluidics-free single-cell genomics with templated emulsification
A microfluidics-free, scalable single-cell RNA-sequencing method produces high-quality transcriptomes.
- Iain C. Clark
- , Kristina M. Fontanez
- & Adam R. Abate
-
Brief Communication
| Open Access Interstrand crosslinking of homologous repair template DNA enhances gene editing in human cells
Interstrand crosslinking of homologous repair template DNA enhances gene editing in human cells
The efficiency of homologous recombination is increased by crosslinking template DNA.
- Hannah I. Ghasemi
- , Julien Bacal
- & Chris D. Richardson
-
Brief Communication
| Open Access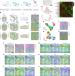 High-plex protein and whole transcriptome co-mapping at cellular resolution with spatial CITE-seq
High-plex protein and whole transcriptome co-mapping at cellular resolution with spatial CITE-seq
Co-indexing of transcriptomes and epitopes is extended to the spatial dimension with large protein panels.
- Yang Liu
- , Marcello DiStasio
- & Rong Fan
-
Article
| Open Access Engineered human hepatocyte organoids enable CRISPR-based target discovery and drug screening for steatosis
Engineered human hepatocyte organoids enable CRISPR-based target discovery and drug screening for steatosis
Organoid models of early liver disease aid target discovery and drug screening.
- Delilah Hendriks
- , Jos F. Brouwers
- & Hans Clevers
-
Article
| Open Access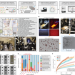 High-throughput microbial culturomics using automation and machine learning
High-throughput microbial culturomics using automation and machine learning
A machine learning isolation and genotyping platform enable high-throughput bacterial culture generation.
- Yiming Huang
- , Ravi U. Sheth
- & Harris H. Wang
-
Article
| Open Access Prediction of prime editing insertion efficiencies using sequence features and DNA repair determinants
Prediction of prime editing insertion efficiencies using sequence features and DNA repair determinants
Prime editing efficiency is affected by insertion sequence features, secondary structure and DNA repair context.
- Jonas Koeppel
- , Juliane Weller
- & Leopold Parts
-
Research Briefing |
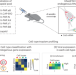 Spatial tropism profiling of AAV vectors by ultrasensitive sequential FISH in tissue
Spatial tropism profiling of AAV vectors by ultrasensitive sequential FISH in tissue
Ultrasensitive sequential fluorescence in situ hybridization (USeqFISH) enables multiplexed detection of the expression of endogenous and exogenous genes delivered by adeno-associated virus (AAV) vectors in intact tissue. USeqFISH provides a spatial map of AAV tropism with high throughput and resolution.
-
Research Briefing |
 CLASH enables large-scale parallel knock-in for cell engineering
CLASH enables large-scale parallel knock-in for cell engineering
CLASH is a CRISPR-based platform that enables the parallel knock-in via homology-directed repair of a large pool of transgene variants encoded in adeno-associated virus vectors. CLASH can be applied to the systematic and unbiased selection of favorable features in T cells and, in principle, other cell types.
-
Article |
 Massively parallel knock-in engineering of human T cells
Massively parallel knock-in engineering of human T cells
Unbiased selection of CAR-T cells is achieved by massively parallel knock-in engineering.
- Xiaoyun Dai
- , Jonathan J. Park
- & Sidi Chen
-
Article
| Open Access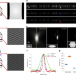 Three-dimensional structured illumination microscopy with enhanced axial resolution
Three-dimensional structured illumination microscopy with enhanced axial resolution
Enhanced axial resolution for 3D SIM is achieved with deep learning or four-beam interference.
- Xuesong Li
- , Yicong Wu
- & Hari Shroff
-
Article |
 Predicting prime editing efficiency and product purity by deep learning
Predicting prime editing efficiency and product purity by deep learning
The design of prime editing guide RNAs is optimized by deep learning.
- Nicolas Mathis
- , Ahmed Allam
- & Gerald Schwank
-
Article |
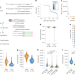 Droplet-based transcriptome profiling of individual synapses
Droplet-based transcriptome profiling of individual synapses
High-throughput profiling of the transcriptomes of individual synapses shows molecular heterogeneity.
- Muchun Niu
- , Wenjian Cao
- & Chenghang Zong
-
Research Briefing |
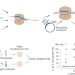 Single-cell recording of cellular RNAs in bacteria
Single-cell recording of cellular RNAs in bacteria
The collection of RNAs in a cell reflects its identity and behavior. Although probing these RNAs in the present is commonplace, peering into the past remains challenging. We developed TIGER (transcribed RNAs inferred by genetically encoded records) to record selected RNAs at the single-cell level, enabling us to connect the past with the present.
Browse broader subjects
Browse narrower subjects
- Analytical biochemistry
- Behavioural methods
- Bioinformatics
- Biological models
- Biophysical methods
- Cytological techniques
- Electrophysiology
- Epigenetics analysis
- Experimental organisms
- Gene delivery
- Gene expression analysis
- Genetic engineering
- Genetic techniques
- Genomic analysis
- High-throughput screening
- Imaging
- Immunological techniques
- Isolation, separation and purification
- Lab-on-a-chip
- Mass spectrometry
- Metabolomics
- Microbiology techniques
- Microscopy
- Molecular engineering
- Nanobiotechnology
- Optogenetics
- Proteomic analysis
- Sensors and probes
- Sequencing
- Software
- Optical spectroscopy
- Structure determination

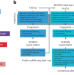 Benchmarking of single-cell ATAC sequencing tools
Benchmarking of single-cell ATAC sequencing tools