Featured
-
-
Letter |
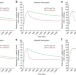 Antibody recycling by engineered pH-dependent antigen binding improves the duration of antigen neutralization
Antibody recycling by engineered pH-dependent antigen binding improves the duration of antigen neutralization
The typical therapeutic antibody binds only two target antigen molecules during its lifetime. Igawa et al. describe a method for engineering antibody recycling in vivo, suggesting an approach to reduce the size and/or frequency of dosage with therapeutic antibodies.
- Tomoyuki Igawa
- , Shinya Ishii
- & Kunihiro Hattori
-
Editorial |
Making a mark
High-throughput technologies are enabling epigenetic modifications to be mapped on a genome-wide scale, but whether such knowledge can be rapidly translated into biomedical applications remains unclear.
-
News & Views |
Taking the measure of the methylome
Two comparative studies from the International Human Epigenome Project find high concordance between different methods for measuring genomic methylation.
- Stephan Beck
-
News & Views |
 Timing is everything in the human embryo
Timing is everything in the human embryo
A noninvasive imaging method for predicting how human embryos will develop may improve the success and safety of in vitro fertilization.
- Ann A Kiessling
-
Correspondence |
 ProHits: integrated software for mass spectrometry–based interaction proteomics
ProHits: integrated software for mass spectrometry–based interaction proteomics
- Guomin Liu
- , Jianping Zhang
- & Anne-Claude Gingras
-
Review Article |
 Genomics tools for unraveling chromosome architecture
Genomics tools for unraveling chromosome architecture
- Bas van Steensel
- & Job Dekker
-
-
Article |
 Non-invasive imaging of human embryos before embryonic genome activation predicts development to the blastocyst stage
Non-invasive imaging of human embryos before embryonic genome activation predicts development to the blastocyst stage
The grading systems used by in vitro fertilization clinics cannot determine reliably whether a given embryo will lead to a successful pregnancy. Wong et al. address one part of this problem by showing that development of an embryo to the blastocyst stage can be predicted with high confidence at day 2 post fertilization.
- Connie C Wong
- , Kevin E Loewke
- & Renee A Reijo Pera
-
Letter |
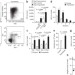 Substrate elasticity provides mechanical signals for the expansion of hemopoietic stem and progenitor cells
Substrate elasticity provides mechanical signals for the expansion of hemopoietic stem and progenitor cells
Biomechanical forces may be an effective approach for controlling the behavior of stem cells in vitro. Holst et al. show that the elasticity of a tropoelastin matrix expands hematopoietic stem and progenitor cells.
- Jeff Holst
- , Sarah Watson
- & John E J Rasko
-
Analysis |
 Comparison of sequencing-based methods to profile DNA methylation and identification of monoallelic epigenetic modifications
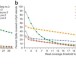
Comparison of sequencing-based methods to profile DNA methylation and identification of monoallelic epigenetic modifications
Methods for profiling DNA methylation differ in the physical principles used to detect modified cytosines. Harris et al. compare the performances of four sequencing-based technologies for genome-wide analysis of DNA methylation and combine two methods to enable detection of allelic differences in epigenetic marks.
- R Alan Harris
- , Ting Wang
- & Joseph F Costello
-
Analysis |
 Quantitative comparison of genome-wide DNA methylation mapping technologies
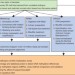
Quantitative comparison of genome-wide DNA methylation mapping technologies
Comparison of the methylation patterns of cells in different developmental or disease states can help to elucidate both normal and pathological regulatory mechanisms. Bock et al. evaluate the ability of three sequencing-based methods and one microarray-based technology to detect differentially methylated regions on a genome-wide scale.
- Christoph Bock
- , Eleni M Tomazou
- & Alexander Meissner
-
-
Correspondence |
Biofortified sorghum in Africa: using problem formulation to inform risk assessment
- Karen E Hokanson
- , Norman C Ellstrand
- & Alan F Raybould
-
Letter |
 Monoclonal antibodies isolated without screening by analyzing the variable-gene repertoire of plasma cells
Monoclonal antibodies isolated without screening by analyzing the variable-gene repertoire of plasma cells
Isolation of antigen-specific antibodies or antibody fragments, whether through B-cell immortalization or recombinant libraries, generally requires laborious screening. Reddy et al. circumvent this step using high-throughput sequencing of plasma cells and bioinformatic analysis of the variable-gene repertoire.
- Sai T Reddy
- , Xin Ge
- & George Georgiou
-
Resource |
 High-throughput generation, optimization and analysis of genome-scale metabolic models
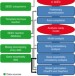
High-throughput generation, optimization and analysis of genome-scale metabolic models
Reconstructing a metabolic model from the genome sequence of an organism is a useful but arduous approach for predicting phenotypes. Henry et al. describe a resource that automates most of this process and apply it to create >100 new metabolic models of microbes.
- Christopher S Henry
- , Matthew DeJongh
- & Rick L Stevens
-
Editorial |
MAQC-II: analyze that!
The MAQC consortium's latest study suggests that human error in handling DNA microarray data analysis software could delay the technology's wider adoption in the clinic.
-
-
News & Views |
 Microarrays in the clinic
Microarrays in the clinic
The MicroArray Quality Control (MAQC) consortium has evaluated methods for making clinically useful predictions from large-scale gene expression data.
- Guy W Tillinghast
-
Article |
 The MicroArray Quality Control (MAQC)-II study of common practices for the development and validation of microarray-based predictive models
The MicroArray Quality Control (MAQC)-II study of common practices for the development and validation of microarray-based predictive models
The Microarray Quality Control consortium pitted 36 teams against each other to evaluate methods for creating genomic classifiers, computational tools for interpreting gene expression profiles. The performance of the classifiers on blinded validation data—and metadata on the analytic methods—reveal the challenges facing the field.
- Leming Shi
- , Gregory Campbell
- & Russell D Wolfinger
-
Analysis |
 Discovery and characterization of chromatin states for systematic annotation of the human genome
Discovery and characterization of chromatin states for systematic annotation of the human genome
Which of the possible combinations of epigenetic marks have biological significance is a major question in epigenetics. Analyzing data from human T-cells, Ernst and Kellis discover 51 distinct, recurring combinations of histone modifications that can be correlated with the functional annotations of the underlying DNA sequences.
- Jason Ernst
- & Manolis Kellis
-
Article |
 Rapid profiling of a microbial genome using mixtures of barcoded oligonucleotides
Rapid profiling of a microbial genome using mixtures of barcoded oligonucleotides
To identify genes affecting traits of interest in E. coli, Warner et al. describe a method to rapidly create and assay rationally mutated versions of every gene. The approach is applied to several traits, including tolerance to cellulosic hydrolysate, a biofuel precursor.
- Joseph R Warner
- , Philippa J Reeder
- & Ryan T Gill
-
Article
| Open Access Genome sequence of the model mushroom Schizophyllum commune
Genome sequence of the model mushroom Schizophyllum commune
Much remains to be learned about the biology of mushrooms, which are an important source of food as well as secondary metabolites and enzymes of biotechnological importance. Ohm et al. report the sequence of the genetically tractable species Schizophyllum commune and identify genes involved in the formation of fruiting bodies and the degradation of lignocellulose.
- Robin A Ohm
- , Jan F de Jong
- & Han A B Wösten
-
Perspective |
 Options and considerations when selecting a quantitative proteomics strategy
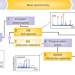
Options and considerations when selecting a quantitative proteomics strategy
- Bruno Domon
- & Ruedi Aebersold
-
Correspondence |
Guidelines for reporting the use of column chromatography in proteomics
- Andrew R Jones
- , Kathleen Carroll
- & Norman W Paton
-
Correspondence |
Guidelines for reporting the use of gel image informatics in proteomics
- Christine Hoogland
- , Martin O'Gorman
- & Andrew R Jones
-
Correspondence |
Guidelines for reporting the use of capillary electrophoresis in proteomics
- Paula J Domann
- , Satoko Akashi
- & Chris F Taylor
-
News & Views |
 Systematic phenotyping of mouse mutants
Systematic phenotyping of mouse mutants
Comprehensive phenotypic screening of knockout mice highlights the pleiotropic functions of secreted and transmembrane proteins.
- Wolfgang Wurst
- & Martin Hrabe de Angelis
-
News & Views |
 Paring down signaling complexity
Paring down signaling complexity
A new method for characterizing signaling responses to pairs of agonists predicts how cells react to higher-order combinations of external stimuli.
- Kevin A Janes
-
Correspondence |
 Minimum information about a protein affinity reagent (MIAPAR)
Minimum information about a protein affinity reagent (MIAPAR)
- Julie Bourbeillon
- , Sandra Orchard
- & David Sherman
-
Feature |
 Proteomics retrenches
Proteomics retrenches
Improvements in technology are making proteomics research less descriptive and more analytic, but the field has yet to deliver on its aspirations.
- Peter Mitchell
-
Resource |
 A mouse knockout library for secreted and transmembrane proteins
A mouse knockout library for secreted and transmembrane proteins
Tang et al. present the first large-scale, gene-specific library of knockout mice. They disrupt 472 genes encoding secreted or transmembrane proteins and report the results of a comprehensive phenotypic analysis.
- Tracy Tang
- , Li Li
- & Frederic J de Sauvage
-
-
News & Views |
 Scalable pluripotent stem cell culture
Scalable pluripotent stem cell culture
Large-scale production of human embryonic stem cells will require improved culture methods.
- Larry A Couture
-
News & Views |
 Raising the bar for cancer therapy models
Raising the bar for cancer therapy models
Can mouse cancer models predict the results of phase 3 clinical trials?
- Giulio Francia
- & Robert S Kerbel
-
Article |
 Assessing therapeutic responses in Kras mutant cancers using genetically engineered mouse models
Assessing therapeutic responses in Kras mutant cancers using genetically engineered mouse models
Genetically engineered mouse models of cancer simulate the spontaneous development of tumors in their native tissue environment. Singh et al. establish their ability to predict the efficacy of different treatment regimens by comparing clinical trial results to equivalent experiments in mutated KRAS-driven mouse models of pancreatic and lung cancer.
- Mallika Singh
- , Anthony Lima
- & Leisa Johnson
-
Analysis |
 Comparative assessment of methods for aligning multiple genome sequences
Comparative assessment of methods for aligning multiple genome sequences
Mining information from genomes often begins by aligning the sequences to identify evolutionarily conserved regions. Chen et al. assess the performance of four commonly used multiple sequence alignment tools.
- Xiaoyu Chen
- & Martin Tompa
-
Letter |
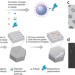 Single-molecule enzyme-linked immunosorbent assay detects serum proteins at subfemtomolar concentrations
Single-molecule enzyme-linked immunosorbent assay detects serum proteins at subfemtomolar concentrations
Rissin et al. increase the sensitivity of sandwich ELISA by segregating beads bearing a single enzyme-labeled immunoconjugate into femtoliter-volume reaction chambers. As the small volume of each well permits detection of extremely low levels of fluorescence, protein abundance is determined by counting the number of fluorescent wells as a percentage of the number of wells containing beads.
- David M Rissin
- , Cheuk W Kan
- & David C Duffy
-
Resource |
 Analysis of a genome-wide set of gene deletions in the fission yeast Schizosaccharomyces pombe
Analysis of a genome-wide set of gene deletions in the fission yeast Schizosaccharomyces pombe
Genome-wide single-gene deletion libraries can be important tools for understanding the molecular workings of an organism, but have only been created for a single eukaryotic species, Saccharomyces cerevisiae. Now, Kim et al. present a second collection of deletion mutants that covers 98.4% of the genes of Schizosaccharomyces pombe, allowing a systematic comparison of gene essentiality and knockout phenotypes between two eukaryotic species.
- Dong-Uk Kim
- , Jacqueline Hayles
- & Kwang-Lae Hoe
-
-
News & Views |
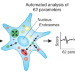 High-content imaging
High-content imaging
Multiparametric imaging of siRNA screening data sheds light on endocytosis.
- Arnold Hayer
- & Tobias Meyer
-
Resource |
 High-resolution DNA analysis of human embryonic stem cell lines reveals culture-induced copy number changes and loss of heterozygosity
High-resolution DNA analysis of human embryonic stem cell lines reveals culture-induced copy number changes and loss of heterozygosity
Cultured human embryonic stem cells often acquire chromosomal abnormalities that could be detrimental in certain applications. Närvä et al. report the highest-resolution genetic analysis of these cells to date and identify genes whose expression is altered by culture-induced genetic changes.
- Elisa Närvä
- , Reija Autio
- & Riitta Lahesmaa
-
Article |
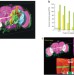 V3D enables real-time 3D visualization and quantitative analysis of large-scale biological image data sets
V3D enables real-time 3D visualization and quantitative analysis of large-scale biological image data sets
High-throughput imaging generates massive data sets that are difficult to quantitatively analyze by hand. Peng et al. describe customizable software for visualizing and working with multi-gigabyte three-dimensional images in real time.
- Hanchuan Peng
- , Zongcai Ruan
- & Eugene W Myers
-
Letter |
 Isotopic labeling of terminal amines in complex samples identifies protein N-termini and protease cleavage products
Isotopic labeling of terminal amines in complex samples identifies protein N-termini and protease cleavage products
Many proteases are important drug targets, but identification of their substrates remains challenging. By using polymers to selectively isolate N-terminal peptides generated by proteolysis of complex samples, Kleifeld et al. identify substrates of clinically relevant proteases with broad specificity.
- Oded Kleifeld
- , Alain Doucet
- & Christopher M Overall
-
Article |
 Directed evolution of a magnetic resonance imaging contrast agent for noninvasive imaging of dopamine
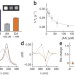
Directed evolution of a magnetic resonance imaging contrast agent for noninvasive imaging of dopamine
Magnetic resonance imaging of hemoglobin in the brain can detect blood flow associated with neural activity, but direct imaging of neurotransmitters would provide a more sensitive measure of neural signal processing. Shapiro et al. use directed evolution to generate a protein probe that enables magnetic resonance imaging of the neurotransmitter dopamine.
- Mikhail G Shapiro
- , Gil G Westmeyer
- & Alan Jasanoff
-
News & Views |
 ChIPs and regulatory bits
ChIPs and regulatory bits
Machine learning reveals combinatorial patterns of transcription factor binding that drive gene expression.
- Xin He
- & Saurabh Sinha
-
News & Views |
 Systematic tracking of cell fate changes
Systematic tracking of cell fate changes
High-throughput measurements across several regulatory levels provide a comprehensive view of ES-cell differentiation.
- Jonghwan Kim
- & Stuart H Orkin
-
Letter |
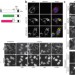 Real-time imaging of hepatitis C virus infection using a fluorescent cell-based reporter system
Real-time imaging of hepatitis C virus infection using a fluorescent cell-based reporter system
Hepatitis C virus research is hampered by the inability to detect individual infected cells. Jones et al. achieve this by imaging the translocation of a fluorescent reporter protein after cleavage by a viral protease in living or fixed cells.
- Christopher T Jones
- , Maria Teresa Catanese
- & Charles M Rice
-
News & Views |
 Enriching quantitative proteomics with SIN
Enriching quantitative proteomics with SIN
A new metric called the normalized spectral index (SIN) provides a simple way to quantify and compare label-free proteomics data.
- Mihaela E Sardiu
- & Michael P Washburn
-
Correspondence |
 High-density resequencing DNA microarrays in public health emergencies
High-density resequencing DNA microarrays in public health emergencies
- Nicolas Berthet
- , India Leclercq
- & Jean-Claude Manuguerra
Browse broader subjects
Browse narrower subjects
- Analytical biochemistry
- Behavioural methods
- Bioinformatics
- Biological models
- Biophysical methods
- Cytological techniques
- Electrophysiology
- Epigenetics analysis
- Experimental organisms
- Gene delivery
- Gene expression analysis
- Genetic engineering
- Genetic techniques
- Genomic analysis
- High-throughput screening
- Imaging
- Immunological techniques
- Isolation, separation and purification
- Lab-on-a-chip
- Mass spectrometry
- Metabolomics
- Microbiology techniques
- Microscopy
- Molecular engineering
- Nanobiotechnology
- Optogenetics
- Proteomic analysis
- Sensors and probes
- Sequencing
- Software
- Optical spectroscopy
- Structure determination

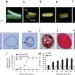 Cultured cambial meristematic cells as a source of plant natural products
Cultured cambial meristematic cells as a source of plant natural products Epigenetic modifications and human disease
Epigenetic modifications and human disease The Cartagena Protocol and genetically modified mosquitoes
The Cartagena Protocol and genetically modified mosquitoes Lawsuits rock Jackson
Lawsuits rock Jackson Detecting methylated bases in real time
Detecting methylated bases in real time