This Month |
Featured
-
-
Article |
 Screening for functional circular RNAs using the CRISPR–Cas13 system
Screening for functional circular RNAs using the CRISPR–Cas13 system
This paper describes a CRISPR–Cas13 system to effectively target circRNAs and screen their functions in vitro and in vivo, which enables the study of relevant circRNA phenotypes in human cell proliferation and in mouse embryogenesis.
- Siqi Li
- , Xiang Li
- & Ling-Ling Chen
-
Article |
 MARS: discovering novel cell types across heterogeneous single-cell experiments
MARS: discovering novel cell types across heterogeneous single-cell experiments
MARS uses a meta-learning strategy for annotating known cell types and identifying novel ones across single-cell RNA-seq datasets.
- Maria Brbić
- , Marinka Zitnik
- & Jure Leskovec
-
Article |
 DeepC: predicting 3D genome folding using megabase-scale transfer learning
DeepC: predicting 3D genome folding using megabase-scale transfer learning
DeepC uses transfer learning-based deep neural networks for predicting genome folding from megabase-scale DNA sequence.
- Ron Schwessinger
- , Matthew Gosden
- & Jim R. Hughes
-
Article |
 Predicting 3D genome folding from DNA sequence with Akita
Predicting 3D genome folding from DNA sequence with Akita
Akita enables three-dimensional genome folding predictions from DNA sequence using a convolutional neural network.
- Geoff Fudenberg
- , David R. Kelley
- & Katherine S. Pollard
-
Brief Communication |
 An efficient KRAB domain for CRISPRi applications in human cells
An efficient KRAB domain for CRISPRi applications in human cells
CRISPRi activities are measured from dCas9 fusions with 57 KRAB domains, which lead to the identification of ZIM3 KRAB–dCas9 fusion as a superior repressor.
- Nader Alerasool
- , Dmitri Segal
- & Mikko Taipale
-
Brief Communication |
 Structure-based validation can drastically underestimate error rate in proteome-wide cross-linking mass spectrometry studies
Structure-based validation can drastically underestimate error rate in proteome-wide cross-linking mass spectrometry studies
The current standard approach for estimating error in proteome-scale cross-linking mass spectrometry datasets has severe limitations. A proposed set of data-quality metrics provides a more accurate assessment of error rate.
- Kumar Yugandhar
- , Ting-Yi Wang
- & Haiyuan Yu
-
Research Highlight |
Mapping RNA–RNA interactions
RIC-seq enables in situ mapping of intra- and intermolecular RNA–RNA interactions.
- Lin Tang
-
Article |
 3D mapping and accelerated super-resolution imaging of the human genome using in situ sequencing
3D mapping and accelerated super-resolution imaging of the human genome using in situ sequencing
OligoFISSEQ combines Oligopaints with fluorescence in situ sequencing to enable the 3D mapping of many regions across the genome in human cells to interrogate genome organization at improved genomic resolution. OligoFISSEQ is compatible with immunochemistry and OligoSTORM for super-resolution imaging.
- Huy Q. Nguyen
- , Shyamtanu Chattoraj
- & C.-ting Wu
-
Research Highlight |
 Radial genome organization
Radial genome organization
Sequencing gradually digested chromatin along the nuclear radius enables the mapping of radial organization of chromatin in human cells.
- Lei Tang
-
-
Perspective |
 Mechanistic modeling of chromatin folding to understand function
Mechanistic modeling of chromatin folding to understand function
This Perspective highlights recently developed computational models for studying chromosome organization, with a focus on how mechanistic modeling helps biologists to interpret the biological function behind the genome structures.
- Chris A. Brackey
- , Davide Marenduzzo
- & Nick Gilbert
-
-
-
Research Highlight |
Charting the human cell landscape
Researchers construct a transcriptome-based single-cell human cell landscape for major human organs.
- Lei Tang
-
Resource
| Open Access Genetic tool development in marine protists: emerging model organisms for experimental cell biology
Genetic tool development in marine protists: emerging model organisms for experimental cell biology
This Resource describes genetic tools for microbial eukaryotes, providing a roadmap for developing genetically tractable organisms.
- Drahomíra Faktorová
- , R. Ellen R. Nisbet
- & Julius Lukeš
-
Research Highlight |
Identifying the chromatin-associated proteome
ChromID identifies proteins associated with various chromatin modifications.
- Arunima Singh
-
Article |
 RecV recombinase system for in vivo targeted optogenomic modifications of single cells or cell populations
RecV recombinase system for in vivo targeted optogenomic modifications of single cells or cell populations
Light-dependent variants of Cre, Dre and Flp enable targeted sparse or single-cell labeling in mouse and zebrafish.
- Shenqin Yao
- , Peng Yuan
- & Ali Cetin
-
Perspective |
 Purification and enrichment of specific chromatin loci
Purification and enrichment of specific chromatin loci
This Perspective discusses the challenges, progress and future of proteome profiling at specific chromatin loci for elucidating genome biology.
- Mathilde Gauchier
- , Guido van Mierlo
- & Jérôme Déjardin
-
-
Comment |
Single-cell biology: beyond the sum of its parts
The field of single-cell RNA sequencing (scRNA-seq) has been paired with genomics, epigenomics, spatial omics, proteomics and imaging to achieve multimodal measurements of individual cellular phenotypes and genotypes. In its purest form, single-cell multimodal omics involves the simultaneous detection of multiple traits in the same cell. More broadly, multimodal omics also encompasses comparative pairing and computational integration of measurements made across multiple distinct cells to reconstruct phenotypes. Here I highlight some of the biological insights gained from multimodal studies and discuss the challenges and opportunities in this emerging field.
- Alexander F. Schier
-
Article |
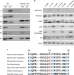 An antibody for analysis of autophagy induction
An antibody for analysis of autophagy induction
A new method of autophagy measurement is based on the detection of phospho-ATG16L1, a conserved early marker of autophagy. Sensitive detection can be achieved in multiple biological systems and assays with advantages over standard methods.
- Wensheng Tian
- , Reham Alsaadi
- & Ryan C. Russell
-
News & Views |
 Mapping m6A and m1A with mutation signatures
Mapping m6A and m1A with mutation signatures
In two novel RNA modification mapping methods, the authors have engineered RNA enzymes and used the enzyme-mediated mutational signatures to map m6A and m1A at single-nucleotide resolution in mammalian RNA.
- Dan Ohtan Wang
-
Article |
 Genome-wide quantification of ADAR adenosine-to-inosine RNA editing activity
Genome-wide quantification of ADAR adenosine-to-inosine RNA editing activity
The Alu editing index (AEI) quantifies genome-wide editing in Alu repeats and allows comparison across samples.
- Shalom Hillel Roth
- , Erez Y. Levanon
- & Eli Eisenberg
-
Article |
 DART-seq: an antibody-free method for global m6A detection
DART-seq: an antibody-free method for global m6A detection
Getting around the limitations of antibody-based N6-methyladenosine (m6A) pulldown, such as high input requirements and cross-reactivity, DART-seq profiles transcriptome-wide m6A occurrences from RNA amounts equivalent to the RNA obtained from 1,000 cells.
- Kate D. Meyer
-
Article |
 Simultaneous profiling of 3D genome structure and DNA methylation in single human cells
Simultaneous profiling of 3D genome structure and DNA methylation in single human cells
Single-nucleus methyl-3C sequencing jointly interrogates 3D chromatin organization and DNA methylation in human cells, and these joint measurements more accurately distinguish different cell types than either unimodal method.
- Dong-Sung Lee
- , Chongyuan Luo
- & Joseph R. Ecker
-
Brief Communication |
 Live imaging of mRNA using RNA-stabilized fluorogenic proteins
Live imaging of mRNA using RNA-stabilized fluorogenic proteins
Pepper, an RNA aptamer that prevents degradation of degron-tagged fluorescent proteins, enables fully genetically encoded fluorescence imaging of mRNA in living cells.
- Jiahui Wu
- , Sara Zaccara
- & Samie R. Jaffrey
-
Brief Communication |
 An efficient auxin-inducible degron system with low basal degradation in human cells
An efficient auxin-inducible degron system with low basal degradation in human cells
An optimized F-box protein–degron pair enables efficient auxin-mediated protein degradation with minimal basal degradation in human cells and is suitable for transmembrane, cytoplasmic and nuclear proteins.
- Shiqian Li
- , Xavier Prasanna
- & Elina Ikonen
-
Article |
 Multiplexed genome engineering by Cas12a and CRISPR arrays encoded on single transcripts
Multiplexed genome engineering by Cas12a and CRISPR arrays encoded on single transcripts
A single transcript encoding Cas12a and an array of CRISPR RNAs enables multiplexed genome engineering, from multiple knockouts to transcriptional activation or repression to orthogonal transcriptional control and editing in the same sample.
- Carlo C. Campa
- , Niels R. Weisbach
- & Randall J. Platt
-
Article |
 FLAM-seq: full-length mRNA sequencing reveals principles of poly(A) tail length control
FLAM-seq: full-length mRNA sequencing reveals principles of poly(A) tail length control
FLAM-seq implements a cDNA library preparation followed by single-molecule sequencing, for determining full-length mRNA molecules, including poly(A) tails.
- Ivano Legnini
- , Jonathan Alles
- & Nikolaus Rajewsky
-
-
-
-
Article |
 LADL: light-activated dynamic looping for endogenous gene expression control
LADL: light-activated dynamic looping for endogenous gene expression control
A fusion of CIBN to dCas9 and a Cry2 bridge enables blue-light-inducible long-range chromatin loops.
- Ji Hun Kim
- , Mayuri Rege
- & Jennifer E. Phillips-Cremins
-
Article |
 MULTI-seq: sample multiplexing for single-cell RNA sequencing using lipid-tagged indices
MULTI-seq: sample multiplexing for single-cell RNA sequencing using lipid-tagged indices
Tagging live single cells and nuclei with lipid- or cholesterol-modified oligonucleotides enables massive scRNA-seq sample multiplexing, identifies doublets and recovers cells with low RNA content.
- Christopher S. McGinnis
- , David M. Patterson
- & Zev J. Gartner
-
Research Highlight |
 Protein translation inside synthetic membraneless organelles
Protein translation inside synthetic membraneless organelles
Phase separation in combination with spatial targeting allows for selective translation of mRNA within synthetic membraneless organelles.
- Lei Tang
-
Brief Communication |
 HiChIRP reveals RNA-associated chromosome conformation
HiChIRP reveals RNA-associated chromosome conformation
HiChIRP combines a modified chromosome conformation capture protocol with enrichment of RNA-associated chromosome conformation to visualize genome-wide looping linked to an RNA of interest.
- Maxwell R. Mumbach
- , Jeffrey M. Granja
- & Howard Y. Chang
-
-
Research Highlight |
Variability in genome structure
Comparisons of genome organization in many individual cells with high-throughput FISH show extensive variation.
- Nicole Rusk
-
-
Article |
 Capturing the dynamics of genome replication on individual ultra-long nanopore sequence reads
Capturing the dynamics of genome replication on individual ultra-long nanopore sequence reads
Replication initiation is stochastic and obscured in population sequencing; D-NAscent reports the use of long nanopore reads to detect base analogs and thus to assess replication initiation at the individual molecule level.
- Carolin A. Müller
- , Michael A. Boemo
- & Conrad A. Nieduszynski
-
Brief Communication |
 Multiplexed detection of proteins, transcriptomes, clonotypes and CRISPR perturbations in single cells
Multiplexed detection of proteins, transcriptomes, clonotypes and CRISPR perturbations in single cells
ECCITE-seq combines the single-cell analysis of multiple modalities, for example transcriptome, immune cell receptors, cell surface proteins and single-guide RNAs.
- Eleni P. Mimitou
- , Anthony Cheng
- & Peter Smibert
-
-
Brief Communication |
 Deep-learning augmented RNA-seq analysis of transcript splicing
Deep-learning augmented RNA-seq analysis of transcript splicing
DARTS first uses public domain data to train a deep neural network to predict differential alternative splicing; the predictions are then combined with observed RNA-seq data in a Bayesian framework to infer changes in alternative splicing between biological samples.
- Zijun Zhang
- , Zhicheng Pan
- & Yi Xing
-
Research Highlight |
Spotlight on Cas12
A search for more type V Cas12 family members turns up unexpected functionality.
- Nicole Rusk
-
Review Article |
 Methods to study RNA–protein interactions
Methods to study RNA–protein interactions
This Review discusses the strengths and limitations of methods to probe RNA–protein interactions.
- Muthukumar Ramanathan
- , Douglas F. Porter
- & Paul A. Khavari
-
-
Research Highlight |
Creating epigenetic memory
A three-component system of epigenetic writers and readers preserves memory over many generations of cells.
- Nicole Rusk
-
Brief Communication |
 Detection of splice isoforms and rare intermediates using multiplexed primer extension sequencing
Detection of splice isoforms and rare intermediates using multiplexed primer extension sequencing
MPE-seq uses pools of reverse-transcription primers to facilitate enrichment of splice isoforms and pre-mRNA splicing intermediates.
- Hansen Xu
- , Benjamin J. Fair
- & Jeffrey A. Pleiss
-
Browse broader subjects
Browse narrower subjects
- Cell division
- Chromatin
- Chromosomes
- CRISPR-Cas systems
- DNA damage and repair
- DNA metabolism
- DNA recombination
- DNA replication
- Epigenetics
- Non-coding RNAs
- Nuclear organization
- Post-translational modifications
- Protein folding
- Proteolysis
- Proteomics
- Riboswitches
- Ribozymes
- RNA metabolism
- RNAi
- Single-molecule biophysics
- Transcription
- Transcriptomics
- Translation
- Transposition

 Ling-Ling Chen
Ling-Ling Chen