Research Highlight |
Featured
-
-
Research Briefing |
 TREX identifies region-specific protein interactors of RNA molecules
TREX identifies region-specific protein interactors of RNA molecules
Interactions between RNA and RNA-binding proteins (RBPs) define the fate and function of every RNA molecule. We present TREX, or targeted RNase H-mediated extraction of crosslinked RBPs, an efficient and accurate method to unbiasedly reveal the protein interactors of specific regions of RNAs isolated from living cells.
-
Article |
 RNA molecular recording with an engineered RNA deaminase
RNA molecular recording with an engineered RNA deaminase
An engineered RNA A-to-I deaminase (rABE) offers low sequence bias, high activity and low background for REMORA (RNA-encoded molecular recording in adenosines) and enables improved molecular recording of RNA–protein interactions.
- Yizhu Lin
- , Samentha Kwok
- & Stephen N. Floor
-
Article
| Open Access Spatiotemporally resolved transcriptomics reveals the subcellular RNA kinetic landscape
Spatiotemporally resolved transcriptomics reveals the subcellular RNA kinetic landscape
TEMPOmap combines pulse-chase metabolic labeling with multiplexed three-dimensional in situ sequencing to simultaneously profile the age and subcellular location of individual RNA molecules from thousands of genes to reveal RNA kinetic landscapes.
- Jingyi Ren
- , Haowen Zhou
- & Xiao Wang
-
Article
| Open Access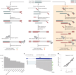 Nano3P-seq: transcriptome-wide analysis of gene expression and tail dynamics using end-capture nanopore cDNA sequencing
Nano3P-seq: transcriptome-wide analysis of gene expression and tail dynamics using end-capture nanopore cDNA sequencing
Nano3P-seq presents a nanopore-based sequencing tool to profile polyA-tailed and non-polyA-tailed transcripts, as well as capture polyA tail length and composition.
- Oguzhan Begik
- , Gregor Diensthuber
- & Eva Maria Novoa
-
Brief Communication |
 An improved imaging system that corrects MS2-induced RNA destabilization
An improved imaging system that corrects MS2-induced RNA destabilization
An improved version of the MS2-MCP system for imaging RNA dynamics involves tethering translation termination factors to tagged mRNAs to bypass destabilization caused by NMD machinery.
- Weihan Li
- , Anna Maekiniemi
- & Robert H. Singer
-
This Month |
Athlete–scientists like to sweat
Some scientists find ways to balance their love of athleticism with research.
- Vivien Marx
-
Research Highlight |
Sequencing RNA isoforms in brain tissue
Single-nuclei isoform RNA sequencing enables cell-type specific isoform analysis in frozen brain samples.
- Lei Tang
-
Research Highlight |
 Sequencing diverse RNA modifications
Sequencing diverse RNA modifications
Nanopore sequencing enables quantitative profiling of pseudouridine and 2′-O-methylation modifications in native RNAs.
- Lei Tang
-
Article |
 Robust single-cell discovery of RNA targets of RNA-binding proteins and ribosomes
Robust single-cell discovery of RNA targets of RNA-binding proteins and ribosomes
Surveying targets by APOBEC-mediated profiling identifies binding sites of RBPs by C-to-U RNA editing. STAMP is isoform specific, can be multiplexed and enables detection of ribosome association in single cells.
- Kristopher W. Brannan
- , Isaac A. Chaim
- & Gene W. Yeo
-
Article |
 Genome-wide quantification of ADAR adenosine-to-inosine RNA editing activity
Genome-wide quantification of ADAR adenosine-to-inosine RNA editing activity
The Alu editing index (AEI) quantifies genome-wide editing in Alu repeats and allows comparison across samples.
- Shalom Hillel Roth
- , Erez Y. Levanon
- & Eli Eisenberg
-
Article |
 DART-seq: an antibody-free method for global m6A detection
DART-seq: an antibody-free method for global m6A detection
Getting around the limitations of antibody-based N6-methyladenosine (m6A) pulldown, such as high input requirements and cross-reactivity, DART-seq profiles transcriptome-wide m6A occurrences from RNA amounts equivalent to the RNA obtained from 1,000 cells.
- Kate D. Meyer
-
Article |
 FLAM-seq: full-length mRNA sequencing reveals principles of poly(A) tail length control
FLAM-seq: full-length mRNA sequencing reveals principles of poly(A) tail length control
FLAM-seq implements a cDNA library preparation followed by single-molecule sequencing, for determining full-length mRNA molecules, including poly(A) tails.
- Ivano Legnini
- , Jonathan Alles
- & Nikolaus Rajewsky
-
-
Brief Communication |
 Deep-learning augmented RNA-seq analysis of transcript splicing
Deep-learning augmented RNA-seq analysis of transcript splicing
DARTS first uses public domain data to train a deep neural network to predict differential alternative splicing; the predictions are then combined with observed RNA-seq data in a Bayesian framework to infer changes in alternative splicing between biological samples.
- Zijun Zhang
- , Zhicheng Pan
- & Yi Xing
-
Brief Communication |
 Detection of splice isoforms and rare intermediates using multiplexed primer extension sequencing
Detection of splice isoforms and rare intermediates using multiplexed primer extension sequencing
MPE-seq uses pools of reverse-transcription primers to facilitate enrichment of splice isoforms and pre-mRNA splicing intermediates.
- Hansen Xu
- , Benjamin J. Fair
- & Jeffrey A. Pleiss
-
Brief Communication |
 COMRADES determines in vivo RNA structures and interactions
COMRADES determines in vivo RNA structures and interactions
In vivo probing of RNA structures with COMRADES yields insight into RNA folding of the ZIKA virus genome and its interaction with host RNAs.
- Omer Ziv
- , Marta M. Gabryelska
- & Eric A. Miska
-
Research Highlight |
Transcripts from a spliceosome
Two new methods provide high-resolution profiles of splicing events in yeast.
- Tal Nawy
-
Research Highlights |
CRISPR–Cas goes RNA
CasRx is a programmable CRISPR–Cas system that targets RNA for efficient knockdown and splicing modulation.
- Stéphane Larochelle
-
Article |
 TimeLapse-seq: adding a temporal dimension to RNA sequencing through nucleoside recoding
TimeLapse-seq: adding a temporal dimension to RNA sequencing through nucleoside recoding
TimeLapse-seq chemically recodes labeled s4U nucleosides to a cytidine derivative and monitors transcriptional dynamics.
- Jeremy A Schofield
- , Erin E Duffy
- & Matthew D Simon
-
Tools in Brief |
Alternative splice atlas
-
Research Highlights |
The proof of splicing is in the proteome
An integrated workflow allows the quantitative interrogation of the impact of alternative splicing at the proteome level.
- Stéphane Larochelle
-
Article |
 Thiol-linked alkylation of RNA to assess expression dynamics
Thiol-linked alkylation of RNA to assess expression dynamics
SLAM seq detects s4U incorporation into RNA and, combined with 3′-end RNA-seq, provides insight into RNA turnover.
- Veronika A Herzog
- , Brian Reichholf
- & Stefan L Ameres
-
Brief Communication |
 Visualizing adenosine-to-inosine RNA editing in single mammalian cells
Visualizing adenosine-to-inosine RNA editing in single mammalian cells
inoFISH uses fluorescence in situ hybridization to visualize and localize adenosine-to-inosine-edited transcripts.
- Ian A Mellis
- , Rohit Gupte
- & Sara H Rouhanifard
-
Method to Watch |
 CRISPR targets RNA
CRISPR targets RNA
Having revolutionized DNA editing, CRISPR turns to RNA.
- Nicole Rusk
-
Commentary |
 Reversible RNA modifications in meiosis and pluripotency
Reversible RNA modifications in meiosis and pluripotency
- Arne Klungland
- , John Arne Dahl
- & Adam Filipczyk
-
Research Highlights |
Topographical transcriptomes
Three experimental methods generate global maps of in vivo RNA interactions.
- Tal Nawy
-
Research Highlights |
Of TRIBE and RNA tagging
Genetic approaches profile RNA–protein interactions.
- Nicole Rusk
-
Research Highlights |
Protein isoforms: more than meets the eye
Alternative splicing imparts distinct functions through isoform-specific protein–protein interactions.
- Stéphane Larochelle
-
Article |
 Robust transcriptome-wide discovery of RNA-binding protein binding sites with enhanced CLIP (eCLIP)
Robust transcriptome-wide discovery of RNA-binding protein binding sites with enhanced CLIP (eCLIP)
Enhanced CLIP yields complex libraries of RNA components of ribonucleoprotein complexes and maintains single-nucleotide resolution of binding sites. eCLIP enables large scale profiling, as demonstrated with the binding profiles of 73 RBPs in two human cancer cell lines.
- Eric L Van Nostrand
- , Gabriel A Pratt
- & Gene W Yeo
-
Research Highlights |
RNA catch and release
The binding of single RNA molecules to individual proteins can be observed in the subcellular compartments of living cells.
- Stéphane Larochelle
-
Article |
 Single-nucleotide-resolution mapping of m6A and m6Am throughout the transcriptome
Single-nucleotide-resolution mapping of m6A and m6Am throughout the transcriptome
Unique mutational signatures induced by cross-linking of m6A-specific antibodies to RNA identify m6A and m6Am residues at single-nucleotide resolution, transcriptome-wide.
- Bastian Linder
- , Anya V Grozhik
- & Samie R Jaffrey
-
Methods in Brief |
RNA caps in bacteria stabilize transcripts
-
Article |
 A superfolding Spinach2 reveals the dynamic nature of trinucleotide repeat–containing RNA
A superfolding Spinach2 reveals the dynamic nature of trinucleotide repeat–containing RNA
The Spinach2 RNA aptamer provides substantially improved imaging over its predecessor Spinach, enabling live-cell imaging of many tagged RNA species including toxic trinucleotide repeat–containing RNAs.
- Rita L Strack
- , Matthew D Disney
- & Samie R Jaffrey
-
Brief Communication |
 Identifying RNA editing sites using RNA sequencing data alone
Identifying RNA editing sites using RNA sequencing data alone
Highly accurate and sensitive predictions of RNA editing sites can be obtained using RNA sequencing data from multiple individuals or species, without relying on matched genomic DNA sequence. Reanalyzing existing RNA sequencing data in this way greatly expands the catalog of human protein recoding events.
- Gokul Ramaswami
- , Rui Zhang
- & Jin Billy Li
-
Article |
 Analysis of alternative cleavage and polyadenylation by 3′ region extraction and deep sequencing
Analysis of alternative cleavage and polyadenylation by 3′ region extraction and deep sequencing
The 3′ region extraction and deep sequencing (3′READS) method accurately identifies cleavage and polyadenylation sites, avoiding common artifacts and detecting sites in A-rich contexts. It was used to greatly expand the number of characterized sites in the mouse genome, including those in long noncoding RNAs.
- Mainul Hoque
- , Zhe Ji
- & Bin Tian
-
Brief Communication |
 Accurate identification of human Alu and non-Alu RNA editing sites
Accurate identification of human Alu and non-Alu RNA editing sites
A computational framework is reported for the accurate and sensitive identification of RNA editing sites from whole-genome DNA and RNA sequences from the same individual.
- Gokul Ramaswami
- , Wei Lin
- & Jin Billy Li
-
Research Highlights |
 Timing an intron's departure
Timing an intron's departure
An in vitro RNA-labeling technique with single-molecule resolution offers a look into the kinetics and the location of splicing.
- Nicole Rusk
-
News & Views |
 Making sense out of nonsense to visualize editing in the fly nervous system
Making sense out of nonsense to visualize editing in the fly nervous system
In vivo methods to capture processing events such as RNA editing in specific cell types are sparse. Researchers have now developed a method to visualize adenosine-to-inosine editing activity in individual fruit fly neurons using a reverse-engineered fluorescent reporter.
- Chammiran Daniel
- & Marie Öhman
-
Article |
 Visualizing adenosine-to-inosine RNA editing in the Drosophila nervous system
Visualizing adenosine-to-inosine RNA editing in the Drosophila nervous system
Adenosine-to-inosine RNA editing modifies expressed sequences and enhances functional protein diversity. The authors report an in vivo fluorescent reporter that provides a readout of adenosine deaminase RNA-editing activity in Drosophila melanogaster neurons, showing evidence of inter-individual variability in editing activity.
- James E C Jepson
- , Yiannis A Savva
- & Robert A Reenan
-
Article |
 Analysis and design of RNA sequencing experiments for identifying isoform regulation
Analysis and design of RNA sequencing experiments for identifying isoform regulation
The mixture of isoforms model (MISO) assesses the confidence in estimates of the abundance of spliced exons or isoforms from paired-end RNA-seq data and detects their differential expression.
- Yarden Katz
- , Eric T Wang
- & Christopher B Burge
-
News & Views |
 Simplifying complexity
Simplifying complexity
A software tool based on gene model profiling improves analysis of alternative splice events in RNA sequencing (RNA-seq) data.
- Nicole Cloonan
- & Sean M Grimmond
-
Research Highlights |
Searching for mismatches in a vast genomic landscape
Raw data of millions of sequences used to assemble the reference genomes of ten organisms are analyzed in search of mismatches indicative of editing events. Findings include candidate sites for in vivo DNA and RNA editing, and a common sequencing error.
- Erika Pastrana
-
Article |
 Conditional gene expression and RNAi using MEC-8–dependent splicing in C. elegans
Conditional gene expression and RNAi using MEC-8–dependent splicing in C. elegans
A conditional gene expression system in Caenorhabditis elegans is reported. It should permit the generation of temperature-sensitive alleles for most genes.
- Andrea Calixto
- , Charles Ma
- & Martin Chalfie
