Brief Communication
|
Open Access
Featured
-
-
Article
| Open Access Spatial landmark detection and tissue registration with deep learning
Spatial landmark detection and tissue registration with deep learning
Effortless landmark detection is an unsupervised deep learning-based approach that addresses key challenges in landmark detection and image registration for accurate performance across diverse tissue imaging datasets.
- Markus Ekvall
- , Ludvig Bergenstråhle
- & Joakim Lundeberg
-
Article |
 scGPT: toward building a foundation model for single-cell multi-omics using generative AI
scGPT: toward building a foundation model for single-cell multi-omics using generative AI
Pretrained using over 33 million single-cell RNA-sequencing profiles, scGPT is a foundation model facilitating a broad spectrum of downstream single-cell analysis tasks by transfer learning.
- Haotian Cui
- , Chloe Wang
- & Bo Wang
-
Perspective |
 Challenges and perspectives in computational deconvolution of genomics data
Challenges and perspectives in computational deconvolution of genomics data
This Perspective provides an overview of major challenges and associated recommendations in computational deconvolution for genomics data.
- Lana X. Garmire
- , Yijun Li
- & Andrew E. Teschendorff
-
Article
| Open Access Toward universal cell embeddings: integrating single-cell RNA-seq datasets across species with SATURN
Toward universal cell embeddings: integrating single-cell RNA-seq datasets across species with SATURN
SATURN performs cross-species integration and analysis using both single-cell gene expression and protein representations generated by protein language models.
- Yanay Rosen
- , Maria Brbić
- & Jure Leskovec
-
Article |
 TISSUE: uncertainty-calibrated prediction of single-cell spatial transcriptomics improves downstream analyses
TISSUE: uncertainty-calibrated prediction of single-cell spatial transcriptomics improves downstream analyses
Transcript Imputation with Spatial Single-cell Uncertainty Estimation (TISSUE) offers a general framework for estimating uncertainty for spatial gene expression predictions, enabling improved downstream analysis of spatially resolved transcriptomics data.
- Eric D. Sun
- , Rong Ma
- & James Zou
-
Brief Communication
| Open Access Correcting PCR amplification errors in unique molecular identifiers to generate accurate numbers of sequencing molecules
Correcting PCR amplification errors in unique molecular identifiers to generate accurate numbers of sequencing molecules
This study introduces a method utilizing homotrimeric nucleotide blocks to achieve accurate counts of RNA molecules in both bulk and single-cell sequencing data.
- Jianfeng Sun
- , Martin Philpott
- & Adam P. Cribbs
-
Resource |
 scPerturb: harmonized single-cell perturbation data
scPerturb: harmonized single-cell perturbation data
scPerturb is an information resource for single-cell perturbation data analysis and comparison.
- Stefan Peidli
- , Tessa D. Green
- & Chris Sander
-
Method to Watch |
 Spatially resolved multiomics
Spatially resolved multiomics
Spatially resolved multimodal omics offers a collective way to capture molecular information in complex tissues.
- Lei Tang
-
-
Article
| Open Access Population-level integration of single-cell datasets enables multi-scale analysis across samples
Population-level integration of single-cell datasets enables multi-scale analysis across samples
By learning representations for both cells and various condition covariates, scPoli facilitates atlas-level integration and analysis of single-cell genomics datasets with improved interpretability.
- Carlo De Donno
- , Soroor Hediyeh-Zadeh
- & Fabian J. Theis
-
This Month |
 Chlamydomonas reinhardtii: a model for photosynthesis and so much more
Chlamydomonas reinhardtii: a model for photosynthesis and so much more
The green alga Chlamydomonas reinhardtii is a useful reference organism for studying photosynthesis, cilia and the cell cycle. Like many other algae, it exhibits daily rhythms in gene expression and behavior that are in sync with the rising and setting of the sun.
- Sunnyjoy Dupuis
- & Sabeeha S. Merchant
-
Research Briefing |
 Evaluating long-read RNA-sequencing analysis tools with in silico mixtures
Evaluating long-read RNA-sequencing analysis tools with in silico mixtures
We conducted a comprehensive long-read RNA sequencing (RNA-seq) benchmarking experiment by combining spike-ins and in silico mixtures to establish a ground-truth dataset. We used long- and short-read RNA-seq technology to deeply sequence samples and compared the performance of a range of analysis tools on these data.
-
Analysis |
 Benchmarking long-read RNA-sequencing analysis tools using in silico mixtures
Benchmarking long-read RNA-sequencing analysis tools using in silico mixtures
This analysis leverages experimentally sequenced data and in silico mixtures to simulate transcript expression differences, which enables a performance assessment of long-read tools developed for isoform detection, differential transcript expression analysis and differential transcript usage analysis.
- Xueyi Dong
- , Mei R. M. Du
- & Matthew E. Ritchie
-
Article
| Open Access A new Bayesian factor analysis method improves detection of genes and biological processes affected by perturbations in single-cell CRISPR screening
A new Bayesian factor analysis method improves detection of genes and biological processes affected by perturbations in single-cell CRISPR screening
Guided sparse factor analysis (GSFA) is a powerful statistical framework to detect changes in gene expression as a result of perturbations in single-cell CRISPR screening.
- Yifan Zhou
- , Kaixuan Luo
- & Xin He
-
Article
| Open Access Deep generative modeling of transcriptional dynamics for RNA velocity analysis in single cells
Deep generative modeling of transcriptional dynamics for RNA velocity analysis in single cells
veloVI enhances RNA velocity analysis with uncertainty quantification and extensibility by deep generative modeling of gene-specific transcriptional dynamics.
- Adam Gayoso
- , Philipp Weiler
- & Nir Yosef
-
Article
| Open Access Spatiotemporal, optogenetic control of gene expression in organoids
Spatiotemporal, optogenetic control of gene expression in organoids
A workflow combining optogenetic perturbations with spatial transcriptomics to program spatiotemporal gene expression patterns in organoids.
- Ivano Legnini
- , Lisa Emmenegger
- & Nikolaus Rajewsky
-
Correspondence |
 Extending support for mouse data in the Molecular Signatures Database (MSigDB)
Extending support for mouse data in the Molecular Signatures Database (MSigDB)
- Anthony S. Castanza
- , Jill M. Recla
- & Jill P. Mesirov
-
Article |
 Recovery of missing single-cell RNA-sequencing data with optimized transcriptomic references
Recovery of missing single-cell RNA-sequencing data with optimized transcriptomic references
This paper presents an improved approach for mapping single-cell RNA-seq reads with optimized transcriptomic references, which markedly recovers previously missing gene expression data.
- Allan-Hermann Pool
- , Helen Poldsam
- & Yuki Oka
-
Article
| Open Access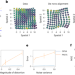 Alignment of spatial genomics data using deep Gaussian processes
Alignment of spatial genomics data using deep Gaussian processes
Gaussian Process Spatial Alignment (GPSA) aligns multiple spatially resolved genomics and histology datasets and improves downstream analysis.
- Andrew Jones
- , F. William Townes
- & Barbara E. Engelhardt
-
News & Views |
 Dissecting gene regulation with multimodal sequencing
Dissecting gene regulation with multimodal sequencing
Recently proposed computational approaches explore casual links between chromatin and transcriptional changes that are provided by single-cell multimodal sequencing to bridge the knowledge gap in transcriptional regulatory control.
- Ivan G. Costa
-
Article |
 Dictys: dynamic gene regulatory network dissects developmental continuum with single-cell multiomics
Dictys: dynamic gene regulatory network dissects developmental continuum with single-cell multiomics
By probabilistic modeling of gene regulation and expression kinetics, Dictys infers dynamic and context-specific gene regulatory networks using single-cell multiomics data.
- Lingfei Wang
- , Nikolaos Trasanidis
- & Luca Pinello
-
Research Highlight |
 Ex utero development of primate embryos
Ex utero development of primate embryos
Two independent studies demonstrate in vitro development of early cynomolgus monkey embryos.
- Madhura Mukhopadhyay
-
Article |
 Significance analysis for clustering with single-cell RNA-sequencing data
Significance analysis for clustering with single-cell RNA-sequencing data
This study presents a significance analysis framework for evaluating single-cell clusters. Application of the method detects cases of over-clustering in reported single-cell RNA-sequencing analysis results.
- Isabella N. Grabski
- , Kelly Street
- & Rafael A. Irizarry
-
Article |
 SCS: cell segmentation for high-resolution spatial transcriptomics
SCS: cell segmentation for high-resolution spatial transcriptomics
Subcellular spatial transcriptomics cell segmentation (SCS) combines information from stained images and sequencing data to improve cell segmentation in high-resolution spatial transcriptomics data.
- Hao Chen
- , Dongshunyi Li
- & Ziv Bar-Joseph
-
Article
| Open Access MultiVI: deep generative model for the integration of multimodal data
MultiVI: deep generative model for the integration of multimodal data
By learning a joint representation using deep generative modeling, MultiVI integrates multimodal and single-modality single-cell datasets, which enhances multiple functionalities.
- Tal Ashuach
- , Mariano I. Gabitto
- & Nir Yosef
-
Article |
 Context-aware transcript quantification from long-read RNA-seq data with Bambu
Context-aware transcript quantification from long-read RNA-seq data with Bambu
Leveraging long-read RNA-seq data and machine learning, Bambu facilitates accurate transcript discovery and quantification.
- Ying Chen
- , Andre Sim
- & Jonathan Göke
-
-
-
Article
| Open Access Nano-DMS-MaP allows isoform-specific RNA structure determination
Nano-DMS-MaP allows isoform-specific RNA structure determination
Nano-DMS-MaP combines the power of DMS mutational profiling and long-read nanopore sequencing to resolve structural differences among RNA isoforms, revealing the structural landscape of HIV-1 transcripts in cells.
- Patrick Bohn
- , Anne-Sophie Gribling-Burrer
- & Redmond P. Smyth
-
Article
| Open Access Spatiotemporally resolved transcriptomics reveals the subcellular RNA kinetic landscape
Spatiotemporally resolved transcriptomics reveals the subcellular RNA kinetic landscape
TEMPOmap combines pulse-chase metabolic labeling with multiplexed three-dimensional in situ sequencing to simultaneously profile the age and subcellular location of individual RNA molecules from thousands of genes to reveal RNA kinetic landscapes.
- Jingyi Ren
- , Haowen Zhou
- & Xiao Wang
-
Analysis
| Open Access Comparison of transformations for single-cell RNA-seq data
Comparison of transformations for single-cell RNA-seq data
This paper compares different transformation approaches for analysis of single-cell RNA-sequencing data and provides recommendations for method selection.
- Constantin Ahlmann-Eltze
- & Wolfgang Huber
-
News & Views |
 Designing spatial transcriptomic experiments
Designing spatial transcriptomic experiments
Optimal design of spatial transcriptomic experiments allows statistical evaluation of the impact of various biological and technological features on the discovery of cell phenotypes.
- Dario Righelli
- , Andrea Sottosanti
- & Davide Risso
-
Article
| Open Access In silico tissue generation and power analysis for spatial omics
In silico tissue generation and power analysis for spatial omics
This paper presents a statistical framework for power analysis of spatial omics studies, facilitated by an in silico tissue-generation method.
- Ethan A. G. Baker
- , Denis Schapiro
- & Aviv Regev
-
Article |
 Combining long-term circuit mapping and network transcriptomics with SiR-N2c
Combining long-term circuit mapping and network transcriptomics with SiR-N2c
A self-inactivating variant of the CVS-N2c rabies virus enables both retrograde viral tracing and transcriptomic analyses, thereby allowing a combination of circuit mapping and molecular studies.
- Hassal Lee
- , Ernesto Ciabatti
- & Marco Tripodi
-
Article
| Open Access Screening cell–cell communication in spatial transcriptomics via collective optimal transport
Screening cell–cell communication in spatial transcriptomics via collective optimal transport
This work presents a computational framework, COMMOT, to spatially infer cell–cell communication from transcriptomics data based on a variant of optimal transport (OT).
- Zixuan Cang
- , Yanxiang Zhao
- & Qing Nie
-
Editorial |
Method of the Year 2022: long-read sequencing
Long-read sequencing powers a more complete reading of genomic information.
-
Comment |
 Long-read sequencing in the era of epigenomics and epitranscriptomics
Long-read sequencing in the era of epigenomics and epitranscriptomics
As long-read sequencing technologies continue to advance, the possibility of obtaining maps of DNA and RNA modifications at single-molecule resolution has become a reality. Here we highlight the opportunities and challenges posed by the use of long-read sequencing technologies to study epigenetic and epitranscriptomic marks and how this will affect the way in which we approach the study of health and disease states.
- Morghan C. Lucas
- & Eva Maria Novoa
-
Research Briefing |
 Parts-based decomposition of spatial genomics data finds distinct tissue regions
Parts-based decomposition of spatial genomics data finds distinct tissue regions
Dimension reduction is a cornerstone of exploratory data analysis; however, traditional methods fail to preserve the spatial context of spatial genomics data. In this work, we develop a nonnegative spatial factorization (NSF) model that allows interpretable, parts-based decomposition of spatial single-cell count data. NSF allows label-free annotation of regions of interest in spatial genomics data and identifies genes and cells that can be used to define those regions.
-
Article
| Open Access Nonnegative spatial factorization applied to spatial genomics
Nonnegative spatial factorization applied to spatial genomics
This paper presents nonnegative spatial factorization, a general framework for spatially aware and interpretable dimension reduction for high-dimensional spatial data, and its application to spatial transcriptomics analysis.
- F. William Townes
- & Barbara E. Engelhardt
-
Brief Communication
| Open Access Multiplexed transcriptome discovery of RNA-binding protein binding sites by antibody-barcode eCLIP
Multiplexed transcriptome discovery of RNA-binding protein binding sites by antibody-barcode eCLIP
Antibody-barcode eCLIP (ABC) uses proximity ligation to couple DNA-barcoded antibodies to RNA-binding protein (RBP)-protected RNA fragments for multiplexed eCLIP. ABC can be used to interrogate several RBPs in a single tube with results on par with eCLIP.
- Daniel A. Lorenz
- , Hsuan-Lin Her
- & Gene W. Yeo
-
Article
| Open Access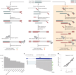 Nano3P-seq: transcriptome-wide analysis of gene expression and tail dynamics using end-capture nanopore cDNA sequencing
Nano3P-seq: transcriptome-wide analysis of gene expression and tail dynamics using end-capture nanopore cDNA sequencing
Nano3P-seq presents a nanopore-based sequencing tool to profile polyA-tailed and non-polyA-tailed transcripts, as well as capture polyA tail length and composition.
- Oguzhan Begik
- , Gregor Diensthuber
- & Eva Maria Novoa
-
Article
| Open Access Detection of m6A from direct RNA sequencing using a multiple instance learning framework
Detection of m6A from direct RNA sequencing using a multiple instance learning framework
This work presents m6Anet, which implements a neural-network-based multiple instance learning model to detect m6A modifications from direct RNA sequencing data.
- Christopher Hendra
- , Ploy N. Pratanwanich
- & Jonathan Göke
-
Research Highlight |
 Imaging-based spatial transcriptomics goes electric
Imaging-based spatial transcriptomics goes electric
Researchers use electric fields to transfer RNA from a tissue sample onto a surface for subsequent fluorescence in situ hybridization-based profiling of transcriptomes at the single-cell level.
- Rita Strack
-
Article |
 Annotation of spatially resolved single-cell data with STELLAR
Annotation of spatially resolved single-cell data with STELLAR
STELLAR (spatial cell learning) is a geometric deep learning model that works with spatially resolved single-cell datasets to both assign cell types in unannotated datasets based on a reference dataset and discover new cell types.
- Maria Brbić
- , Kaidi Cao
- & Jure Leskovec
-
Article
| Open Access Light-Seq: light-directed in situ barcoding of biomolecules in fixed cells and tissues for spatially indexed sequencing
Light-Seq: light-directed in situ barcoding of biomolecules in fixed cells and tissues for spatially indexed sequencing
Light-Seq uses light-directed DNA barcoding in fixed cells and tissues for multiplexed spatial indexing and subsequent next generation sequencing. This approach blends spatial and omics information to enable analysis of rare cell types in complex tissues.
- Jocelyn Y. Kishi
- , Ninning Liu
- & Peng Yin
-
Editorial |
Decoding noncoding RNAs
Research interest in noncoding RNAs and their biological implications in a variety of cellular contexts has been growing. In this issue, we present a series of pieces discussing recent method advances and future directions for deciphering the regulatory roles of noncoding RNAs.
-
-
Research Briefing |
 A statistical method to uncover gene expression changes in spatial transcriptomics
A statistical method to uncover gene expression changes in spatial transcriptomics
Cell type-specific inference of differential expression (C-SIDE) is a statistical model that identifies which genes (within a determined cell type) are differentially expressed on the basis of spatial position, pathological changes or cell–cell interactions. C-SIDE facilitates differential expression analysis in spatial transcriptomics by jointly modeling cell type mixtures and spatially varying gene expression.
-
Research Highlight |
Towards a full picture of the total transcriptome
VASA-seq offers a single cell sequencing tool for capturing full-length coverage of coding sequences and supplementing non-coding RNA molecules.
- Lei Tang

 Assessing GPT-4 for cell type annotation in single-cell RNA-seq analysis
Assessing GPT-4 for cell type annotation in single-cell RNA-seq analysis Modeling morphogenesis
Modeling morphogenesis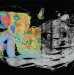 A new member of the spatial omics family
A new member of the spatial omics family