Featured
-
-
Perspective |
 Can AlphaFold’s breakthrough in protein structure help decode the fundamental principles of adaptive cellular immunity?
Can AlphaFold’s breakthrough in protein structure help decode the fundamental principles of adaptive cellular immunity?
This Perspective discusses the potential of protein structure-prediction models for exploring the structural landscape and specificity of TCR–pMHC interactions.
- Benjamin McMaster
- , Christopher Thorpe
- & Hashem Koohy
-
Editorial |
AlphaFold and beyond
A great deal has happened in the protein structure prediction field since Nature Methods selected this topic as our Method of the Year 2021. Here’s a quick, non-comprehensive update.
-
-
Article
| Open Access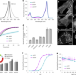 Chemically stable fluorescent proteins for advanced microscopy
Chemically stable fluorescent proteins for advanced microscopy
The engineered hyperfolder YFP (hfYFP) and variants offer unprecedented chemical and thermal stability, making them versatile probes for microscopy as well as for challenging applications like correlative light and electron microscopy and expansion microscopy.
- Benjamin C. Campbell
- , Maria G. Paez-Segala
- & Ce Feng Liu
-
Editorial |
Method of the Year 2021: Protein structure prediction
Deep Learning based approaches for protein structure prediction have sent shock waves through the structural biology community. We anticipate far-reaching and long-lasting impact.
-
Technology Feature |
 Method of the Year: protein structure prediction
Method of the Year: protein structure prediction
Nature Methods has named protein structure prediction the Method of the Year 2021.
- Vivien Marx
-
Research Highlight |
GFP lights up amyloid fibrils
Fluorescent proteins can bind amyloid fibrils formed from natural peptides and proteins.
- Rita Strack
-
-
Research Highlight |
 Fluorescent proteins from scratch
Fluorescent proteins from scratch
Researchers have carried out the first de novo design of a β-barrel protein and engineered the interior to bind and activate a fluorogenic dye.
- Rita Strack
-
Resource |
 Systematic characterization of maturation time of fluorescent proteins in living cells
Systematic characterization of maturation time of fluorescent proteins in living cells
This work characterizes the maturation kinetics of 50 cyan to far-red fluorescent proteins and provides evidence that proteins that mature faster than their brighter but slower counterparts are more useful for quantitative evaluation of fast processes.
- Enrique Balleza
- , J Mark Kim
- & Philippe Cluzel
-
Research Highlights |
 Protein folding studies go global
Protein folding studies go global
A large-scale approach to analyze how protein sequence determines folding provides new insights into an old question.
- Allison Doerr
-
Research Highlights |
Watching proteins fold on the ribosome
Researchers monitor cotranslational protein folding in real time.
- Rita Strack
-
Research Highlights |
STOMPing at the bits
Spatially targeted optical microproteomics identifies novel amyloid plaque components.
- Stéphane Larochelle
-
Article |
 Mutational interference mapping experiment (MIME) for studying RNA structure and function
Mutational interference mapping experiment (MIME) for studying RNA structure and function
Functionally important residues in a long RNA can be identified by mutational interference mapping experiment (MIME), a method which uses random mutagenesis of RNA followed by selection for function and high-throughput sequencing.
- Redmond P Smyth
- , Laurence Despons
- & Roland Marquet
-
Article |
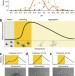 ProteoPlex: stability optimization of macromolecular complexes by sparse-matrix screening of chemical space
ProteoPlex: stability optimization of macromolecular complexes by sparse-matrix screening of chemical space
ProteoPlex optimizes buffer conditions for the isolation and purification of macromolecular complexes. The concurrent complex stabilization is beneficial for structure determination using X-ray crystallography or cryo-electron microscopy.
- Ashwin Chari
- , David Haselbach
- & Holger Stark
-
Methods in Brief |
Fast reaction kinetics with time-resolved mass spectrometry
-
Method to Watch |
 Nanopores for proteins
Nanopores for proteins
Nanopores hold promise for single-protein characterization.
- Tal Nawy
-
Methods in Brief |
Four-color FRET follows Hsp90 states
-
Brief Communication |
 Tracking protein aggregation and mislocalization in cells with flow cytometry
Tracking protein aggregation and mislocalization in cells with flow cytometry
Protein localization changes in cells are monitored at high-throughput applying pulse-shape analysis to flow-cytometry data. The authors use the technique in combination with tetracysteine-based oligomer sensors to monitor toxic protein aggregation in a cellular model of Huntington's disease.
- Yasmin M Ramdzan
- , Saskia Polling
- & Danny M Hatters
-
Brief Communication |
 Visualizing a one-way protein encounter complex by ultrafast single-molecule mixing
Visualizing a one-way protein encounter complex by ultrafast single-molecule mixing
A laminar flow mixing microfluidic device enables single-molecule fluorescence resonance energy transfer (FRET) kinetic measurements with a time resolution of 0.2 ms, enabling the study of early binding-coupled folding and unfolding events of an intrinsically disordered protein, α-synuclein. Also in this issue, Kim et al. describe another microfluidic mixing device for single-molecule experiments.
- Yann Gambin
- , Virginia VanDelinder
- & Ashok A Deniz
-
Research Highlights |
News in brief
-
Research Highlights |
Following the fold
A specialized supercomputer allows molecular dynamics simulations to be carried out for much longer periods of time than previously possible, yielding new insights into protein folding and dynamics.
- Allison Doerr
-
Brief Communication |
 Estimating prion concentration in fluids and tissues by quantitative PMCA
Estimating prion concentration in fluids and tissues by quantitative PMCA
The misfolded form of the prion protein, PrPSc, can be quantified in a variety of tissues and fluids using a quantitative version of the popular protein misfolding cyclic amplification (PMCA) assay.
- Baian Chen
- , Rodrigo Morales
- & Claudio Soto
-
News & Views |
 Waltz, an exciting new move in amyloid prediction
Waltz, an exciting new move in amyloid prediction
A new amyloid-prediction tool, Waltz, offers advantages over previous amyloid-prediction tools for distinguishing 'true' amyloids from amorphous aggregates.
- Mikael Oliveberg
-
Article |
 Protein folding stability and dynamics imaged in a living cell
Protein folding stability and dynamics imaged in a living cell
Protein dynamics can be studied in single living cells by time-resolved fluorescent imaging of the unfolding of a fluorescence resonance energy transfer (FRET) probe—labeled protein as fast temperature jumps are applied.
- Simon Ebbinghaus
- , Apratim Dhar
- & Martin Gruebele
-
Article |
 Exploring the sequence determinants of amyloid structure using position-specific scoring matrices
Exploring the sequence determinants of amyloid structure using position-specific scoring matrices
Waltz is a position-specific amyloid-propensity prediction tool developed to distinguish between true amyloid sequences and amorphous β-aggregates.
- Sebastian Maurer-Stroh
- , Maja Debulpaep
- & Frederic Rousseau

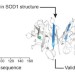
 OpenFold: retraining AlphaFold2 yields new insights into its learning mechanisms and capacity for generalization
OpenFold: retraining AlphaFold2 yields new insights into its learning mechanisms and capacity for generalization