Featured
-
-
Article
| Open Access Compact engineered human mechanosensitive transactivation modules enable potent and versatile synthetic transcriptional control
Compact engineered human mechanosensitive transactivation modules enable potent and versatile synthetic transcriptional control
Synthetic CRISPR-based transactivation domains engineered using mechanosensitive transcription factors enable robust transcriptional control.
- Barun Mahata
- , Alan Cabrera
- & Isaac B. Hilton
-
Research Highlight |
 A closer look at chromatin
A closer look at chromatin
An expansion microscopy technique called ChromExM offers detailed views into the organization chromatin and associated gene expression machinery in embryos.
- Rita Strack
-
Article
| Open Access Learning consistent subcellular landmarks to quantify changes in multiplexed protein maps
Learning consistent subcellular landmarks to quantify changes in multiplexed protein maps
CAMPA (Conditional Autoencoder for Multiplexed Pixel Analysis) learns representations of molecular pixel profiles from multiplexed images that can be clustered to quantify subcellular landmarks and capture interpretable cellular phenotypes.
- Hannah Spitzer
- , Scott Berry
- & Fabian J. Theis
-
Brief Communication
| Open Access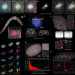 RS-FISH: precise, interactive, fast, and scalable FISH spot detection
RS-FISH: precise, interactive, fast, and scalable FISH spot detection
RS-FISH is a user-friendly software for accurate spot detection that is applicable to smFISH experiments, spatial transcriptomics, and spatial genomics. The approach enables fast spot detection in even very large volumetric datasets.
- Ella Bahry
- , Laura Breimann
- & Stephan Preibisch
-
Article |
 Persistence and plasticity in bacterial gene regulation
Persistence and plasticity in bacterial gene regulation
This work presents biotin-DNA affinity purification (DAP) sequencing, that is, an in vitro, clone-free workflow to profile transcription factor (TF) DNA binding, as well as multiDAP to simultaneously characterize TF DNA binding in multiple bacterial genomes.
- Leo A. Baumgart
- , Ji Eun Lee
- & Ronan C. O’Malley
-
Research Highlight |
 Measuring condensation forces
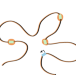
Measuring condensation forces
Researchers use an in vitro assay in combination with a polymer model to study protein–DNA condensation and how condensates bring distant DNA elements into close proximity.
- Lei Tang
-
Technology Feature |
 Tick-tock, it’s RNA o’clock
Tick-tock, it’s RNA o’clock
There’s an evolving choice of ways to track the temporal dynamic of RNA biographies.
- Vivien Marx
-
Comment |
 A new era of synchrotron-enabled macromolecular crystallography
A new era of synchrotron-enabled macromolecular crystallography
The future of macromolecular crystallography includes new X-ray sources, enhanced remote-accessible capabilities and time-resolved methods to capture intermediate structures along reaction pathways.
- Aina E. Cohen
-
Brief Communication |
 An efficient KRAB domain for CRISPRi applications in human cells
An efficient KRAB domain for CRISPRi applications in human cells
CRISPRi activities are measured from dCas9 fusions with 57 KRAB domains, which lead to the identification of ZIM3 KRAB–dCas9 fusion as a superior repressor.
- Nader Alerasool
- , Dmitri Segal
- & Mikko Taipale
-
Perspective |
 Mechanistic modeling of chromatin folding to understand function
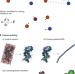
Mechanistic modeling of chromatin folding to understand function
This Perspective highlights recently developed computational models for studying chromosome organization, with a focus on how mechanistic modeling helps biologists to interpret the biological function behind the genome structures.
- Chris A. Brackey
- , Davide Marenduzzo
- & Nick Gilbert
-
Article |
 FLAM-seq: full-length mRNA sequencing reveals principles of poly(A) tail length control
FLAM-seq: full-length mRNA sequencing reveals principles of poly(A) tail length control
FLAM-seq implements a cDNA library preparation followed by single-molecule sequencing, for determining full-length mRNA molecules, including poly(A) tails.
- Ivano Legnini
- , Jonathan Alles
- & Nikolaus Rajewsky
-
Research Highlight |
Recording transcriptional activity
Acquisition of RNA into CRISPR arrays allows the recording of transcriptional dynamics in cells.
- Nicole Rusk
-
Tools in Brief |
Big Papi uncovers genetic interactions
-
Article |
 Resolving systematic errors in widely used enhancer activity assays in human cells
Resolving systematic errors in widely used enhancer activity assays in human cells
Using the ORI of plasmids used in enhancer assays as the sole core promoter and inhibiting the interferon I response triggered by plasmid transfection greatly reduces false positive and negative results in single-candidate and massively parallel enhancer assays and enables genome-wide enhancer screens.
- Felix Muerdter
- , Łukasz M Boryń
- & Alexander Stark
-
Brief Communication |
 Inducible and multiplex gene regulation using CRISPR–Cpf1-based transcription factors
Inducible and multiplex gene regulation using CRISPR–Cpf1-based transcription factors
Inducible dCpf1-based transcriptional activators achieve tunable, multiplexed target gene expression.
- Y Esther Tak
- , Benjamin P Kleinstiver
- & J Keith Joung
-
Brief Communication |
 CRISPR-STOP: gene silencing through base-editing-induced nonsense mutations
CRISPR-STOP: gene silencing through base-editing-induced nonsense mutations
Early STOP codons created with CRISPR base editors leads to gene knockout with high efficiency and does not stress cells with double-strand DNA breaks. CRISPR-STOP can target the majority of human genes and is useful for genetic screens.
- Cem Kuscu
- , Mahmut Parlak
- & Mazhar Adli
-
Methods in Brief |
Improved 3D single-molecule imaging
-
Tools in Brief |
CRiNCL unveils noncoding RNA functions
-
Methods in Brief |
A functional assay for promoters
-
Methods in Brief |
Are super-enhancers really super?
-
Research Highlights |
 Spatial transcriptomics
Spatial transcriptomics
By capturing and barcoding RNA in its native tissue location, researchers can visualize and quantify gene expression in situ.
- Nicole Rusk
-
Research Highlights |
The human transient transcriptome
Fragmenting transcripts after 4-thiouridine incorporation allows the quantification of even short-lived noncoding RNA.
- Nicole Rusk
-
Research Highlights |
Capturing transcription factors in the wild
An assay that snares transcription factors with genomic DNA fragments reveals how genetic and epigenetic factors shape binding behavior.
- Michael Eisenstein
-
Analysis |
 Comparison of Cas9 activators in multiple species
Comparison of Cas9 activators in multiple species
A comparison of seven dCas9-based transcriptional activators shows that VPR, SAM, and Suntag perform best in cell lines from a variety of organisms.
- Alejandro Chavez
- , Marcelle Tuttle
- & George Church
-
Methods in Brief |
Multiplexed single-cell protein and RNA measurements
-
Brief Communication |
 Characterizing transcriptional heterogeneity through pathway and gene set overdispersion analysis
Characterizing transcriptional heterogeneity through pathway and gene set overdispersion analysis
By testing potential sources of biological signal that drive population-level variation in single-cell gene expression, the PAGODA software enables cells to be characterized on the basis of multiple functionally relevant features such as cell type, signaling state and cell cycle state.
- Jean Fan
- , Neeraj Salathia
- & Peter V Kharchenko
-
News & Views |
 Removing roadblocks to deep sequencing of modified RNAs
Removing roadblocks to deep sequencing of modified RNAs
By removing modified nucleotides that block reverse transcriptase, two methods have now made tRNAs amenable to RNA-seq.
- Jeremy E Wilusz
-
Brief Communication |
 Efficient and quantitative high-throughput tRNA sequencing
Efficient and quantitative high-throughput tRNA sequencing
The combination of an engineered demthylase and a highly processive reverse transcriptase during tRNA library preparation for high-throughput sequencing allows comprehensive profiling of the small RNAs.
- Guanqun Zheng
- , Yidan Qin
- & Tao Pan
-
Research Highlights |
 Catching Pol II in the act
Catching Pol II in the act
Two methods capture the pausing behavior of RNA polymerase II as it generates messages from mammalian genomes.
- Tal Nawy
-
Brief Communication |
 Combining protein and mRNA quantification to decipher transcriptional regulation
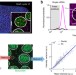
Combining protein and mRNA quantification to decipher transcriptional regulation
The combination of immunofluorescence and single-molecule FISH (smFISH) enables a quantitative description of how transcription factor binding drives the kinetics of mRNA production.
- Heng Xu
- , Leonardo A Sepúlveda
- & Ido Golding
-
Research Highlights |
Splice and sequence
A ligation-based sequencing method reveals distant exon connections within long RNA molecules.
- Tal Nawy
-
Research Highlights |
RNA that activates transcription
Synthetic small RNA transcriptional activators can regulate gene transcription in Escherichia coli.
- Nicole Rusk
-
Brief Communication |
 Improved and expanded Q-system reagents for genetic manipulations
Improved and expanded Q-system reagents for genetic manipulations
Binary expression systems such as the Gal4-UAS or Q-systems are useful tools for genetic manipulation in Drosophila. Here, improved reagents for the Q-system are described.
- Olena Riabinina
- , David Luginbuhl
- & Christopher J Potter
-
Methods in Brief |
Following transcription in real time
-
Research Highlights |
FISHing for faster findings
Quick-hybridizing probes help scientists image the high-speed events leading up to gene transcription.
- Michael Eisenstein
-
Article |
 FIREWACh: high-throughput functional detection of transcriptional regulatory modules in mammalian cells
FIREWACh: high-throughput functional detection of transcriptional regulatory modules in mammalian cells
Screening of cells transduced with a lentivirus library harboring nucleosome-free regions of mammalian genomes identifies cis elements that regulate transcription.
- Matthew Murtha
- , Zeynep Tokcaer-Keskin
- & Lisa Dailey
-
Tools in Brief |
Optimized optogenetic gene expression
-
Research Highlights |
Orthogonal logic gates
A set of 16 TetR proteins specific for their synthetic promoters will allow the construction of complex genetic circuits.
- Nicole Rusk
-
Article |
 High-resolution mapping of transcription factor binding sites on native chromatin
High-resolution mapping of transcription factor binding sites on native chromatin
A method to map transcription factor binding across the genome, at high resolution and without cross-linking.
- Sivakanthan Kasinathan
- , Guillermo A Orsi
- & Steven Henikoff
-
Article |
 Orthogonal Cas9 proteins for RNA-guided gene regulation and editing
Orthogonal Cas9 proteins for RNA-guided gene regulation and editing
A set of Cas9 endonucleases orthogonal to the Streptococcus pyogenes enzyme is identified. This will enable simultaneous addressing of multiple RNA-guided activities to different genomic target sites with the CRISPR-Cas9 system.
- Kevin M Esvelt
- , Prashant Mali
- & George M Church
-
Research Highlights |
The influence of a host
Dissecting the effect of a host's genetic background on circuit performance will allow better design.
- Nicole Rusk
-
Brief Communication |
 CRISPR RNA–guided activation of endogenous human genes
CRISPR RNA–guided activation of endogenous human genes
Synthetic transcription factors based on the RNA-guided CRISPR-Cas9 system are used to activate endogenous genes in human cells. Also online, Gersbach and colleagues report similar developments at multiple other loci.
- Morgan L Maeder
- , Samantha J Linder
- & J Keith Joung
-
Brief Communication |
 RNA-guided gene activation by CRISPR-Cas9–based transcription factors
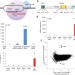
RNA-guided gene activation by CRISPR-Cas9–based transcription factors
Synthetic transcription factors based on the RNA-guided CRISPR-Cas9 system are used to activate specific endogenous genes in human cells. Also online, Joung and colleagues report similar developments at two other loci.
- Pablo Perez-Pinera
- , D Dewran Kocak
- & Charles A Gersbach
-
Article |
 TALE-mediated modulation of transcriptional enhancers in vivo
TALE-mediated modulation of transcriptional enhancers in vivo
Transcription activator–like effectors are used for in vivo activation and repression of endogenous promoters and enhancers in the fruit fly.
- Justin Crocker
- & David L Stern
-
Research Highlights |
 Complex logic in a single layer
Complex logic in a single layer
Two independently controlled regulators modulate the output in amplifying Boolean logic gates.
- Nicole Rusk
-
Brief Communication |
 Robust, synergistic regulation of human gene expression using TALE activators
Robust, synergistic regulation of human gene expression using TALE activators
TAL effector (TALE)–based transcription factors robustly induce transcription when targeted to DNase I hypersensitive sites in a gene promoter in human cells, even more so when several TALE transcription factors work in synergy at the same promoter. Tunable expression of endogenous genes will be of great interest in basic research and in synthetic biology.
- Morgan L Maeder
- , Samantha J Linder
- & J Keith Joung
-
Tools in Brief |
Programming transcription
-
Brief Communication |
 Single-allele analysis of transcription kinetics in living mammalian cells
Single-allele analysis of transcription kinetics in living mammalian cells
Single integration of a target gene expressing MS2-binding stem loops allows the real time quantification of transcriptional bursts, promoter firings and cell cycle–dependent transcription rates.
- Sharon Yunger
- , Liat Rosenfeld
- & Yaron Shav-Tal
-
News & Views |
 Illuminating eukaryotic transcription start sites
Illuminating eukaryotic transcription start sites
Simplified methods to map transcription start sites allow the exploration of transcription-initiation landscapes in rare cell types.
- John A Stamatoyannopoulos

 Combining compact human protein domains with CRISPR systems for robust gene activation
Combining compact human protein domains with CRISPR systems for robust gene activation