Research Briefing |
Featured
-
-
Brief Communication
| Open Access Multiplexed transcriptome discovery of RNA-binding protein binding sites by antibody-barcode eCLIP
Multiplexed transcriptome discovery of RNA-binding protein binding sites by antibody-barcode eCLIP
Antibody-barcode eCLIP (ABC) uses proximity ligation to couple DNA-barcoded antibodies to RNA-binding protein (RBP)-protected RNA fragments for multiplexed eCLIP. ABC can be used to interrogate several RBPs in a single tube with results on par with eCLIP.
- Daniel A. Lorenz
- , Hsuan-Lin Her
- & Gene W. Yeo
-
Article
| Open Access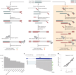 Nano3P-seq: transcriptome-wide analysis of gene expression and tail dynamics using end-capture nanopore cDNA sequencing
Nano3P-seq: transcriptome-wide analysis of gene expression and tail dynamics using end-capture nanopore cDNA sequencing
Nano3P-seq presents a nanopore-based sequencing tool to profile polyA-tailed and non-polyA-tailed transcripts, as well as capture polyA tail length and composition.
- Oguzhan Begik
- , Gregor Diensthuber
- & Eva Maria Novoa
-
Comment |
 Profiling the epigenetic landscape of the antigen receptor repertoire: the missing epi-immunogenomics data
Profiling the epigenetic landscape of the antigen receptor repertoire: the missing epi-immunogenomics data
High-resolution sequencing methods that capture the epigenetic landscape within the T cell receptor (TCR) gene loci are pivotal for a fundamental understanding of the epigenetic regulatory mechanisms of the TCR repertoire. In our opinion, filling the gaps in our understanding of the epigenetic mechanisms regulating the TCR repertoire will benefit the development of strategies that can modulate the TCR repertoire composition by leveraging the dynamic nature of epigenetic modifications.
- Rayyan Aburajab
- , Mateusz Pospiech
- & Houda Alachkar
-
Brief Communication
| Open Access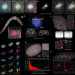 RS-FISH: precise, interactive, fast, and scalable FISH spot detection
RS-FISH: precise, interactive, fast, and scalable FISH spot detection
RS-FISH is a user-friendly software for accurate spot detection that is applicable to smFISH experiments, spatial transcriptomics, and spatial genomics. The approach enables fast spot detection in even very large volumetric datasets.
- Ella Bahry
- , Laura Breimann
- & Stephan Preibisch
-
Article
| Open Access Detection of m6A from direct RNA sequencing using a multiple instance learning framework
Detection of m6A from direct RNA sequencing using a multiple instance learning framework
This work presents m6Anet, which implements a neural-network-based multiple instance learning model to detect m6A modifications from direct RNA sequencing data.
- Christopher Hendra
- , Ploy N. Pratanwanich
- & Jonathan Göke
-
Brief Communication |
 An improved imaging system that corrects MS2-induced RNA destabilization
An improved imaging system that corrects MS2-induced RNA destabilization
An improved version of the MS2-MCP system for imaging RNA dynamics involves tethering translation termination factors to tagged mRNAs to bypass destabilization caused by NMD machinery.
- Weihan Li
- , Anna Maekiniemi
- & Robert H. Singer
-
This Month |
Athlete–scientists like to sweat
Some scientists find ways to balance their love of athleticism with research.
- Vivien Marx
-
Article |
 Annotation of spatially resolved single-cell data with STELLAR
Annotation of spatially resolved single-cell data with STELLAR
STELLAR (spatial cell learning) is a geometric deep learning model that works with spatially resolved single-cell datasets to both assign cell types in unannotated datasets based on a reference dataset and discover new cell types.
- Maria Brbić
- , Kaidi Cao
- & Jure Leskovec
-
Article
| Open Access Light-Seq: light-directed in situ barcoding of biomolecules in fixed cells and tissues for spatially indexed sequencing
Light-Seq: light-directed in situ barcoding of biomolecules in fixed cells and tissues for spatially indexed sequencing
Light-Seq uses light-directed DNA barcoding in fixed cells and tissues for multiplexed spatial indexing and subsequent next generation sequencing. This approach blends spatial and omics information to enable analysis of rare cell types in complex tissues.
- Jocelyn Y. Kishi
- , Ninning Liu
- & Peng Yin
-
Editorial |
Decoding noncoding RNAs
Research interest in noncoding RNAs and their biological implications in a variety of cellular contexts has been growing. In this issue, we present a series of pieces discussing recent method advances and future directions for deciphering the regulatory roles of noncoding RNAs.
-
Comment |
 Towards higher-resolution and in vivo understanding of lncRNA biogenesis and function
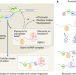
Towards higher-resolution and in vivo understanding of lncRNA biogenesis and function
Recent studies have revealed multifaceted roles of long noncoding RNAs (lncRNAs) in gene regulation, accompanying an increased understanding of lncRNA processing, localization, interacting macromolecules and structural modules. Here, progress and recently developed technological advances for understanding lncRNA biogenesis, modes of action and cellular phenotypes are highlighted, and challenges and opportunities towards higher-resolution and in vivo studies in this field are discussed.
- Ling-Ling Chen
-
Comment |
 Regulatory non-coding RNAs: everything is possible, but what is important?
Regulatory non-coding RNAs: everything is possible, but what is important?
In recent years, the number of annotated noncoding RNAs (ncRNAs) and RNA-binding proteins (RBPs) has increased dramatically. The wide range of RBPs identified highlights the enormous potential for RNA in virtually all aspects of cell biology, from transcriptional regulation to metabolic control. Yet, there is a growing gap between what is possible and what has been demonstrated to be functionally important. Here we highlight recent methodological developments in the study of RNA–protein interactions, discuss the challenges and opportunities for exploring their functional roles, and provide our perspectives on what is needed to bridge the gap in this rapidly expanding field.
- Jimmy K. Guo
- & Mitchell Guttman
-
Article |
 RNA secondary structure packages evaluated and improved by high-throughput experiments
RNA secondary structure packages evaluated and improved by high-throughput experiments
The EternaBench dataset of synthetic RNA constructs was used to directly compare RNA secondary structure prediction software packages on ensemble-oriented prediction tasks and used to train the EternaFold model for improved performance.
- Hannah K. Wayment-Steele
- , Wipapat Kladwang
- & Rhiju Das
-
Research Highlight |
Towards a full picture of the total transcriptome
VASA-seq offers a single cell sequencing tool for capturing full-length coverage of coding sequences and supplementing non-coding RNA molecules.
- Lei Tang
-
Article |
 Cyclic immonium ion of lactyllysine reveals widespread lactylation in the human proteome
Cyclic immonium ion of lactyllysine reveals widespread lactylation in the human proteome
This article reports the cyclic immonium ion as a diagnostic fragment ion for lysine lactylation. The approach was used for identifying lactylation in various enriched and unenriched proteome databases, demonstrating prevalence of lactylation beyond histones.
- Ning Wan
- , Nian Wang
- & Hui Ye
-
Research Briefing |
 A neural network for large-scale clustering of peptide mass spectra
A neural network for large-scale clustering of peptide mass spectra
Repository-scale analysis of hundreds of millions to billions of mass spectra is a challenging endeavor due to the complexity and volume of associated data. A deep neural network embedding method is presented that enables large-scale investigation of repeatedly observed yet consistently unidentified mass spectra.
-
Perspective |
 Best practice standards for circular RNA research
Best practice standards for circular RNA research
This Perspective focuses on experimental techniques developed for circRNA studies, and provides practical guidelines for performing circRNA research.
- Anne F. Nielsen
- , Albrecht Bindereif
- & Jørgen Kjems
-
News & Views |
 Mapping beads on strings
Mapping beads on strings
DiMeLo-seq leverages immunotethered DNA methyltransferases with long-read sequencing to map the locations of chromatin proteins in their natural context.
- Kami Ahmad
-
Research Briefing |
 Novel spike-ins for single-cell sequencing enable ground-truth RNA counting
Novel spike-ins for single-cell sequencing enable ground-truth RNA counting
Counting of RNA molecules using unique molecular identifiers (UMIs) is ubiquitous in single-cell sequencing. Here, we introduce molecular spikes, a new type of RNA spike-ins with in-built UMIs. These versatile molecular spikes have many uses in experimental and computational method development and routine biological applications.
-
Research Highlight |
Enhanced protein isoform characterization
Long-read proteogenomics helps delineate isoform diversity in full-length proteins.
- Arunima Singh
-
Research Highlight |
Sequencing RNA isoforms in brain tissue
Single-nuclei isoform RNA sequencing enables cell-type specific isoform analysis in frozen brain samples.
- Lei Tang
-
Article |
 DiMeLo-seq: a long-read, single-molecule method for mapping protein–DNA interactions genome wide
DiMeLo-seq: a long-read, single-molecule method for mapping protein–DNA interactions genome wide
DiMeLo-seq uses native long-read sequencing to examine protein–DNA interactions by mapping exogenous methylation marks generated by a nonspecific DNA methyltransferase, as well as profile endogenous CpG methylation simultaneously.
- Nicolas Altemose
- , Annie Maslan
- & Aaron Streets
-
Brief Communication |
 The SpliZ generalizes ‘percent spliced in’ to reveal regulated splicing at single-cell resolution
The SpliZ generalizes ‘percent spliced in’ to reveal regulated splicing at single-cell resolution
The Splicing Z Score (SpliZ) provides single-cell-level quantification of alternative splicing with improved statistical power.
- Julia Eve Olivieri
- , Roozbeh Dehghannasiri
- & Julia Salzman
-
Article |
 Global profiling of phosphorylation-dependent changes in cysteine reactivity
Global profiling of phosphorylation-dependent changes in cysteine reactivity
This article describes a chemical proteomic approach to quantitatively relate serine/threonine phosphorylation to changes in the reactivity of cysteine residues, thereby affecting their potential to be post-translationally modified and/or targeted by electrophilic small molecules.
- Esther K. Kemper
- , Yuanjin Zhang
- & Benjamin F. Cravatt
-
Method to Watch |
 Greater detail in the nucleus
Greater detail in the nucleus
Probing spatially organized DNA and its interacting elements in single cells will deepen our understanding of cell-type-specific gene regulation.
- Lei Tang
-
Research Highlight |
Recapitulating miRNA biogenesis in cells
MapToCleave enables the large-scale study of miRNA precursor processing in living cells.
- Lei Tang
-
Research Highlight |
Resolving human gastrulation
The first single-cell transcriptome of the gastrulating human embryo.
- Madhura Mukhopadhyay
-
Research Highlight |
Profiling double-strand break repair
Repair-seq enables efficient profiling of genetic factors involved in DNA double-strand break repair.
- Lin Tang
-
Article |
 Persistence and plasticity in bacterial gene regulation
Persistence and plasticity in bacterial gene regulation
This work presents biotin-DNA affinity purification (DAP) sequencing, that is, an in vitro, clone-free workflow to profile transcription factor (TF) DNA binding, as well as multiDAP to simultaneously characterize TF DNA binding in multiple bacterial genomes.
- Leo A. Baumgart
- , Ji Eun Lee
- & Ronan C. O’Malley
-
News & Views |
 Integrative analysis methods for spatial transcriptomics
Integrative analysis methods for spatial transcriptomics
Computational methods use different integrative strategies to tackle the challenges of spatially resolved transcriptomics data analysis.
- Shaina Lu
- , Daniel Fürth
- & Jesse Gillis
-
Research Highlight |
 Mapping genome structures in single cells
Mapping genome structures in single cells
Researchers develop single-cell SPRITE to detect higher-order 3D genome structures in single cells.
- Lei Tang
-
Article |
 Systematic calibration of epitranscriptomic maps using a synthetic modification-free RNA library
Systematic calibration of epitranscriptomic maps using a synthetic modification-free RNA library
This work describes the generation of a modification-free RNA library that resembles endogenous transcriptome sequence and expression level, which can be used as a negative control in epitranscriptomic sequencing methods to obtain high-confidence and quantitative maps of various RNA modifications.
- Zhang Zhang
- , Tao Chen
- & Guan-Zheng Luo
-
Research Highlight |
 Measuring condensation forces
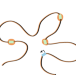
Measuring condensation forces
Researchers use an in vitro assay in combination with a polymer model to study protein–DNA condensation and how condensates bring distant DNA elements into close proximity.
- Lei Tang
-
Article |
 Nascent Ribo-Seq measures ribosomal loading time and reveals kinetic impact on ribosome density
Nascent Ribo-Seq measures ribosomal loading time and reveals kinetic impact on ribosome density
This work describes nascent Ribo-Seq, which measures the speed of ribosome loading and polysome assembly on endogenous mRNA through a combination of 4-thiouridine labeling and ribosome footprint sequencing.
- Johanna Schott
- , Sonja Reitter
- & Georg Stoecklin
-
Brief Communication
| Open Access SnapHiC: a computational pipeline to identify chromatin loops from single-cell Hi-C data
SnapHiC: a computational pipeline to identify chromatin loops from single-cell Hi-C data
SnapHiC offers a computational tool for improving detection of chromatin loops from single-cell Hi-C data.
- Miao Yu
- , Armen Abnousi
- & Ming Hu
-
Research Highlight |
 Towards resolving proteomes in single cells
Towards resolving proteomes in single cells
Development of methods for single-cell proteomics increases depth and throughput
- Arunima Singh
-
Research Highlight |
 Sequencing diverse RNA modifications
Sequencing diverse RNA modifications
Nanopore sequencing enables quantitative profiling of pseudouridine and 2′-O-methylation modifications in native RNAs.
- Lei Tang
-
Review Article |
 The triumphs and limitations of computational methods for scRNA-seq
The triumphs and limitations of computational methods for scRNA-seq
This review provides an overview of recent computational developments in scRNA-seq analysis and highlights packages and tools applied in executing these analyses.
- Peter V. Kharchenko
-
Matters Arising |
 Impact of phosphorylation on thermal stability of proteins
Impact of phosphorylation on thermal stability of proteins
- Clément M. Potel
- , Nils Kurzawa
- & Mikhail M. Savitski
-
Technology Feature |
 Tick-tock, it’s RNA o’clock
Tick-tock, it’s RNA o’clock
There’s an evolving choice of ways to track the temporal dynamic of RNA biographies.
- Vivien Marx
-
Comment |
 A new era of synchrotron-enabled macromolecular crystallography
A new era of synchrotron-enabled macromolecular crystallography
The future of macromolecular crystallography includes new X-ray sources, enhanced remote-accessible capabilities and time-resolved methods to capture intermediate structures along reaction pathways.
- Aina E. Cohen
-
Review Article |
 Understanding the invisible hands of sample preparation for cryo-EM
Understanding the invisible hands of sample preparation for cryo-EM
The quality of structural data obtained in cryo-EM is affected by multiple factors pertaining to sample preparation. This Review discusses available techniques and current challenges.
- Giulia Weissenberger
- , Rene J. M. Henderikx
- & Peter J. Peters
-
Article |
 Robust single-cell discovery of RNA targets of RNA-binding proteins and ribosomes
Robust single-cell discovery of RNA targets of RNA-binding proteins and ribosomes
Surveying targets by APOBEC-mediated profiling identifies binding sites of RBPs by C-to-U RNA editing. STAMP is isoform specific, can be multiplexed and enables detection of ribosome association in single cells.
- Kristopher W. Brannan
- , Isaac A. Chaim
- & Gene W. Yeo
-
Article |
 Single-cell joint detection of chromatin occupancy and transcriptome enables higher-dimensional epigenomic reconstructions
Single-cell joint detection of chromatin occupancy and transcriptome enables higher-dimensional epigenomic reconstructions
This paper reports CoTECH, which couples chromatin binding enrichment with RNA sequencing for concurrent measurements of histone modification and transcriptome in single cells, offering a multiomics tool for studying epigenetic regulations.
- Haiqing Xiong
- , Yingjie Luo
- & Aibin He
-
Article |
 PCprophet: a framework for protein complex prediction and differential analysis using proteomic data
PCprophet: a framework for protein complex prediction and differential analysis using proteomic data
PCprophet combines complex-level scoring and machine learning to predict novel protein complexes from protein cofractionation mass spectrometry data and to perform differential analysis across experimental conditions.
- Andrea Fossati
- , Chen Li
- & Ruedi Aebersold
-
Brief Communication |
 Genome-scale deconvolution of RNA structure ensembles
Genome-scale deconvolution of RNA structure ensembles
The DRACO algorithm deconvolutes coexisting RNA structures from mutational profiling experiments, and can be applied to bacterial regulatory structures and elements from the SARS-CoV-2 genome.
- Edoardo Morandi
- , Ilaria Manfredonia
- & Danny Incarnato
-
Research Highlight |
 Sequencing DNA bendability
Sequencing DNA bendability
Loop-seq is a high-throughput sequencing assay that measures DNA looping and can help explain how DNA bendability contributes to nucleosome organization.
- Lei Tang
-
Article |
 High-accuracy long-read amplicon sequences using unique molecular identifiers with Nanopore or PacBio sequencing
High-accuracy long-read amplicon sequences using unique molecular identifiers with Nanopore or PacBio sequencing
This work presents a sequencing strategy based on unique molecular identifiers that improves long-read consensus sequence accuracy of targeted amplicons as well as shotgun whole-genome fragments.
- Søren M. Karst
- , Ryan M. Ziels
- & Mads Albertsen
-
Comment |
 Spatially resolved transcriptomics adds a new dimension to genomics
Spatially resolved transcriptomics adds a new dimension to genomics
As single-cell omics continue to advance, the field of spatially resolved transcriptomics has emerged with a set of experimental and computational methods to map out the positions of cells and their gene expression profiles in space. Here we summarize current transcriptome-wide and sequencing-based methodologies and their applications in genomics research.
- Ludvig Larsson
- , Jonas Frisén
- & Joakim Lundeberg
Browse broader subjects
Browse narrower subjects
- Cell division
- Chromatin
- Chromosomes
- CRISPR-Cas systems
- DNA damage and repair
- DNA metabolism
- DNA recombination
- DNA replication
- Epigenetics
- Non-coding RNAs
- Nuclear organization
- Post-translational modifications
- Protein folding
- Proteolysis
- Proteomics
- Riboswitches
- Ribozymes
- RNA metabolism
- RNAi
- Single-molecule biophysics
- Transcription
- Transcriptomics
- Translation
- Transposition

 Parts-based decomposition of spatial genomics data finds distinct tissue regions
Parts-based decomposition of spatial genomics data finds distinct tissue regions