Abstract
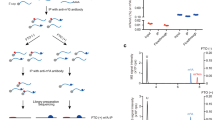
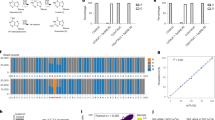
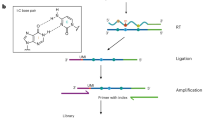
N6-methyladenosine (m6A) is the most abundant RNA modification in mammalian cells and the best-studied epitranscriptomic mark. Despite the development of various tools to map m6A, a transcriptome-wide method that enables absolute quantification of m6A at single-base resolution is lacking. Here we use glyoxal and nitrite-mediated deamination of unmethylated adenosines (GLORI) to develop an absolute m6A quantification method that is conceptually similar to bisulfite-sequencing-based quantification of DNA 5-methylcytosine. We apply GLORI to quantify the m6A methylomes of mouse and human cells and reveal clustered m6A modifications with differential distribution and stoichiometry. In addition, we characterize m6A dynamics under stress and examine the quantitative landscape of m6A modification in gene expression regulation. GLORI is an unbiased, convenient method for the absolute quantification of the m6A methylome.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
The sequence data generated in this study have been deposited in the NCBI Gene Expression Omnibus (GEO), under accession code GSE210563. The public MAZTER-seq, miCLIP, and m6A-seq datasets were downloaded from the GEO database (GSE122961, GSE63753, SRA280261, and GSE165690). The published ribosome profiling datasets of HEK293T, HeLa, and MEF were download from the GEO database (GSE129194, GSE63591, and SRA280261). The life-time data of HeLa cell line were downloaded from the GEO database (GSE49339). The reference genome GRCh38 and GRCm38 were download from the following link: https://hgdownload.soe.ucsc.edu/downloads.html.
Code availability
The code of the GLORI-tools is available on GitHub from https://github.com/liucongcas/GLORI-tools.
References
Roundtree, I. A., Evans, M. E., Pan, T. & He, C. Dynamic RNA modifications in gene expression regulation. Cell 169, 1187–1200 (2017).
Frye, M., Jaffrey, S. R., Pan, T., Rechavi, G. & Suzuki, T. RNA modifications: what have we learned and where are we headed? Nat. Rev. Genet. 17, 365–372 (2016).
Perry, R. P. & Kelley, D. E. Existence of methylated messenger RNA in mouse L cells. Cell 1, 37–42 (1974).
Perry, R. P., Kelley, D. E., Friderici, K. & Rottman, F. The methylated constituents of L cell messenger RNA: evidence for an unusual cluster at the 5′ terminus. Cell 4, 387–394 (1975).
Bokar, J. A., Rath-Shambaugh, M. E., Ludwiczak, R., Narayan, P. & Rottman, F. Characterization and partial purification of mRNA N6-adenosine methyltransferase from HeLa cell nuclei. internal mRNA methylation requires a multisubunit complex. J. Biol. Chem. 269, 17697–17704 (1994).
Bokar, J. A., Shambaugh, M. E., Polayes, D., Matera, A. G. & Rottman, F. M. Purification and cDNA cloning of the AdoMet-binding subunit of the human mRNA (N6-adenosine)-methyltransferase. RNA 3, 1233–1247 (1997).
Liu, J. et al. A METTL3–METTL14 complex mediates mammalian nuclear RNA N6-adenosine methylation. Nat. Chem. Biol. 10, 93–95 (2014).
Ping, X. L. et al. Mammalian WTAP is a regulatory subunit of the RNA N6-methyladenosine methyltransferase. Cell Res. 24, 177–189 (2014).
Schwartz, S. et al. Perturbation of m6A writers reveals two distinct classes of mRNA methylation at internal and 5– sites. Cell Rep 8, 284–296 (2014).
Patil, D. P. et al. m6A RNA methylation promotes XIST-mediated transcriptional repression. Nature 537, 369–373 (2016).
Jia, G. et al. N6-methyladenosine in nuclear RNA is a major substrate of the obesity-associated FTO. Nat. Chem. Biol. 7, 885–887 (2011).
Zheng, G. et al. ALKBH5 is a mammalian RNA demethylase that impacts RNA metabolism and mouse fertility. Mol. Cell 49, 18–29 (2013).
Barbieri, I. & Kouzarides, T. Role of RNA modifications in cancer. Nat. Rev. Cancer 20, 303–322 (2020).
Livneh, I., Moshitch-Moshkovitz, S., Amariglio, N., Rechavi, G. & Dominissini, D. The m6A epitranscriptome: transcriptome plasticity in brain development and function. Nat. Rev. Neurosci. 21, 36–51 (2020).
Li, X., Xiong, X. & Yi, C. Epitranscriptome sequencing technologies: decoding RNA modifications. Nat. Methods 14, 23–31 (2016).
Dominissini, D. et al. Topology of the human and mouse m6A RNA methylomes revealed by m6A-seq. Nature 485, 201–206 (2012).
Meyer, K. D. et al. Comprehensive analysis of mRNA methylation reveals enrichment in 3′ UTRs and near stop codons. Cell 149, 1635–1646 (2012).
Chen, K. et al. High-resolution N6-methyladenosine (m6A) map using photo-crosslinking-assisted m6A sequencing. Angew. Chem. Int. Ed. Engl. 54, 1587–1590 (2015).
Linder, B. et al. Single-nucleotide-resolution mapping of m6A and m6Am throughout the transcriptome. Nat. Methods 12, 767–772 (2015).
Molinie, B. et al. m6A-LAIC-seq reveals the census and complexity of the m6A epitranscriptome. Nat. Methods 13, 692–698 (2016).
Dierks, D. et al. Multiplexed profiling facilitates robust m6A quantification at site, gene and sample resolution. Nat. Methods 18, 1060–1067 (2021).
McIntyre, A. B. R. et al. Limits in the detection of m6A changes using MeRIP/m6A-seq. Sci Rep. 10, 6590 (2020).
Sun, H. et al. m6Am-seq reveals the dynamic m6Am methylation in the human transcriptome. Nat. Commun. 12, 4778 (2021).
Garcia-Campos, M. A. et al. Deciphering the ‘m6A Code’ via antibody-independent quantitative profiling. Cell 178, 731–747 e716 (2019).
Zhang, Z. et al. Single-base mapping of m6A by an antibody-independent method. Sci. Adv. 5, eaax0250 (2019).
Zhang, Z. et al. Systematic calibration of epitranscriptomic maps using a synthetic modification-free RNA library. Nat. Methods 18, 1213–1222 (2021).
Meyer, K. D. DART-seq: an antibody-free method for global m6A detection. Nat. Methods 16, 1275–1280 (2019).
Wang, Y., Xiao, Y., Dong, S., Yu, Q. & Jia, G. Antibody-free enzyme-assisted chemical approach for detection of N(6)-methyladenosine. Nat. Chem. Biol. 16, 896–903 (2020).
Shu, X. et al. A metabolic labeling method detects m6A transcriptome-wide at single base resolution. Nat. Chem. Biol. 16, 887–895 (2020).
Hu, L. et al. m6A RNA modifications are measured at single-base resolution across the mammalian transcriptome. Nat. Biotechnol. 40, 1210–1219 (2022).
Liu, N. et al. Probing N6-methyladenosine RNA modification status at single nucleotide resolution in mRNA and long noncoding RNA. RNA 19, 1848–1856 (2013).
Xiao, Y. et al. An elongation- and ligation-based qPCR amplification method for the radiolabeling-free detection of locus-specific N6-methyladenosine modification. Angew. Chem. Int. Ed. Engl. 57, 15995–16000 (2018).
Aschenbrenner, J. et al. Engineering of a DNA polymerase for direct m6A sequencing. Angew. Chem. Int. Ed. Engl. 57, 417–421 (2018).
Liu, W. et al. Identification of a selective DNA ligase for accurate recognition and ultrasensitive quantification of N6-methyladenosine in RNA at one-nucleotide resolution. Chem. Sci. 9, 3354–3359 (2018).
Raiber, E. A., Hardisty, R., van Delft, P. & Balasubramanian, S. Mapping and elucidating the function of modified bases in DNA. Nat. Rev. Chem. 1, 0069 (2017).
Shapiro, R. & Pohl, S. H. The reaction of ribonucleosides with nitrous acid. Side products and kinetics. Biochemistry 7, 448–455 (1968).
Schuster, H. & Wilhelm, R. C. Reaction differences between tobacco mosaic virus and its free ribonucleic acid with nitrous acid. Biochim. Biophys. Acta 68, 554–560 (1963).
Shapiro, R. & Hachmann, J. The reaction of guanine derivatives with 1,2-dicarbonyl compounds. Biochemistry 5, 2799–2807 (1966).
Nakaya, K., Takenaka, O., Horinishi, H. & Shibata, K. Reactions of glyoxal with nucleic acids. Nucleotides and their component bases. Biochim. Biophys. Acta 161, 23–31 (1968).
Broude, N. E. & Budowsky, E. I. The reaction of glyoxal with nucleic acid components. 3. Kinetics of the reaction with monomers. Biochim. Biophys. Acta 254, 380–388 (1971).
Cattenoz, P. B., Taft, R. J., Westhof, E. & Mattick, J. S. Transcriptome-wide identification of A > I RNA editing sites by inosine specific cleavage. RNA 19, 257–270 (2013).
Knutson, S. D., Arthur, R. A., Johnston, H. R. & Heemstra, J. M. Selective enrichment of A-to-I edited transcripts from cellular RNA using endonuclease V. J. Am. Chem. Soc. 142, 5241–5251 (2020).
Kivioja, T. et al. Counting absolute numbers of molecules using unique molecular identifiers. Nat. Methods 9, 72–74 (2011).
Islam, S. et al. Quantitative single-cell RNA-seq with unique molecular identifiers. Nat. Methods 11, 163–166 (2014).
Wei, J. et al. Differential m6A, m6Am, and m1A demethylation mediated by FTO in the cell nucleus and cytoplasm. Mol. Cell 71, 973–985 e975 (2018).
Levanon, E. Y. et al. Systematic identification of abundant A-to-I editing sites in the human transcriptome. Nat. Biotechnol. 22, 1001–1005 (2004).
Tan, M. H. et al. Dynamic landscape and regulation of RNA editing in mammals. Nature 550, 249–254 (2017).
Huang, T., Chen, W., Liu, J., Gu, N. & Zhang, R. Genome-wide identification of mRNA 5-methylcytosine in mammals. Nat. Struct. Mol. Biol. 26, 380–388 (2019).
Yang, X. et al. 5-methylcytosine promotes mRNA export—NSUN2 as the methyltransferase and ALYREF as an m5C reader. Cell Res. 27, 606–625 (2017).
Yankova, E. et al. Small-molecule inhibition of METTL3 as a strategy against myeloid leukaemia. Nature 593, 597–601 (2021).
Zhou, J. et al. Dynamic m6A mRNA methylation directs translational control of heat shock response. Nature 526, 591–594 (2015).
Meyer, K. D. et al. 5′ UTR m6A promotes cap-independent translation. Cell 163, 999–1010 (2015).
Mahdavi-Amiri, Y., Chung Kim Chung, K. & Hili, R. Single-nucleotide resolution of N6-adenine methylation sites in DNA and RNA by nitrite sequencing. Chem. Sci. 12, 606–612 (2020).
Werner, S. et al. NOseq: amplicon sequencing evaluation method for RNA m6A sites after chemical deamination. Nucleic Acids Res. 49, e23 (2021).
Moore, L. D., Le, T. & Fan, G. DNA methylation and its basic function. Neuropsychopharmacology 38, 23–38 (2013).
Mohn, F. et al. Lineage-specific polycomb targets and de novo DNA methylation define restriction and potential of neuronal progenitors. Mol. Cell 30, 755–766 (2008).
Dawson, M. A. & Kouzarides, T. Cancer epigenetics: from mechanism to therapy. Cell 150, 12–27 (2012).
Esteller, M. Cancer epigenomics: DNA methylomes and histone-modification maps. Nat. Rev. Genet. 8, 286–298 (2007).
Jones, P. A. Functions of DNA methylation: islands, start sites, gene bodies and beyond. Nat. Rev. Genet. 13, 484–492 (2012).
Horvath, S. & Raj, K. DNA methylation-based biomarkers and the epigenetic clock theory of ageing. Nat. Rev. Genet. 19, 371–384 (2018).
Park, Y., Figueroa, M. E., Rozek, L. S. & Sartor, M. A. MethylSig: a whole genome DNA methylation analysis pipeline. Bioinformatics 30, 2414–2422 (2014).
Bailey, T. L. et al. MEME SUITE: tools for motif discovery and searching. Nucleic Acids Res. 37, W202–W208 (2009).
Kim, D., Paggi, J. M., Park, C., Bennett, C. & Salzberg, S. L. Graph-based genome alignment and genotyping with HISAT2 and HISAT-genotype. Nat. Biotechnol. 37, 907–915 (2019).
Feng, J., Liu, T., Qin, B., Zhang, Y. & Liu, X. S. Identifying ChIP-seq enrichment using MACS. Nat. Protoc. 7, 1728–1740 (2012).
Meng, J. et al. A protocol for RNA methylation differential analysis with MeRIP-Seq data and exomePeak R/Bioconductor package. Methods 69, 274–281 (2014).
Acknowledgements
The authors would like to thank T. Luo, J. Liu, J. Zhou, P. Du, and G. Luo for discussions and help. We also thank P. Chai and Y. Cheng for assistance in the hypoxia experiment. We thank the National Center for Protein Sciences at Peking University in Beijing, China for assistance with quantification of library size distribution. We thank the State Key Laboratory of Natural and Biomimetic Drugs at Peking University for advice and support in technology. We thank the High Performance Computing Platform of the Center for Life Science for assistance with the analysis. This work was supported by the National Key R&D Program of China (2019YFA0802201 to C.Y., 2017YFA0505202 to J.W., 2019YFA0110902 to C.Y., and 2020YFA0710401 to J.P.) and the National Natural Science Foundation of China (22077006 to J.W., 91940304 to C.Y., 21825701 to C.Y., and 91853107 to J.W.).
Author information
Authors and Affiliations
Contributions
J.W. and C.Y. conceived the project and designed the experiments; C.Y., J.W., C.L., H.S., K.L., and W.S. wrote the manuscript with the help of J.P.; H.S., Y.Y., and W.S. performed the experiments with the help of F.L., Y.X., and Y.H.; C.L developed the bioinformatics pipeline of GLORI; J.W. and Y.Y. developed the chemical method of GLORI; C.L. and K.L. designed and performed the bioinformatics analysis with the help of C.Y., H.M., B.L., and W.L.; H.S. designed and conducted all sample preparation for next-generation sequencing with the help of Y.X.; W.S. and H.S. further optimized the reaction condition with the help of F.L. and Y.L.; J.W. and C.Y. supervised the project.
Corresponding authors
Ethics declarations
Competing interests
A patent application has been filed by Peking University for the technology disclosed in this publication.
Peer review
Peer review information
Nature Biotechnology thanks Kathy Liu and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Figs 1–15.
Supplementary Data 1
GLORI-identified m6A sites in HEK293T cells.
Supplementary Data 2
GLORI-identified m6A sites in METTL3, METTL14, WTAP, and control knockdown cells.
Supplementary Data 3
GLORI-identified m6A sites in STM2457 treated cells.
Supplementary Data 4
GLORI-identified m6A sites in YTHDF2 and control knockdown cells.
Supplementary Data 5
Hypoxia stress-inducible m6A sites.
Supplementary Data 6
Heat shock stress-inducible m6A sites.
Supplementary Data 7
Spike-in model sequences and siRNA sequences.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Liu, C., Sun, H., Yi, Y. et al. Absolute quantification of single-base m6A methylation in the mammalian transcriptome using GLORI. Nat Biotechnol 41, 355–366 (2023). https://doi.org/10.1038/s41587-022-01487-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41587-022-01487-9
This article is cited by
-
Dissecting the sequence and structural determinants guiding m6A deposition and evolution via inter- and intra-species hybrids
Genome Biology (2024)
-
Decoding epitranscriptomic regulation of viral infection: mapping of RNA N6-methyladenosine by advanced sequencing technologies
Cellular & Molecular Biology Letters (2024)
-
Programmable protein expression using a genetically encoded m6A sensor
Nature Biotechnology (2024)
-
GLORI for absolute quantification of transcriptome-wide m6A at single-base resolution
Nature Protocols (2024)
-
Detecting m6A at single-molecular resolution via direct RNA sequencing and realistic training data
Nature Communications (2024)