Featured
-
-
Letter |
 TLR recognition of self nucleic acids hampers glucocorticoid activity in lupus
TLR recognition of self nucleic acids hampers glucocorticoid activity in lupus
Glucocorticoids are widely used to treat patients with autoimmune diseases such as systemic lupus erythematosus (SLE), but many treatment regimens cannot maintain disease control in SLE patients. Here it is shown that the stimulation of plasmacytoid dendritic cells through toll-like receptors TLR7 and TLR9 can account for the reduced activity of glucocorticoids to inhibit the type I interferon pathway in SLE patients. Thus inhibitors of TLR7 and TLR9 signalling might prove to be effective corticosteroid-sparing drugs.
- Cristiana Guiducci
- , Mei Gong
- & Franck J. Barrat
-
Letter |
 Small regulatory RNAs inhibit RNA polymerase II during the elongation phase of transcription
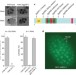
Small regulatory RNAs inhibit RNA polymerase II during the elongation phase of transcription
Small regulatory RNAs function both in the cytoplasm, inhibiting expression from messenger RNAs, and in the nucleus, silencing heterochromatin and preventing genome rearrangement. Now a new protein involved in RNA interference in the nucleus has been characterized. This protein, NRDE-2, associates with NRDE-3 and short interfering RNAs on nascent transcripts. This association prevents elongation of the transcripts by RNA polymerase II, making this a co-transcriptional form of gene silencing.
- Shouhong Guang
- , Aaron F. Bochner
- & Scott Kennedy
-
Letter |
 Tumour angiogenesis is reduced in the Tc1 mouse model of Down’s syndrome
Tumour angiogenesis is reduced in the Tc1 mouse model of Down’s syndrome
Down's syndrome is caused by trisomy of chromosome 21, and it is known that the growth of certain tumours is reduced in this genetic disorder. Using a mouse model of Down's syndrome, several individual genes on chromosome 21 are now being proposed to mediate the effect on tumour growth and angiogenesis.
- Louise E. Reynolds
- , Alan R. Watson
- & Kairbaan M. Hodivala-Dilke
-
Article |
 Crystal structure of HIV-1 Tat complexed with human P-TEFb
Crystal structure of HIV-1 Tat complexed with human P-TEFb
Here the 2.1 Å crystal structure of human immunodeficiency virus (HIV) Tat protein complexed with the positive transcription elongation factor P-TEFb is reported. This shows that Tat binding changes the structure of P-TEFb, which may suggest opportunities for developing inhibitors that block only the form of P-TEFb used by the virus.
- Tahir H. Tahirov
- , Nigar D. Babayeva
- & David H. Price
-
-
Letter |
 Termination of autophagy and reformation of lysosomes regulated by mTOR
Termination of autophagy and reformation of lysosomes regulated by mTOR
When cells are starved, the enzyme TOR is inhibited, inducing autophagy. In this process, autophagosomes sequester intracellular components and then fuse with lysosomes, producing autolysosomes in which cargo is degraded to regenerate nutrients. Now, a mechanism is revealed by which lysosomes are re-formed. When starvation conditions are prolonged, mTOR is re-activated; this attenuates autophagy and results in tubules and vesicles extruding from the autolysosome and maturing into functional lysosomes.
- Li Yu
- , Christina K. McPhee
- & Michael J. Lenardo
-
Letter |
 A novel and unified two-metal mechanism for DNA cleavage by type II and IA topoisomerases
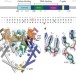
A novel and unified two-metal mechanism for DNA cleavage by type II and IA topoisomerases
Topoisomerases are enzymes that transiently make breaks in DNA, to prevent the build-up of topological stress and tangles as the genome is transcribed, replicated or repaired. Type II topoisomerases have been postulated to use a two-metal mechanism to break duplex DNA. Now, the structure of the DNA-binding and cleavage core of a yeast type II topoisomerase has been solved, showing that the enzyme uses a variation of the classical mechanism, and can also carry out the type of cleavage performed by type IA topoisomerases.
- Bryan H. Schmidt
- , Alex B. Burgin
- & James M. Berger
-
Article |
 HIF-1 antagonizes p53-mediated apoptosis through a secreted neuronal tyrosinase
HIF-1 antagonizes p53-mediated apoptosis through a secreted neuronal tyrosinase
When oxygen levels drop in a tissue, the transcription factor hypoxia-inducible factor (HIF) is activated to regulate the cellular response. HIFα levels are increased in most solid tumours and this correlates with a poor prognosis, for unknown reasons. Here it is shown that HIF-1, the worm version of HIFα, protects germ cells from DNA-damage-induced death. It does this remotely, by increasing the production of the TYR-2 protein in distant neurons. Inhibiting a human TYR-2 homologue promotes apoptosis in melanoma cells.
- Ataman Sendoel
- , Ines Kohler
- & Michael O. Hengartner
-
Research Highlights |
Genomics: Transposition trends
-
Letter |
 Relationship between nucleosome positioning and DNA methylation
Relationship between nucleosome positioning and DNA methylation
Nucleosomes are composed of around 147 bases of DNA wrapped around an octamer of histone proteins. Here, a genome-wide analysis of nucleosome positioning in Arabidopsis thaliana has been combined with profiles of DNA methylation at single base resolution, revealing 10-base periodicities in the DNA methylation status of nucleosome-bound DNA. The results indicate that nucleosome positioning influences the pattern of DNA methylation throughout the genome.
- Ramakrishna K. Chodavarapu
- , Suhua Feng
- & Matteo Pellegrini
-
Letter |
 Principles of stop-codon reading on the ribosome
Principles of stop-codon reading on the ribosome
Stop codons in messenger RNA define when a protein sequence has been completely synthesized; such codons bind release factors (RFs), which cause the newly made protein to be released. Structures of RFs alone and in combination with the ribosome have been reported, but the energetics of the reaction in the presence of codons had not been determined. Here, molecular dynamics simulations of 14 termination complexes are used to define how termination is achieved and how the RFs distinguish different sequences.
- Johan Sund
- , Martin Andér
- & Johan Åqvist
-
Letter |
 Phenotypic robustness conferred by apparently redundant transcriptional enhancers
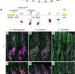
Phenotypic robustness conferred by apparently redundant transcriptional enhancers
Transcriptional enhancers are segments of regulatory DNA located some distance from the coding region of a gene, and several of them may sometimes serve apparently redundant functions. These authors demonstrate in Drosophila that such 'redundant' enhancers, by contributing higher overall levels of transcription, ensure robustness of phenotypes against both genetic and environmental perturbations, for example mutations in other genes or temperature changes that would otherwise lead to aberrant development.
- Nicolás Frankel
- , Gregory K. Davis
- & David L. Stern
-
Letter |
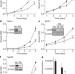 CD95 promotes tumour growth
CD95 promotes tumour growth
CD95 is a classical death receptor protein that regulates tissue homeostasis by inducing cell death. Here it is shown, however, that cancer cells depend on CD95 for optimal growth. Without CD95, the incidence of ovarian cancer and liver cancer in mice is reduced, as is the size of any tumours. So CD95 is a double-edged sword, and it may be necessary to reduce, rather than enhance, its activity in order to kill tumour cells.
- Lina Chen
- , Sun-Mi Park
- & Marcus E. Peter
-
Research Highlights |
Genomics: Not-so-dark genome
-
Letter |
 Structural basis for 5′-nucleotide base-specific recognition of guide RNA by human AGO2
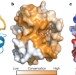
Structural basis for 5′-nucleotide base-specific recognition of guide RNA by human AGO2
The association of microRNAs with Argonaute proteins (AGOs) yields complexes regulating gene expression. Although bacterial and archaeal miRNAs show no sequence preference at their 5′ ends, eukaryotic miRNAs tend to have a 5′ U or A. Here the structure of the human AGO2 MID domain complexed with ribonucleotide monophosphates is solved, revealing specific interaction of UMP and AMP with a loop that discriminates against CMP or GMP, and explaining the observed preference.
- Filipp Frank
- , Nahum Sonenberg
- & Bhushan Nagar
-
Letter |
 Structure of the bifunctional isocitrate dehydrogenase kinase/phosphatase
Structure of the bifunctional isocitrate dehydrogenase kinase/phosphatase
The Escherichia coli isocitrate dehydrogenase kinase/phosphatase (AceK) is a bifunctional enzyme that can phosphorylate or dephosphorylate isocitrate dehydrogenase (ICDH) to either inactivate or activate it in response to environmental changes. Now the structures of AceK and the AceK–ICDH complex have been solved, revealing the conformational changes that occur when AceK changes from a kinase to a phosphatase and vice versa.
- Jimin Zheng
- & Zongchao Jia
-
Letter |
 Distinct FGFs promote differentiation of excitatory and inhibitory synapses
Distinct FGFs promote differentiation of excitatory and inhibitory synapses
Proper functioning of the brain requires a balance between the formation of excitatory and inhibitory synapses, but how this is achieved during development is unclear. Here FGF22 and FGF7, two fibroblast growth factor cell–cell signalling molecules, are shown to promote the formation of excitatory and inhibitory synapses, respectively, through their effect on epilepsy in mice. These findings should inform other neurological and psychiatric disorders involving defects in synapse formation.
- Akiko Terauchi
- , Erin M. Johnson-Venkatesh
- & Hisashi Umemori
-
Letter |
 The folding cooperativity of a protein is controlled by its chain topology
The folding cooperativity of a protein is controlled by its chain topology
Proteins often comprise domains that can be distinguished as relatively separate regions in the three-dimensional structure. Communication between these domains is important for catalysis, regulation and folding, but how they communicate is largely unclear. Here, single-molecule optical tweezers were used to pull on a protein while monitoring the energetics of unfolding and refolding events in disparate regions. By comparing topological variations of the same protein, new rules of cooperation between domains were derived.
- Elizabeth A. Shank
- , Ciro Cecconi
- & Carlos Bustamante
-
Letter |
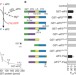 eIF5 has GDI activity necessary for translational control by eIF2 phosphorylation
eIF5 has GDI activity necessary for translational control by eIF2 phosphorylation
The initiation of protein synthesis requires the eukaryotic translation initiation factor (eIF) 2, which uses energy from the hydrolysis of GTP. Another factor, eIF5, accelerates the GTP-hydrolysing activity of eIF2. Here, two other roles for eIF5 have been defined. One involves stabilizing GDP, the product of GTP hydrolysis, on eIF2. In its other role, eIF5 works with phosphorylated eIF2 to inhibit the guanine-nucleotide exchange factor eIF2B. These results clarify our understanding of how the initiation of translation is regulated.
- Martin D. Jennings
- & Graham D. Pavitt
-
Article |
 Chemical genetics of Plasmodium falciparum
Chemical genetics of Plasmodium falciparum
Here, a library of more than 300,000 chemicals was screened for activity against Plasmodium falciparum growing in red blood cells. Of these chemicals, 172 representative candidates were profiled in detail; one exemplar compound showed efficacy in a mouse model of malaria. The findings provide the scientific community with new starting points for drug discovery.
- W. Armand Guiguemde
- , Anang A. Shelat
- & R. Kiplin Guy
-
News & Views |
 Roots respond to an inner calling
Roots respond to an inner calling
In plant roots, patterning of two types of water-conducting xylem tissue is determined by a signalling system that involves the reciprocal dance of a mobile transcription factor and mobile microRNAs.
- Ben Scheres
-
News & Views |
 Decision at the break point
Decision at the break point
Many decisions affect the fate of damaged DNA — for example, how to repair the damage, or whether to repair it at all and instead let the damaged cell die. An intricate web of molecular interactions affects such decisions.
- Simon J. Boulton
-
Article |
 Helical assembly in the MyD88–IRAK4–IRAK2 complex in TLR/IL-1R signalling
Helical assembly in the MyD88–IRAK4–IRAK2 complex in TLR/IL-1R signalling
Toll-like receptors (TLRs) are crucial to innate immunity. Activation of these proteins, and of receptors for the pro-inflammatory cytokines IL-1 and IL-18, leads to the recruitment of adaptor proteins such as MyD88. These in turn interact with further proteins such as IRAK2 and IRAK4. The crystal structure of the MyD88–IRAK2–IRAK4 death domain complex is now reported, explaining how these three proteins cooperate in TLR/IL-1R signalling.
- Su-Chang Lin
- , Yu-Chih Lo
- & Hao Wu
-
News |
 Existence of RNA 'dark matter' in doubt
Existence of RNA 'dark matter' in doubt
The abundance of transcripts from the genome may have been overestimated.
- Melissa Lee Phillips
-
Letter |
 Moonlighting bacteriophage proteins derepress staphylococcal pathogenicity islands
Moonlighting bacteriophage proteins derepress staphylococcal pathogenicity islands
Staphylococcal superantigens can lead to toxic shock syndrome. They are encoded on pathogenicity islands and with the aid of helper phages can be excised and packaged into highly transmissable phage particles. Here it is shown that a specific, non-essential helper phage protein is responsible for derepression of the pathogenicity island, thereby providing the mechanism for the first step of its mobilization.
- María Ángeles Tormo-Más
- , Ignacio Mir
- & José R. Penadés
-
Letter |
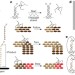 Molecular robots guided by prescriptive landscapes
Molecular robots guided by prescriptive landscapes
Traditional robots need to store internal representations of their goals and environment, and to coordinate sensing and the movement of components required in response. Individual molecules are limited in their ability to store complex information, but robotic behaviour can still be realized — as has now been shown with DNA walkers, which can carry out a sequence of actions such as 'start', 'follow', 'turn' and 'stop' that are programmed into the DNA landscape on which the walkers move.
- Kyle Lund
- , Anthony J. Manzo
- & Hao Yan
-
Letter |
 Self-assembly of spider silk proteins is controlled by a pH-sensitive relay
Self-assembly of spider silk proteins is controlled by a pH-sensitive relay
Spider silk proteins are remarkably soluble when stored at high concentration and yet can be converted to extremely sturdy fibres, through unknown molecular mechanisms. Here, the X-ray structure of the amino-terminal domain of a silk protein is presented, revealing how evolutionarily conserved polar surfaces might control self-assembly as the pH is lowered along the spider's silk extrusion duct. Such a mechanism might be applicable to the design of versatile fibrous materials.
- Glareh Askarieh
- , My Hedhammar
- & Stefan D. Knight
-
Letter |
 Induction of tumour immunity by targeted inhibition of nonsense-mediated mRNA decay
Induction of tumour immunity by targeted inhibition of nonsense-mediated mRNA decay
The main reason why tumours are not controlled by the immune system is that they do not express potent tumour rejection antigens. Tumour vaccination aims to provoke a response to any antigens that are expressed. Here, a new approach is described: nonsense-mediated messenger RNA decay in tumour cells is inhibited, leading to the expression of new antigens and to significant inhibition of tumour growth in mice.
- Fernando Pastor
- , Despina Kolonias
- & Eli Gilboa
-
Research Highlights |
Synthetic biology: Search and destroy
-
News & Views |
 Enhancers make non-coding RNA
Enhancers make non-coding RNA
Genomes don't just encode protein-coding RNAs. They also give rise to various groups of RNAs that can regulate gene expression. Short RNAs that form from enhancer sequences might be one such class of regulatory RNA.
- Bing Ren
-
News & Views |
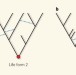 Common ancestry put to the test
Common ancestry put to the test
The question of whether or not all life on Earth has an ultimate common origin is a subtle one, complicated by the phenomenon of lateral gene transfer. It has now been tackled with a formal statistical analysis.
- Mike Steel
- & David Penny
-
News |
 Genomics goes beyond DNA sequence
Genomics goes beyond DNA sequence
A technology that simultaneously reads a DNA sequence and its crucial modifications makes its debut.
- Alla Katsnelson
-
Letter |
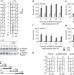 Ubiquitin-dependent DNA damage bypass is separable from genome replication
Ubiquitin-dependent DNA damage bypass is separable from genome replication
Post-replicative repair (PRR) enables cells to bypass or overcome DNA damage during DNA replication. In eukaryotes, ubiquitylation of the replication clamp PCNA by components of the RAD6 pathway activates damage bypass. When this occurs has been debated. It is now shown that PRR can be postponed until much of the undamaged genome is replicated. Moreover, it seems that PRR occurs mainly by an error-prone process, with error-free bypass playing a minor role.
- Yasukazu Daigaku
- , Adelina A. Davies
- & Helle D. Ulrich
-
Article |
 Deciphering the splicing code
Deciphering the splicing code
The coding capacity of the genome is greatly expanded by the process of alternative splicing, which enables a single gene to produce more than one distinct protein. Can the expression of these different proteins be predicted from sequence data? Here, modelling based on information theory has been used to develop a 'splicing code', which can predict, with good accuracy, tissue-dependent changes in alternative splicing.
- Yoseph Barash
- , John A. Calarco
- & Brendan J. Frey
-
Letter |
 A three-dimensional model of the yeast genome
A three-dimensional model of the yeast genome
The topologies of, and spatial relationships between, chromosomes are important but poorly understood. Here, a high-throughput method is used to study intra- and inter-chromosomal interactions in Saccharomyces cerevisiae. A map of the haploid genome is generated at kilobase resolution, and is used to construct a three-dimensional model of the yeast genome. The findings provide a glimpse of the interface between the form and function of a eukaryotic genome.
- Zhijun Duan
- , Mirela Andronescu
- & William S. Noble
-
Letter |
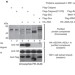 Histone H2A deubiquitinase activity of the Polycomb repressive complex PR-DUB
Histone H2A deubiquitinase activity of the Polycomb repressive complex PR-DUB
Polycomb group (PcG) proteins are transcriptional repressors that modify chromatin and regulate important developmental genes. One PcG-associated, chromatin-modifying activity is an enzyme that ubiquitinates histone H2A of chromatin. Here, a fruitfly PcG complex that is associated with H2A deubiquitination, and thereby with gene repression, is identified. PcG-mediated gene silencing might thus involve a dynamic balance between ubiquitination and deubiquitination of H2A.
- Johanna C. Scheuermann
- , Andrés Gaytán de Ayala Alonso
- & Jürg Müller
-
-
Article |
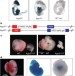 A dicer-independent miRNA biogenesis pathway that requires Ago catalysis
A dicer-independent miRNA biogenesis pathway that requires Ago catalysis
MicroRNAs, which regulate gene expression, are transcribed as longer sequences that are processed to produce the mature form. Two nuclease enzymes, Drosha and Dicer, are known to act sequentially to trim the microRNA to size. Here, however, a subset of microRNAs that includes miR-451, important for erythropoiesis, is found to be processed independently of Dicer. Rather, the Argonaute protein — part of the complex that aligns microRNA and messenger RNA — carries out the secondary cleavage.
- Sihem Cheloufi
- , Camila O. Dos Santos
- & Gregory J. Hannon
-
Letter |
 Cis-interactions between Notch and Delta generate mutually exclusive signalling states
Cis-interactions between Notch and Delta generate mutually exclusive signalling states
Notch and Delta are transmembrane proteins that allow neighbouring cells to communicate during development. Here, quantitative time-lapse microscopy has been used to show that the response of Notch to Delta on a neighbouring cell is graded, whereas its response to Delta on the same cell is sharp and occurs at a fixed threshold. A mathematical model explores how this new design principle enhances the sharpness of developmental boundaries set by classical lateral inhibition.
- David Sprinzak
- , Amit Lakhanpal
- & Michael B. Elowitz
-
Article |
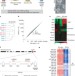 Aberrant silencing of imprinted genes on chromosome 12qF1 in mouse induced pluripotent stem cells
Aberrant silencing of imprinted genes on chromosome 12qF1 in mouse induced pluripotent stem cells
Induced pluripotent stem cells (iPSCs) are generated by the enforced expression of particular transcription factors in somatic cells. The extent to which such cells are equivalent to embryonic stem (ES) cells is an open question. Here, genetically identical mouse ES cells and iPSCs have been compared; the overall expression patterns of messenger RNAs and microRNAs are the same, with the exception of a few transcripts encoded within an imprinted gene cluster on chromosome 12qF1.
- Matthias Stadtfeld
- , Effie Apostolou
- & Konrad Hochedlinger
-
Letter |
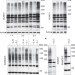 Listeria monocytogenes impairs SUMOylation for efficient infection
Listeria monocytogenes impairs SUMOylation for efficient infection
SUMOylation is a post-translational protein modification that affects many eukaryotic cellular processes. It is shown here that cellular infection with Listeria monocytogenes induces degradation of one of the essential SUMOylation enzymes, Ubc9, through a mechanism that involves a bacterial toxin, listeriolysin O. This effect on SUMOylation may support efficient infection by Listeria.
- David Ribet
- , Mélanie Hamon
- & Pascale Cossart
-
Technology Feature |
 MicroRNAs as biomarkers
MicroRNAs as biomarkers
-
Research Highlights |
Cancer biology: Cells combat chemo
-
Letter |
 An RNA polymerase II- and AGO4-associated protein acts in RNA-directed DNA methylation
An RNA polymerase II- and AGO4-associated protein acts in RNA-directed DNA methylation
DNA methylation is an important epigenetic mark in many eukaryotes. In Arabidopsis plants, small interfering RNAs bound to the Argonaute 4 (AGO4) protein can direct de novo DNA methylation and consequent gene silencing. Here, a new regulator of RNA-directed DNA methylation has been discovered. This protein, RDM1, is proposed to bind to methylated DNA and to function in the AGO4 effector complex.
- Zhihuan Gao
- , Hai-Liang Liu
- & Jian-Kang Zhu
-
Letter |
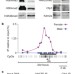 CpG islands influence chromatin structure via the CpG-binding protein Cfp1
CpG islands influence chromatin structure via the CpG-binding protein Cfp1
Most human gene promoters are embedded within CpG islands that lack DNA methylation and coincide with sites at which histone H3 lysine 4 is trimethylated (H3K4me3 sites). Here, a zinc-finger protein, Cfp1, is found to be associated with non-methylated CpG islands and H3K4me3 sites throughout the genome in the mouse brain. A primary function of non-methylated CpG islands might be to genetically determine the local chromatin modification state by interaction with Cfp1 and perhaps other CpG-binding proteins.
- John P. Thomson
- , Peter J. Skene
- & Adrian Bird
-
Article |
 Real-time tRNA transit on single translating ribosomes at codon resolution
Real-time tRNA transit on single translating ribosomes at codon resolution
Single-molecule studies allow biological processes to be examined one molecule at a time, as they occur. Here, zero-mode waveguides have been used to concentrate reactions in zeptolitre-sized volumes, making it possible to study real-time translocation by the ribosome. The binding of transfer RNAs (tRNAs) to the ribosome could be followed; the results show that tRNA release from the exit site is uncoupled from tRNA binding to the aminoacyl-tRNA site.
- Sotaro Uemura
- , Colin Echeverría Aitken
- & Joseph D. Puglisi
-
Letter |
 Long non-coding RNA HOTAIR reprograms chromatin state to promote cancer metastasis
Long non-coding RNA HOTAIR reprograms chromatin state to promote cancer metastasis
Large intervening non-coding RNAs (lincRNAs) are pervasively transcribed in the genome. Here it is shown that lincRNAs in the HOX genetic loci are dysregulated during breast cancer progression in human cells, and that expression levels of the lincRNA called HOTAIR can predict whether a tumour will metastasize. Moreover, enforced expression of HOTAIR can lead to altered patterns of binding of the PRC2 protein to the genome.
- Rajnish A. Gupta
- , Nilay Shah
- & Howard Y. Chang
-
News & Views |
 A ribosome in action
A ribosome in action
The manufacture of proteins by ribosomes involves complex interactions of diverse nucleic-acid and protein ligands. Single-molecule studies allow us, for the first time, to follow the synthesis of full-length proteins in real time.
- Susanne Brakmann
-
Article |
 Widespread transcription at neuronal activity-regulated enhancers
Widespread transcription at neuronal activity-regulated enhancers
Regulatory proteins bind non-coding DNA either at promoters (near to a gene's transcription start site) or at enhancers (far away). Binding at enhancers helps to bring the transcription enzyme RNA polymerase to promoters. Here, studies of some 12,000 enhancers that respond to electrical activity in neurons show that binding to enhancers also brings the polymerase to the enhancers themselves, where it transcribes a novel class of non-coding RNAs. Enhancers may thus be more similar to promoters than hitherto appreciated.
- Tae-Kyung Kim
- , Martin Hemberg
- & Michael E. Greenberg
Browse broader subjects
Browse narrower subjects
- Cell division
- Chromatin
- Chromosomes
- CRISPR-Cas systems
- DNA damage and repair
- DNA metabolism
- DNA recombination
- DNA replication
- Epigenetics
- Non-coding RNAs
- Nuclear organization
- Post-translational modifications
- Protein folding
- Proteolysis
- Proteomics
- Riboswitches
- Ribozymes
- RNA metabolism
- RNAi
- Single-molecule biophysics
- Transcription
- Transcriptomics
- Translation
- Transposition

 TFIIA and the transactivator Rap1 cooperate to commit TFIID for transcription initiation
TFIIA and the transactivator Rap1 cooperate to commit TFIID for transcription initiation Brain function and chromatin plasticity
Brain function and chromatin plasticity