Article
|
Open Access
Featured
-
-
Article
| Open Access Structures, functions and adaptations of the human LINE-1 ORF2 protein
Structures, functions and adaptations of the human LINE-1 ORF2 protein
X-ray crystallography, cryo-electron microscopy, structural modelling, biochemistry, cell biology, and evolutionary analysis enable characterization of ORF2p, the reverse transcriptase of the ancient ‘parasitic’ LINE-1 retrotransposon that has written around one-third of the human genome.
- Eric T. Baldwin
- , Trevor van Eeuwen
- & Martin S. Taylor
-
Article
| Open Access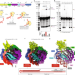 Template and target-site recognition by human LINE-1 in retrotransposition
Template and target-site recognition by human LINE-1 in retrotransposition
Human LINE-1 ORF2p relies on upstream single-stranded target DNA to position the adjacent duplex in the endonuclease active site for nicking of the longer DNA strand, with a single nick generating a staggered DNA break.
- Akanksha Thawani
- , Alfredo Jose Florez Ariza
- & Kathleen Collins
-
Article |
 Transposon-encoded nucleases use guide RNAs to promote their selfish spread
Transposon-encoded nucleases use guide RNAs to promote their selfish spread
TnpB and IscB nucleases use transposon-encoded guide RNAs to target genomic sequences for cleavage, thereby favouring copying and spreading of transposable elements.
- Chance Meers
- , Hoang C. Le
- & Samuel H. Sternberg
-
Article |
 Retrotransposons hijack alt-EJ for DNA replication and eccDNA biogenesis
Retrotransposons hijack alt-EJ for DNA replication and eccDNA biogenesis
Retrotransposons hijack the alternative end-joining DNA repair process of the host for a circularization step to synthesize their second-strand DNA.
- Fu Yang
- , Weijia Su
- & ZZ Zhao Zhang
-
Article
| Open Access Cryo-EM structure of the transposon-associated TnpB enzyme
Cryo-EM structure of the transposon-associated TnpB enzyme
Cryo-electron microscopy analysis of the Deinococcus radiodurans ISDra2 TnpB in complex with its cognate ωRNA and target DNA provides insights into the mechanism of TnpB function and the evolution of CRISPR–Cas12 effectors.
- Ryoya Nakagawa
- , Hisato Hirano
- & Osamu Nureki
-
Article
| Open Access Structures of the holo CRISPR RNA-guided transposon integration complex
Structures of the holo CRISPR RNA-guided transposon integration complex
Structural studies of the CRISPR-associated transposon comprising Cas12k, TnsC, TnsB and TniQ from Scytonema hofmannii using cryo-electron microscopy reveal insights into the architecture and mechanism of RNA-guided DNA transposition.
- Jung-Un Park
- , Amy Wei-Lun Tsai
- & Elizabeth H. Kellogg
-
Article |
 Selective TnsC recruitment enhances the fidelity of RNA-guided transposition
Selective TnsC recruitment enhances the fidelity of RNA-guided transposition
CRISPR-associated transposons exploit a proofreading checkpoint to ensure high-fidelity selection of genomic sites for DNA insertion through specific recruitment of TnsC oligomers by the Cascade complex.
- Florian T. Hoffmann
- , Minjoo Kim
- & Samuel H. Sternberg
-
Article
| Open Access Genome surveillance by HUSH-mediated silencing of intronless mobile elements
Genome surveillance by HUSH-mediated silencing of intronless mobile elements
The human silencing hub (HUSH) complex uses introns to distinguish intronless foreign DNA from intron-containing host DNA and modifies chromatin to silence transcription of retrotransposons and retroviruses.
- Marta Seczynska
- , Stuart Bloor
- & Paul J. Lehner
-
Article |
 Target site selection and remodelling by type V CRISPR-transposon systems
Target site selection and remodelling by type V CRISPR-transposon systems
Structural studies on Scytonema hofmanni CRISPR-associated transposon protein complexes indicate a mechanism for RNA-guided DNA transposition involving Cas12k, TnsC and TnsB.
- Irma Querques
- , Michael Schmitz
- & Martin Jinek
-
Article
| Open Access Transposon-associated TnpB is a programmable RNA-guided DNA endonuclease
Transposon-associated TnpB is a programmable RNA-guided DNA endonuclease
The RNA-directed nuclease TnpB from Deinococcus radiodurans can be reprogrammed to cleave DNA target sites in human cells.
- Tautvydas Karvelis
- , Gytis Druteika
- & Virginijus Siksnys
-
Review Article |
 The role of retrotransposable elements in ageing and age-associated diseases
The role of retrotransposable elements in ageing and age-associated diseases
This Review discusses how the activity of retrotransposons influences ageing and the role of these mobile genetic elements in age-related diseases and their treatment.
- Vera Gorbunova
- , Andrei Seluanov
- & John M. Sedivy
-
Article |
 Structural basis of DNA targeting by a transposon-encoded CRISPR–Cas system
Structural basis of DNA targeting by a transposon-encoded CRISPR–Cas system
Cryo-electron microscopy structures of the TniQ–Cascade complex encoded by the Vibrio cholerae Tn6677 transposon reveal the mechanistic basis of the functional association of CRISPR- and transposon-associated machineries.
- Tyler S. Halpin-Healy
- , Sanne E. Klompe
- & Israel S. Fernández
-
Article |
 Transposon-encoded CRISPR–Cas systems direct RNA-guided DNA integration
Transposon-encoded CRISPR–Cas systems direct RNA-guided DNA integration
A programmable transposase integrates donor DNA at user-defined genomic target sites with high fidelity, revealing a new approach for genetic engineering that obviates the need for DNA double-strand breaks and homologous recombination.
- Sanne E. Klompe
- , Phuc L. H. Vo
- & Samuel H. Sternberg
-
Article |
 L1 drives IFN in senescent cells and promotes age-associated inflammation
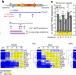
L1 drives IFN in senescent cells and promotes age-associated inflammation
During cellular senescence in human and mouse cells, L1 transposons become transcriptionally derepressed and trigger a type-1 interferon response, which contributes to age-associated inflammation and age-related phenotypes.
- Marco De Cecco
- , Takahiro Ito
- & John M. Sedivy
-
Letter |
 Selective silencing of euchromatic L1s revealed by genome-wide screens for L1 regulators
Selective silencing of euchromatic L1s revealed by genome-wide screens for L1 regulators
The retrotransposition of L1 is controlled by functionally diverse genes at the transcriptional or post-transcriptional levels, and its silencing can lead to the downregulation of host gene expression.
- Nian Liu
- , Cameron H. Lee
- & Joanna Wysocka
-
Brief Communications Arising |
 Elsässer et al. reply
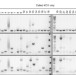
Elsässer et al. reply
- Simon J. Elsässer
- , Kyung-Min Noh
- & Laura A. Banaszynski
-
Brief Communications Arising |
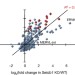 On the role of H3.3 in retroviral silencing
On the role of H3.3 in retroviral silencing
- Gernot Wolf
- , Rita Rebollo
- & Todd S. Macfarlan
-
Letter |
 Histone H3.3 is required for endogenous retroviral element silencing in embryonic stem cells
Histone H3.3 is required for endogenous retroviral element silencing in embryonic stem cells
Transposable elements in mammalian genomes need to be silenced to avoid detrimental genome instability; here, the histone variant H3.3 is shown to have an important role in silencing endogenous retroviral elements.
- Simon J. Elsässer
- , Kyung-Min Noh
- & Laura A. Banaszynski
-
Letter |
 An evolutionary arms race between KRAB zinc-finger genes ZNF91/93 and SVA/L1 retrotransposons
An evolutionary arms race between KRAB zinc-finger genes ZNF91/93 and SVA/L1 retrotransposons
The authors show that two primate-specific genes encoding KRAB domain containing zinc finger proteins, ZNF91 and ZNF93, have evolved during the last 25 million years to repress retrotransposon families that emerged during this time period; according to the new data KZNF gene expansion limits the activity of newly emerged retrotransposons, which subsequently mutate to evade repression.
- Frank M. J. Jacobs
- , David Greenberg
- & David Haussler
-
Article |
 The Mu transpososome structure sheds light on DDE recombinase evolution
The Mu transpososome structure sheds light on DDE recombinase evolution
The structure of the bacteriophage transposase MuA bound to DNA sequences that mimic both the transposon ends and the target DNA ends is solved; the picture of this synaptic complex illustrates the intricacy of Mu transposition, and exposes the architectural diversity among DDE recombinases in complex with substrate DNAs.
- Sherwin P. Montaño
- , Ying Z. Pigli
- & Phoebe A. Rice
-
Letter |
 An siRNA pathway prevents transgenerational retrotransposition in plants subjected to stress
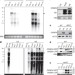
An siRNA pathway prevents transgenerational retrotransposition in plants subjected to stress
The transcription of repetitive elements such as retrotransposons is normally repressed, to prevent their unchecked dissemination throughout the genome. This study shows that heat stress induces the transcription of the ONSEN retroelement. The accumulation of ONSEN is suppressed by small interfering RNAs. When siRNAs were absent, new ONSEN insertions were found in the progeny of heat-stressed plants, having transposed during differentiation. These results imply a memory of stress that can be counteracted by siRNAs.
- Hidetaka Ito
- , Hervé Gaubert
- & Jerzy Paszkowski
-
Letter |
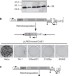 Epigenetic silencing of engineered L1 retrotransposition events in human embryonic carcinoma cells
Epigenetic silencing of engineered L1 retrotransposition events in human embryonic carcinoma cells
The ability of retrotransposons to mobilize and insert into genes presents a challenge to a cell needing to maintain its genomic integrity. These authors have studied retrotransposition in embryonic carcinoma-derived cells. On insertion into DNA, the retrotransposon is quickly silenced, but the retrotransposon-specificity of this process implies that multiple silencing mechanisms may exist. Once cells differentiate, the ability to silence newly introduced retrotransposons is lost but previously inactivated retrotransposons remain inactive.
- Jose L. Garcia-Perez
- , Maria Morell
- & John V. Moran
-
Research Highlights |
Genomics: Transposition trends
 Autonomous transposons tune their sequences to ensure somatic suppression
Autonomous transposons tune their sequences to ensure somatic suppression