Article
|
Open Access
Featured
-
-
Article
| Open Access Translation selectively destroys non-functional transcription complexes
Translation selectively destroys non-functional transcription complexes
Translation actively dislodges stalled transcription elongation complexes (ECs) from damaged DNA, which enables lesion repair and restoration of transcription activity, and coupled ribosomes discriminate between active ECs and stalled ECs, ensuring destruction of only the latter.
- Jason Woodgate
- , Hamed Mosaei
- & Nikolay Zenkin
-
Article
| Open Access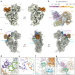 Structural basis of ribosomal 30S subunit degradation by RNase R
Structural basis of ribosomal 30S subunit degradation by RNase R
Cryo-electron microscopy structures of intermediates formed during the degradation of the 30S ribosomal unit shed light on how the 3′ to 5′ exonuclease ribonuclease R controls the ribosomal degradation process.
- Lyudmila Dimitrova-Paternoga
- , Sergo Kasvandik
- & Helge Paternoga
-
Article
| Open Access Adding α,α-disubstituted and β-linked monomers to the genetic code of an organism
Adding α,α-disubstituted and β-linked monomers to the genetic code of an organism
tRNA display enables the direct selection of orthogonal aminoacyl-tRNA synthetases that acylate orthogonal tRNAs with non-canonical monomers, enabling in vivo synthesis of proteins that include these monomers and expanding the repertoire of the genetic code.
- Daniel L. Dunkelmann
- , Carlos Piedrafita
- & Jason W. Chin
-
Article |
 mRNA reading frame maintenance during eukaryotic ribosome translocation
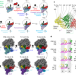
mRNA reading frame maintenance during eukaryotic ribosome translocation
The accuracy of eukaryotic ribosome translocation relies on eukaryote-specific elements of the 80S ribosome, elongation factor 2 and transfer RNAs, all of which contribute to the maintenance of the messenger RNA reading frame.
- Nemanja Milicevic
- , Lasse Jenner
- & Gulnara Yusupova
-
Article |
 Evolutionarily divergent mTOR remodels translatome for tissue regeneration
Evolutionarily divergent mTOR remodels translatome for tissue regeneration
Rapid activation of protein synthesis in the axolotl highlights the unanticipated impact of a translatome on orchestrating the early steps of wound healing and provides a missing link in our understanding of vertebrate regenerative potential.
- Olena Zhulyn
- , Hannah D. Rosenblatt
- & Maria Barna
-
Article
| Open Access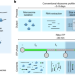 Single-cell quantification of ribosome occupancy in early mouse development
Single-cell quantification of ribosome occupancy in early mouse development
A single-cell ribosome profiling method can provide data at the level of allele-specific ribosome engagement in early development.
- Hakan Ozadam
- , Tori Tonn
- & Can Cenik
-
Article |
 Nuclear export of pre-60S particles through the nuclear pore complex
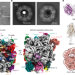
Nuclear export of pre-60S particles through the nuclear pore complex
We report the cryo-electron microscopy structure of native pre-60S particles trapped in the channel of the yeast nuclear pore complex, suggesting a translocation model for the export of pre-60S particles through the complex.
- Zongqiang Li
- , Shuaijiabin Chen
- & Sen-Fang Sui
-
Article
| Open Access Engineered tRNAs suppress nonsense mutations in cells and in vivo
Engineered tRNAs suppress nonsense mutations in cells and in vivo
Suppressor tRNAs adapted to the amino acid that they carry enable readthrough of premature termination codons introduced by nonsense mutations and show potential for the treatment of genetic diseases such as cystic fibrosis.
- Suki Albers
- , Elizabeth C. Allen
- & Zoya Ignatova
-
Article |
 Alternative CDC20 translational isoforms tune mitotic arrest duration
Alternative CDC20 translational isoforms tune mitotic arrest duration
Human cells modulate the duration of their mitotic arrest through the presence of conserved alternative CDC20 translational isoforms.
- Mary-Jane Tsang
- & Iain M. Cheeseman
-
Article |
 Noncoding translation mitigation
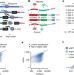
Noncoding translation mitigation
Combining genome-wide CRISPR screens with massively parallel analyses of human and random DNA sequences reveal a unified mechanism for the surveillance and evolution of translation products from annotated noncoding DNA.
- Jordan S. Kesner
- , Ziheng Chen
- & Xuebing Wu
-
Article
| Open Access mRNA decoding in human is kinetically and structurally distinct from bacteria
mRNA decoding in human is kinetically and structurally distinct from bacteria
The reaction coordinate of aminoacyl-tRNA movement is altered on the human ribosome and the process is an order of magnitude slower compared with bacteria due to eukaryote-specific structural elements in the human ribosome and in the elongation factor eEF1A.
- Mikael Holm
- , S. Kundhavai Natchiar
- & Scott C. Blanchard
-
Article |
 A molecular network of conserved factors keeps ribosomes dormant in the egg
A molecular network of conserved factors keeps ribosomes dormant in the egg
Mass spectrometry and structural studies demonstrate the specific changes in protein composition that accompany the transition of ribosomes in zebrafish and Xenopus eggs from a dormant to an active state during early embryogenesis.
- Friederike Leesch
- , Laura Lorenzo-Orts
- & Andrea Pauli
-
Article |
 Short tRNA anticodon stem and mutant eRF1 allow stop codon reassignment
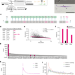
Short tRNA anticodon stem and mutant eRF1 allow stop codon reassignment
Analyses of in-frame stop codons in protein-coding genes of Blastocrithidia nonstop with all three stop codons reassigned reveal a mechanism for UGA reassignment in eukaryotes involving shortening of the tRNA anticodon stem and a mutant eRF1 release factor.
- Ambar Kachale
- , Zuzana Pavlíková
- & Julius Lukeš
-
Article |
 Structural basis of regulated m7G tRNA modification by METTL1–WDR4
Structural basis of regulated m7G tRNA modification by METTL1–WDR4
Structures of the human METTL1–WDR4 complex are revealed, providing molecular insights into substrate recognition, modification and catalytic regulation by the N7-methylguanosine methyltransferase complex.
- Jiazhi Li
- , Longfei Wang
- & Richard I. Gregory
-
Article |
 A male germ-cell-specific ribosome controls male fertility
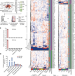
A male germ-cell-specific ribosome controls male fertility
RibosomeST—a ribosome with a specialized nascent polypeptide exit tunnel—cotranslationally regulates the folding of a subset of male germ-cell-specific proteins that are essential for the formation of sperm.
- Huiling Li
- , Yangao Huo
- & Jiahao Sha
-
Article
| Open Access Visualizing translation dynamics at atomic detail inside a bacterial cell
Visualizing translation dynamics at atomic detail inside a bacterial cell
Cryo-electron tomography is used to reveal the structural dynamics and functional diversity of translating ribosomes in Mycoplasma pneumoniae, providing insight into the translation elongation cycle inside cells and how it is reshaped by antibiotics.
- Liang Xue
- , Swantje Lenz
- & Julia Mahamid
-
Article |
 eIF5B and eIF1A reorient initiator tRNA to allow ribosomal subunit joining
eIF5B and eIF1A reorient initiator tRNA to allow ribosomal subunit joining
Single-molecule spectroscopy and structural studies were used to examine the dynamics of association of eIF1A and eIF5B with the human translation initiation complex and their role in presenting tRNA to the complex to initiate translation.
- Christopher P. Lapointe
- , Rosslyn Grosely
- & Joseph D. Puglisi
-
Article
| Open Access Mechanism of mitoribosomal small subunit biogenesis and preinitiation
Mechanism of mitoribosomal small subunit biogenesis and preinitiation
Structural analysis of several small mitoribosomal subunit intermediates reveals a sequential mechanism of biogenesis, and how assembly links to initiation to form active mitoribosomes.
- Yuzuru Itoh
- , Anas Khawaja
- & Alexey Amunts
-
Article
| Open Access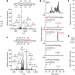 Reversible RNA phosphorylation stabilizes tRNA for cellular thermotolerance
Reversible RNA phosphorylation stabilizes tRNA for cellular thermotolerance
Reversible internal RNA phosphrylation contributes to thermal stability and nuclease resistance of tRNA, and cellular thermotolerance of hyperthermophiles.
- Takayuki Ohira
- , Keiichi Minowa
- & Tsutomu Suzuki
-
Matters Arising |
 Evaluating ribosomal frameshifting in CCR5 mRNA decoding
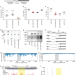
Evaluating ribosomal frameshifting in CCR5 mRNA decoding
- Yousuf A. Khan
- , Gary Loughran
- & John F. Atkins
-
Article |
 Ribosome collisions induce mRNA cleavage and ribosome rescue in bacteria
Ribosome collisions induce mRNA cleavage and ribosome rescue in bacteria
In bacteria, a newly identified endonuclease activated by ribosome collisions truncates mRNA to trigger rescue of stalled ribosomes.
- Kazuki Saito
- , Hanna Kratzat
- & Allen R. Buskirk
-
Article |
 Bacterial ribosome collision sensing by a MutS DNA repair ATPase paralogue
Bacterial ribosome collision sensing by a MutS DNA repair ATPase paralogue
Bacterial MutS2, a paralogue of the DNA mismatch-repair protein MutS, is found to bind collided ribosomes and function in translational quality control.
- Federico Cerullo
- , Sebastian Filbeck
- & Claudio A. P. Joazeiro
-
Article |
 Ageing exacerbates ribosome pausing to disrupt cotranslational proteostasis
Ageing exacerbates ribosome pausing to disrupt cotranslational proteostasis
Ageing alters the kinetics of translation elongation in both Caenorhabditis elegans and Saccharomyces cerevisiae.
- Kevin C. Stein
- , Fabián Morales-Polanco
- & Judith Frydman
-
Article |
 Valine tRNA levels and availability regulate complex I assembly in leukaemia
Valine tRNA levels and availability regulate complex I assembly in leukaemia
Restriction of dietary valine reduces growth of T cell acute lymphoblastic leukaemia through altered valine tRNA biogenesis and reduced translation of mRNAs that encode subunits of mitochondrial complex I.
- Palaniraja Thandapani
- , Andreas Kloetgen
- & Iannis Aifantis
-
Article
| Open Access Accuracy mechanism of eukaryotic ribosome translocation
Accuracy mechanism of eukaryotic ribosome translocation
Structural analysis of the Saccharomyces cerevisiae 80S ribosome trapped in an intermediate translocation state shows stabilization of codon–anticodon interactions by eukaryote-specific elements of the 80S ribosome, eEF2 and tRNA and demonstrates a major role for eEF2 in maintaining the directionality of translocation.
- Muminjon Djumagulov
- , Natalia Demeshkina
- & Gulnara Yusupova
-
Article |
 Single-cell Ribo-seq reveals cell cycle-dependent translational pausing
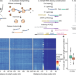
Single-cell Ribo-seq reveals cell cycle-dependent translational pausing
Highly sensitive ribosome profiling of single cells at single-codon resolution enables identification of distinct cell cycle-dependent translational dynamic states in individual cells.
- Michael VanInsberghe
- , Jeroen van den Berg
- & Alexander van Oudenaarden
-
Article |
 Cotranslational prolyl hydroxylation is essential for flavivirus biogenesis
Cotranslational prolyl hydroxylation is essential for flavivirus biogenesis
A proteomics-based strategy is used to examine how three different RNA viruses (polio, Zika and dengue) remodel translation in the host to recruit host machineries necessary for the production of viral proteins.
- Ranen Aviner
- , Kathy H. Li
- & Raul Andino
-
Article
| Open Access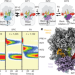 Structural basis of early translocation events on the ribosome
Structural basis of early translocation events on the ribosome
Cryo-electron microscopy and single-molecule fluorescence methods are used to elucidate the mechanism of early translocation events on the bacterial ribosome.
- Emily J. Rundlet
- , Mikael Holm
- & Scott C. Blanchard
-
Article |
 MIR-NATs repress MAPT translation and aid proteostasis in neurodegeneration
MIR-NATs repress MAPT translation and aid proteostasis in neurodegeneration
The natural antisense transcript MAPT-AS1 interferes with translation of mRNA transcript into tau protein in the brain and may represent a general mechanism for controlling levels of intrinsically disordered proteins, with particular relevance for neurodegeneration.
- Roberto Simone
- , Faiza Javad
- & Rohan de Silva
-
Article |
 Anti-tumour immunity induces aberrant peptide presentation in melanoma
Anti-tumour immunity induces aberrant peptide presentation in melanoma
Tryptophan depletion in melanoma cells after prolonged treatment with interferon-γ (IFNγ) results in ribosomal frameshifting and the production of aberrant peptides that can be presented to T cells and induce an immune response.
- Osnat Bartok
- , Abhijeet Pataskar
- & Reuven Agami
-
Article |
 Structural basis for the final steps of human 40S ribosome maturation
Structural basis for the final steps of human 40S ribosome maturation
Studies of five cryo-electron microscopy structures reveal the composition and conformational progression in the final maturation events of human 40S ribosomal subunit assembly.
- Michael Ameismeier
- , Ivo Zemp
- & Roland Beckmann
-
Article |
 The coding capacity of SARS-CoV-2
The coding capacity of SARS-CoV-2
A high-resolution map of coding regions in the SARS-CoV-2 genome enables the identification of 23 unannotated open reading frames and quantification of the expression of canonical viral open reading frames.
- Yaara Finkel
- , Orel Mizrahi
- & Noam Stern-Ginossar
-
Article |
 Functionally uncoupled transcription–translation in Bacillus subtilis
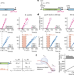
Functionally uncoupled transcription–translation in Bacillus subtilis
In Bacillus subtilis, unlike in Escherichia coli, transcription and translation of genes are not tightly coupled, and pioneering ribosomes lag substantially behind RNA polymerases.
- Grace E. Johnson
- , Jean-Benoît Lalanne
- & Gene-Wei Li
-
Article |
 Cryo-EM of elongating ribosome with EF-Tu•GTP elucidates tRNA proofreading
Cryo-EM of elongating ribosome with EF-Tu•GTP elucidates tRNA proofreading
Time-resolved cryogenic electron microscopy structures of a ribosome during the delivery of aminoacyl-tRNA by EF-Tu•GTP capture 33 ribosomal states, enabling visualization of the initial selection, proofreading and peptidyl transfer stages.
- Anna B. Loveland
- , Gabriel Demo
- & Andrei A. Korostelev
-
Article |
 DNA-PKcs has KU-dependent function in rRNA processing and haematopoiesis
DNA-PKcs has KU-dependent function in rRNA processing and haematopoiesis
The catalytic subunit of DNA-PK autophosphorylates and contributes to ribosome biogenesis and haematopoiesis by binding to the U3 small nucleolar RNA.
- Zhengping Shao
- , Ryan A. Flynn
- & Eliezer Calo
-
Letter |
 eIF5B gates the transition from translation initiation to elongation
eIF5B gates the transition from translation initiation to elongation
Single-molecule dynamics reveal that the GTPase activity of eukaryotic initiation factor eIF5B serves as a kinetic checkpoint for the transition from translation initiation to elongation.
- Jinfan Wang
- , Alex G. Johnson
- & Joseph D. Puglisi
-
Letter |
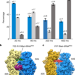 Late steps in bacterial translation initiation visualized using time-resolved cryo-EM
Late steps in bacterial translation initiation visualized using time-resolved cryo-EM
A time-resolved cryo-electron microscopy approach is used to visualize, at near-atomic resolution and on a sub-second timescale, short-lived intermediate states of a fundamental biomolecular reaction.
- Sandip Kaledhonkar
- , Ziao Fu
- & Joachim Frank
-
Article |
 Total synthesis of Escherichia coli with a recoded genome
Total synthesis of Escherichia coli with a recoded genome
High-fidelity convergent total synthesis is used to produce
Escherichia coli with a 61-codon synthetic genome that uses 59 codons to encode all of the canonical amino acids.- Julius Fredens
- , Kaihang Wang
- & Jason W. Chin
-
Letter |
 The translation of non-canonical open reading frames controls mucosal immunity
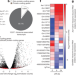
The translation of non-canonical open reading frames controls mucosal immunity
In mouse macrophages, a range of short and non-ATG-initiated open reading frames that can generate proteins are identified, one of which is shown to be essential for host immunity to enteric mucosal infection and inflammation.
- Ruaidhrí Jackson
- , Lina Kroehling
- & Richard A. Flavell
-
Letter |
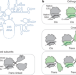 Controlling orthogonal ribosome subunit interactions enables evolution of new function
Controlling orthogonal ribosome subunit interactions enables evolution of new function
Orthogonal ribosomes are engineered in which the two subunits are stapled together in a way that limits association with endogenous subunits in cells, enabling the evolution of new functionality in the orthogonal ribosome.
- Wolfgang H. Schmied
- , Zakir Tnimov
- & Jason W. Chin
-
Letter |
 m6A facilitates hippocampus-dependent learning and memory through YTHDF1
m6A facilitates hippocampus-dependent learning and memory through YTHDF1
Neuronal stimulation induces protein translation of m6A-methylated neuronal mRNAs facilitated by YTHDF1, and this process contributes to learning and memory.
- Hailing Shi
- , Xuliang Zhang
- & Tao Zhou
-
Letter |
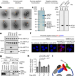 mRNA circularization by METTL3–eIF3h enhances translation and promotes oncogenesis
mRNA circularization by METTL3–eIF3h enhances translation and promotes oncogenesis
METTL3, the enzyme responsible for m6A modification, influences translation by interacting with eIF3h to mediate looping between the regions near the stop codon and 5′ cap of mRNA.
- Junho Choe
- , Shuibin Lin
- & Richard I. Gregory
-
Letter |
 Cotranslational assembly of protein complexes in eukaryotes revealed by ribosome profiling
Cotranslational assembly of protein complexes in eukaryotes revealed by ribosome profiling
Cotranslational assembly is a prevalent mechanism for the formation of oligomeric complexes in Saccharomyces cerevisiae, with one subunit serving as scaffold for the translation of partner subunits.
- Ayala Shiber
- , Kristina Döring
- & Bernd Bukau
-
Article |
 Autism-like phenotype and risk gene mRNA deadenylation by CPEB4 mis-splicing
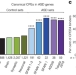
Autism-like phenotype and risk gene mRNA deadenylation by CPEB4 mis-splicing
CPEB4 binds the mRNA of genes known to be associated with autism and shows an isoform imbalance in individuals with autism, and an equivalent imbalance in mice induces an autism-like phenotype.
- Alberto Parras
- , Héctor Anta
- & José J. Lucas
-
Letter |
 Unique features of mammalian mitochondrial translation initiation revealed by cryo-EM
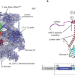
Unique features of mammalian mitochondrial translation initiation revealed by cryo-EM
A cryo-electron microscopy structure of the mammalian mitochondrial translation initiation complex reveals unique features that are necessary to initiate mitochondrial translation and highlights a possible membrane-targeting peptide.
- Eva Kummer
- , Marc Leibundgut
- & Nenad Ban
-
Letter |
 The helicase Ded1p controls use of near-cognate translation initiation codons in 5′ UTRs
The helicase Ded1p controls use of near-cognate translation initiation codons in 5′ UTRs
The helicase Ded1p associates with the pre-initiation complex and influences translation from near-cognate initiation codons by controlling the levels of mRNA secondary structure in 5′ untranslated regions.
- Ulf-Peter Guenther
- , David E. Weinberg
- & Eckhard Jankowsky
-
Letter |
 Codon-specific translation reprogramming promotes resistance to targeted therapy
Codon-specific translation reprogramming promotes resistance to targeted therapy
Enzymes that catalyse modifications of wobble uridine 34 tRNA are essential for the survival of melanoma cells that rely on HIF1α-dependent metabolism through codon-dependent regulation of the translation of HIF1A mRNA.
- Francesca Rapino
- , Sylvain Delaunay
- & Pierre Close
-
Article |
 Visualizing late states of human 40S ribosomal subunit maturation
Visualizing late states of human 40S ribosomal subunit maturation
Cryo-EM structures of late intermediates in the assembly of human 40S ribosomal subunits help to define the principles by which immature rRNA conformations and ribosomal biogenesis factors shape the 40S maturation process.
- Michael Ameismeier
- , Jingdong Cheng
- & Roland Beckmann
-
Article |
 ANKRD16 prevents neuron loss caused by an editing-defective tRNA synthetase
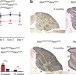
ANKRD16 prevents neuron loss caused by an editing-defective tRNA synthetase
ANKRD16 attenuates neurodegeneration induced by a mutation in the editing domain of alanyl tRNA synthetase by directly accepting mis-activated serine from the synthetase before transfer to the tRNA, establishing a new mechanism by which editing defects are prevented.
- My-Nuong Vo
- , Markus Terrey
- & Susan L. Ackerman

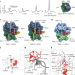 A new family of bacterial ribosome hibernation factors
A new family of bacterial ribosome hibernation factors