« Prev Next »
Say you are studying three individuals who carry the same disease-causing mutation; two of these individuals suffer from the disease but exhibit different symptoms, while the third person is completely unaffected. How would you explain this phenomenon? For many inherited diseases, the same mutation is not always expressed in all individuals who carry it; moreover, when the mutation is expressed, it is not always expressed in the same way. These findings are the basis for the concepts of penetrance and expressivity. Penetrance measures the proportion of a population of individuals who carry a disease-causing allele and express the related disease phenotype. Expressivity measures the extent to which a genotype shows its phenotypic expression. Moreover, both the penetrance and expressivity of different diseases can be partially explained by the action of gene modifiers.
Penetrance
Retinoblastoma is a cancer of the retina that primarily affects children and is caused by mutations in the Rb gene. Interestingly, not all people who carry this mutation suffer from retinoblastoma. For instance, two sisters could inherit the same mutation from their parents, and one might be affected by the disease, while the other is not (OMIM, 2008d). Perhaps even more interestingly, retinoblastoma is not alone in this regard; there are numerous other inherited diseases in which the same mutation is not expressed in all the individuals who carry it. These findings illustrate a concept that geneticists refer to as penetrance.
Penetrance is a measurement of the proportion of individuals in a population who carry a disease-causing allele and express the disease phenotype. In the case of retinoblastoma, because not all individuals who carry the mutation are affected by the disease, this condition is said to display incomplete or reduced penetrance. Other diseases with incomplete penetrance include Huntington's disease and breast cancer. In contrast to these examples, a disease shows complete penetrance if all of the individuals who carry the related gene are affected by the disease. For instance, the inheritance of a specific point mutation in the fibroblast growth factor receptor 3 gene (FGFR3) always results in the disorder achondroplasia, which is characterized by abnormal bone growth and dwarfism (OMIM, 2008a).
Expressivity
While measures of penetrance focus on whether or not a disease is expressed in a population, the degree to which a genotype is phenotypically expressed in individuals can also be measured. For instance, one patient who is suffering from a disease may have more severe symptoms than another patient who carries the same mutated allele. Meanwhile, a third patient who also has the mutated allele may seem almost normal. This phenomenon is described as variable expressivity. Expressivity measures the extent to which a genotype exhibits its phenotypic expression.
Just as individuals with a certain mutation can exhibit differences in disease severity, they can also show differences in many other aspects of their phenotypes, such as their age of onset of disease symptoms. Some diseases, such as Huntington's disease, may have an earlier onset with more severe symptoms in subsequent generations. This type of variable expressivity is called anticipation.
To better understand the concept of expressivity, it is helpful to consider a few diseases that display a range of phenotypes, including neurofibromatosis, holoprosencephaly, and Van der Woude syndrome.
Neurofibromatosis
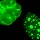
Neurofibromatosis is a disease caused by mutations in the neurofibromin gene (OMIM, 2008b). These mutations can cause the Schwann cells in an affected individual's nervous system to grow into tumors called neurofibromas, which appear as café-au-lait colored spots or bumps under the skin. These tumors can result in skeletal abnormalities and neurological problems. However, not all people who have the mutated neurofibromin gene are equally affected by this condition. Research has shown that family members who carry the same mutated gene can exhibit a range of symptoms, with some individuals experiencing much more severe symptoms than others, although they all carry the same allele.
Holoprosencephaly
Holoprosencephaly is a condition in which the embryonic forebrain does not correctly divide into two lobes (OMIM, 2008c). To date, scientists have described nine types of holoprosencephaly caused by deletions or duplications in various genes and chromosomal regions.
In 1993, researcher Amanda Collins and her colleagues studied a single large family with multiple children affected with holoprosencephaly, and they noted variable expressivity of the disease in the affected family members (Figure 1; Collins et al., 1993). In the most severe cases, holoprosencephaly resulted in death. In contrast, in the mild cases, the family members had near-normal to normal brain development and only minor facial deformities, such as a single upper central incisor (Collins et al., 1993). Other studies have shown that this disease also displays incomplete penetrance (Muenke et al., 1994).
Van der Woude Syndrome
Van der Woude syndrome is a dominantly inherited developmental disorder that causes cleft lip and/or cleft palate (OMIM, 2008e). This condition results from mutations in a single gene that encodes for a protein called interferon regulatory factor 6. Studying a large family of seven generations affected with Van der Woude syndrome, Janku et al. noted that the disorder displayed both incomplete penetrance and variable expressivity. Sometimes, individuals who did not realize they were affected were diagnosed with mild versions of the disorder upon careful examination. Most patients had characteristic lip pits, which were often observed alone and sometimes with other symptoms. Meanwhile, lip and palate clefts were observed in around one-fifth of individuals (Janku et al., 1980).
Why Do Phenotypes Show Differences in Penetrance and Expressivity?
So, how does a scientist relate differences in penetrance and expressivity that are observed at the phenotypic level to changes at the molecular level? For example, how is it possible that one family member carrying a retinoblastoma mutation has the disease, while another carrying the same mutation does not? What accounts for such a difference in disease penetrance? In the case of neurofibromatosis, family members carrying the same mutated neurofibromin gene are unequally affected by the condition, with some family members showing more severe symptoms than others. Why does this disease show variable expressivity? Answering these questions is not an easy task. Nonetheless, research has shown that variable phenotypes can be caused by a number of factors, including the following:
- Modifier genes
- Environmental factors
- Allelic variation
- Complex genetic and environmental interactions
In most cases in which a particular genotype is inherited, it is not fully known why the same allele can cause subtly different or profoundly different phenotypes. In some cases, however, there is genetic evidence that modifier genes influence phenotypic variation.
Modifier genes can affect penetrance, dominance, and expressivity. A genetic modifier, when expressed, is able to alter the expression of another gene. Modifier genes can affect transcription and alter the immediate gene transcript expression, or they can affect phenotypes at other levels of organization by altering phenotypes at the cellular or organismal level (Nadeau, 2001).
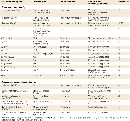
Some examples of modifier genes identified in mice and humans, along with their modifier effects and phenotypic consequences, are shown in Table 1. As you can see from the table, many more modifiers have been identified in mice than in humans because of the ability to perform gene targeting experiments on inbred mouse strains. For example, an unknown modifier associated with specific mouse genetic backgrounds can alter the penetrance of an allele (called disorganization), which causes birth defects. Likewise, another unknown modifier that is associated with specific mouse genetic backgrounds affects the expressivity of an allele (called brachyury), resulting in mice with different tail lengths. A third modifier gene, named dsu, suppresses the expression of a number of different coat color alleles in mice, including ashen, leaden, ruby-eye, and ruby-eye-2.
To better understand the role of modifier genes in humans, Saima Riazuddin and her colleagues studied 141 members of a family afflicted with isolated deafness (i.e., deafness was the only clinical finding in this family), which is caused by the DFNB26 gene. The researchers noticed that the disease was incompletely penetrant. While most individuals who were homozygous for the gene were deaf, seven homozygous family members had normal hearing. Through linkage studies, the scientists identified a dominant modifier, called DFNM1, which suppressed deafness in homozygous individuals (Riazuddin et al., 2000). It is not yet known how this modifier gene suppresses deafness. It may suppress the mutant gene's expression or prevent deafness through another mechanism.
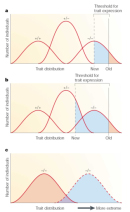
A model of modification shows different ways that modifier genes can alter the phenotypes in an organism (Figure 2; Nadeau, 2001). As shown in Figure 2, a modifier gene can move the threshold for trait expression, which means that the modifier causes a smaller or greater proportion of individuals to express the disease. In this way, a modifier gene can change the level of penetrance. For example, the modifier gene DFNM1 moves the threshold for trait expression to the right, so that some individuals homozygous for the DFNB26 allele are not deaf, which results in incomplete penetrance. Meanwhile, other modifiers can increase the proportion of individuals affected by a disease-causing allele by decreasing the threshold for trait expression (Figures 2a and 2b). Modifiers may also shift the trait distribution, or the range of disease phenotypes, which causes more individuals carrying a disease-causing allele to express a more (or less) severe disease phenotype (Figure 2c). This process results in variable expressivity.
For example, patients with thalassemia, a disorder caused by defective beta-globin synthesis, have diverse clinical characteristics and variable expressivity. Patients with severe cases have profound anemia and require regular blood transfusions, while other individuals who carry the same allele have mild and undetectable symptoms. A number of factors underlie this phenotypic diversity, including the involvement of numerous modifier genes at other genetic loci that affect the production of globins (Weatherall, 2001).
As with thalassemia, other human diseases also vary according to genetic and environmental factors, which can lead to both incomplete penetrance and variable expressivity. In fact, for most diseases, variable expressivity of the disease phenotype is the norm among individuals who carry the same disease-causing allele or alleles (Nadeau, 2001), though the causes are not always clear. Thus, the identification of additional modifier genes will help scientists better understand the nature of a wide range of human diseases.
References and Recommended Reading
Collins, A. L., et al. Holoprosencephaly: A family showing dominant inheritance and variable expression. Journal of Medical Genetics 30, 36–40 (1993)
Janku, P., et al. The Van der Woude syndrome in a large kindred: Variability, penetrance, genetic risks. American Journal of Medical Genetics 5, 117–123 (1980)
Muenke, M., et al. Linkage of a human brain malformation, familial holoprosencephaly, to chromosome 7 and evidence for genetic heterogeneity. Proceedings of the National Academy of Sciences 91, 8102–8106 (1994)
Nadeau, J. H. Modifier genes in mice and humans. Nature Reviews Genetics 2, 165–174 (2001) doi:10.1038/35056009 (link to article)
OMIM. "Achondroplasia; ACH." Johns Hopkins University (2008a) (accessed Aug. 28, 2008)
OMIM. "Neurofibromatosis, type I; NF1." Johns Hopkins University (2008b) (accessed May 24, 2008)
OMIM. "Holoprosencephaly." Johns Hopkins University (2008c) (accessed on Aug. 28, 2008)
OMIM. "Retinoblastoma; RB1." Johns Hopkins University (2008d) (accessed on Aug. 28, 2008)
OMIM. "Van der Woude syndrome." Johns Hopkins University (2008e) (accessed on Aug. 28, 2008)
Riazuddin, S., et al. Dominant modifier DFNM1 suppresses recessive deafness DFNB26. Nature Genetics 26, 431–434 (2000) doi:10.1038/82558 (link to article)
Weatherall, D. J. Phenotype-genotype relationships in monogenic disease: Lessons from the thalassaemias. Nature Reviews Genetics 2, 245–255 (2001) doi:10.1038/35066048 (link to article)



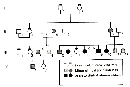
 Figure 1
Figure 1