Article
|
Open Access
Featured
-
-
Research Briefing |
 Simplifying deep learning to enhance accessibility of large-scale 3D brain imaging analysis
Simplifying deep learning to enhance accessibility of large-scale 3D brain imaging analysis
We created DELiVR, a deep-learning pipeline for 3D brain-cell mapping that is trained with virtual reality-generated reference annotations. It can be deployed via the user-friendly interface of the open-source software Fiji, which makes the analysis of large-scale 3D brain images widely accessible to scientists without computational expertise.
-
Article |
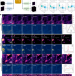 Pretraining a foundation model for generalizable fluorescence microscopy-based image restoration
Pretraining a foundation model for generalizable fluorescence microscopy-based image restoration
A pretrained foundation model (UniFMIR) enables versatile and generalizable performance across diverse fluorescence microscopy image reconstruction tasks.
- Chenxi Ma
- , Weimin Tan
- & Bo Yan
-
Article |
 Selective-plane-activation structured illumination microscopy
Selective-plane-activation structured illumination microscopy
The combination of light sheet illumination and reversibly switchable fluorophores enables improved structured illumination microscopy for fast, low-background super-resolution imaging in cells and spheroids.
- Kenta Temma
- , Ryosuke Oketani
- & Katsumasa Fujita
-
Brief Communication
| Open Access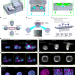 Open-top multisample dual-view light-sheet microscope for live imaging of large multicellular systems
Open-top multisample dual-view light-sheet microscope for live imaging of large multicellular systems
This work presents a highly versatile open-top, dual-view and dual-illumination light-sheet microscope for live imaging of large specimens.
- Franziska Moos
- , Simon Suppinger
- & Prisca Liberali
-
Brief Communication
| Open Access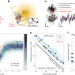 MINSTED tracking of single biomolecules
MINSTED tracking of single biomolecules
MINSTED quantifies tiny movements of individual biomolecules with high spatiotemporal precision to successfully resolve the steps of the molecular motor protein kinesin-1 labeled with a single fluorophore as it switches protofilaments.
- Lukas Scheiderer
- , Henrik von der Emde
- & Stefan W. Hell
-
-
Article
| Open Access An optogenetic method for the controlled release of single molecules
An optogenetic method for the controlled release of single molecules
An optogenetic system enables the controlled release of soluble and transmembrane proteins for precise exploration of cellular protein function at the single-molecule level and streamlined single-molecule imaging.
- Purba Kashyap
- , Sara Bertelli
- & Helge Ewers
-
Article
| Open Access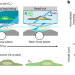 Super-sectioning with multi-sheet reversible saturable optical fluorescence transitions (RESOLFT) microscopy
Super-sectioning with multi-sheet reversible saturable optical fluorescence transitions (RESOLFT) microscopy
Multi-sheet RESOLFT combines the speed and optical sectioning of light-sheet fluorescence microscopy with reversibly photoswitchable fluorescent proteins to enable fast, volumetric super-resolution imaging in live cells.
- Andreas Bodén
- , Dirk Ollech
- & Ilaria Testa
-
Editorial |
Where imaging and metrics meet
When it comes to bioimaging and image analysis, details matter. Papers in this issue offer guidance for improved robustness and reproducibility.
-
Article
| Open Access Image restoration of degraded time-lapse microscopy data mediated by near-infrared imaging
Image restoration of degraded time-lapse microscopy data mediated by near-infrared imaging
InfraRed-mediated Image Restoration (IR2) uses deep learning to combine the benefits of deep-tissue imaging with NIR probes and the convenience of imaging with GFP for improved time-lapse imaging of embryogenesis.
- Nicola Gritti
- , Rory M. Power
- & Jan Huisken
-
Article
| Open Access The Cousa objective: a long-working distance air objective for multiphoton imaging in vivo
The Cousa objective: a long-working distance air objective for multiphoton imaging in vivo
The Cousa objective is an ultra-long working distance air objective optimized for two- and three-photon imaging. Bypassing challenges caused by water immersion and short working distances, the Cousa enables and improves imaging of diverse specimens.
- Che-Hang Yu
- , Yiyi Yu
- & Spencer LaVere Smith
-
Article
| Open Access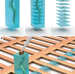 Serial Lift-Out: sampling the molecular anatomy of whole organisms
Serial Lift-Out: sampling the molecular anatomy of whole organisms
Serial Lift-Out creates a series of lamellae from one lift-out volume for cryo-ET, increasing the ease and throughput of cryo-lift-out and enabling the study of molecular anatomy in multicellular systems including C. elegans larvae.
- Oda Helene Schiøtz
- , Christoph J. O. Kaiser
- & Jürgen M. Plitzko
-
Article |
 DeepMainmast: integrated protocol of protein structure modeling for cryo-EM with deep learning and structure prediction
DeepMainmast: integrated protocol of protein structure modeling for cryo-EM with deep learning and structure prediction
DeepMainmast is a protein structure modeling protocol for cryo-EM that combines the strengths of a deep-learning-based de novo protein main-chain-tracing approach with AlphaFold2-based structure predictions for improved performance.
- Genki Terashi
- , Xiao Wang
- & Daisuke Kihara
-
Method to Watch |
 Imaging across scales
Imaging across scales
New twists on established methods and multimodal imaging are poised to bridge gaps between cellular and organismal imaging.
- Rita Strack
-
Method to Watch |
 Visual proteomics
Visual proteomics
Advances will enable proteome-scale structure determination in cells.
- Rita Strack
-
-
Research Briefing |
 Inferring how animals deform improves cell tracking
Inferring how animals deform improves cell tracking
Tracking cells is a time-consuming part of biological image analysis, and traditional manual annotation methods are prohibitively laborious for tracking neurons in the deforming and moving Caenorhabditis elegans brain. By leveraging machine learning to develop a ‘targeted augmentation’ method, we substantially reduced the number of labeled images required for tracking.
-
Article
| Open Access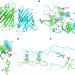 StayGold variants for molecular fusion and membrane-targeting applications
StayGold variants for molecular fusion and membrane-targeting applications
Monomeric and tandem dimer derivatives of the bright and photostable green fluorescent protein StayGold offer versatile tools for tagging target proteins and membranes in extended live-cell imaging.
- Ryoko Ando
- , Satoshi Shimozono
- & Atsushi Miyawaki
-
Article
| Open Access High-fidelity 3D live-cell nanoscopy through data-driven enhanced super-resolution radial fluctuation
High-fidelity 3D live-cell nanoscopy through data-driven enhanced super-resolution radial fluctuation
Enhanced super-resolution radial fluctuations (eSRRF) offers improved image fidelity and resolution compared to the popular SRRF method and further enables volumetric live-cell super-resolution imaging at high speeds.
- Romain F. Laine
- , Hannah S. Heil
- & Ricardo Henriques
-
-
Article
| Open Access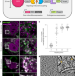 Genetically encoded multimeric tags for subcellular protein localization in cryo-EM
Genetically encoded multimeric tags for subcellular protein localization in cryo-EM
Genetically encoded multimeric particles (GEMs) are 25-nm tags with recognizable structural signatures, which can be used to label specific proteins in mammalian cells to facilitate their subcellular localization in cryo-ET.
- Herman K. H. Fung
- , Yuki Hayashi
- & Julia Mahamid
-
Article
| Open Access nextPYP: a comprehensive and scalable platform for characterizing protein variability in situ using single-particle cryo-electron tomography
nextPYP: a comprehensive and scalable platform for characterizing protein variability in situ using single-particle cryo-electron tomography
nextPYP is a turn-key framework for single-particle cryo-electron tomography that streamlines complex data analysis pipelines, from pre-processing of tilt series to high-resolution refinement, for efficient analysis and visualization of large datasets.
- Hsuan-Fu Liu
- , Ye Zhou
- & Alberto Bartesaghi
-
Perspective |
 Artificial intelligence-enabled quantitative phase imaging methods for life sciences
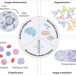
Artificial intelligence-enabled quantitative phase imaging methods for life sciences
This Perspective introduces advances in quantitative phase imaging and artificial intelligence-based image analysis and further describes how the two technologies intersect and synergize to enable biomedical research.
- Juyeon Park
- , Bijie Bai
- & YongKeun Park
-
Article
| Open Access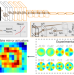 Deep learning-driven adaptive optics for single-molecule localization microscopy
Deep learning-driven adaptive optics for single-molecule localization microscopy
A deep learning approach bypasses iterative trials associated with sensorless adaptive optics to compensate for wavefront deformations when imaging biological specimens, enabling improved deep tissue localization microscopy.
- Peiyi Zhang
- , Donghan Ma
- & Fang Huang
-
Research Briefing |
 Capturing detailed cellular landscapes by montage cryo-electron tomography
Capturing detailed cellular landscapes by montage cryo-electron tomography
To capture expansive, seamless fields of view from frozen hydrated specimens by cryo-electron tomography, we developed methods for the collection and processing of montage data. This approach enables rapid acquisition of contiguous regions of specimens using a montaged tilt series collection scheme.
-
Research Briefing |
 Mapping deformations and increasing quantitative accuracy in expansion microscopy
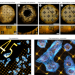
Mapping deformations and increasing quantitative accuracy in expansion microscopy
We introduce GelMap, a flexible workflow for reporting deformations and anisotropy in expansion microscopy. By intrinsically calibrating the expansion hydrogel using a fluorescent grid that scales with expansion and deforms with anisotropy, GelMap enables the reliable quantification of expansion factors and correction of deformations.
-
Article
| Open Access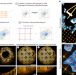 GelMap: intrinsic calibration and deformation mapping for expansion microscopy
GelMap: intrinsic calibration and deformation mapping for expansion microscopy
The GelMap workflow adds a fluorescent grid into samples before expansion, allowing for precise determination of expansion factor and subsequent deformation correction in ExM. GelMap works with diverse samples and expansion methods.
- Hugo G. J. Damstra
- , Josiah B. Passmore
- & Lukas C. Kapitein
-
Article
| Open Access Correlative montage parallel array cryo-tomography for in situ structural cell biology
Correlative montage parallel array cryo-tomography for in situ structural cell biology
Montage parallel array cryo-tomography adopts principles of montage tomography via regular array beam-image-shift montage acquisition and is robust for imaging large fields of view while retaining high-resolution structural information in cryo-electron tomography.
- Jie E. Yang
- , Matthew R. Larson
- & Elizabeth R. Wright
-
Perspective |
 Community-developed checklists for publishing images and image analyses
Community-developed checklists for publishing images and image analyses
Community-developed checklists offer best-practice guidance for biologists preparing light microscopy images and describing image analyses for publications.
- Christopher Schmied
- , Michael S. Nelson
- & Helena Klara Jambor
-
Research Highlight |
 A closer look at chromatin
A closer look at chromatin
An expansion microscopy technique called ChromExM offers detailed views into the organization chromatin and associated gene expression machinery in embryos.
- Rita Strack
-
Correspondence |
 scNodes: a correlation and processing toolkit for super-resolution fluorescence and electron microscopy
scNodes: a correlation and processing toolkit for super-resolution fluorescence and electron microscopy
- Mart G. F. Last
- , Lenard M. Voortman
- & Thomas H. Sharp
-
Brief Communication
| Open Access Segmentation metric misinterpretations in bioimage analysis
Segmentation metric misinterpretations in bioimage analysis
This study shows the importance of proper metrics for comparing algorithms for bioimage segmentation and object detection by exploring the impact of metrics on the relative performance of algorithms in three image analysis competitions.
- Dominik Hirling
- , Ervin Tasnadi
- & Peter Horvath
-
Article |
 DBlink: dynamic localization microscopy in super spatiotemporal resolution via deep learning
DBlink: dynamic localization microscopy in super spatiotemporal resolution via deep learning
DBlink uses deep learning to capture long-term dependencies between different frames in single-molecule localization microscopy data, yielding super spatiotemporal resolution videos of fast dynamic processes in living cells.
- Alon Saguy
- , Onit Alalouf
- & Yoav Shechtman
-
Editorial |
 What’s next for bioimage analysis?
What’s next for bioimage analysis?
Advanced bioimage analysis tools are poised to disrupt the way in which microscopy images are acquired and analyzed. This Focus issue shares the hopes and opinions of experts on the near and distant future of image analysis.
-
Comment |
Towards effective adoption of novel image analysis methods
The bridging of domains such as deep learning-driven image analysis and biology brings exciting promises of previously impossible discoveries as well as perils of misinterpretation and misapplication. We encourage continual communication between method developers and application scientists that emphases likely pitfalls and provides validation tools in conjunction with new techniques.
- Talley Lambert
- & Jennifer Waters
-
Comment |
 Phasor plots and the future of spectral and lifetime imaging
Phasor plots and the future of spectral and lifetime imaging
I share my opinions on the benefits of and bottlenecks for hyperspectral and time-resolved imaging. I also discuss current and future perspectives for analyzing these types of data using the phasor approach.
- Leonel Malacrida
-
Research Briefing |
 LIONESS enables 4D nanoscale reconstruction of living brain tissue
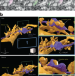
LIONESS enables 4D nanoscale reconstruction of living brain tissue
We developed LIONESS, a technology that leverages improvements to optical super-resolution microscopy and prior information on sample structure via machine learning to overcome the limitations (in 3D-resolution, signal-to-noise ratio and light exposure) of optical microscopy of living biological specimens. LIONESS enables dense reconstruction of living brain tissue and morphodynamics visualization at the nanoscale.
-
Article
| Open Access Dense 4D nanoscale reconstruction of living brain tissue
Dense 4D nanoscale reconstruction of living brain tissue
A combination of gentle stimulated emission depletion microscopy imaging and deep-learning-based improvements in signal-to-noise ratio enables high-resolution reconstruction of neuronal architecture in living tissue.
- Philipp Velicky
- , Eder Miguel
- & Johann G. Danzl
-
Research Highlight |
 Capturing hyperspectral images
Capturing hyperspectral images
A single-shot hyperspectral phasor camera (SHy-Cam) enables fast, multiplexed volumetric imaging.
- Rita Strack
-
-
Research Highlight |
 Rethinking microscope objectives
Rethinking microscope objectives
A microscope objective inspired by the Schmidt telescope offers a large field of view, high numerical aperture, long working distance and compatibility with all homogeneous immersion media for versatile bioimaging.
- Rita Strack
-
Brief Communication
| Open Access High-contrast en bloc staining of mouse whole-brain and human brain samples for EM-based connectomics
High-contrast en bloc staining of mouse whole-brain and human brain samples for EM-based connectomics
For EM-based connectomics applications, a staining protocol for large tissue samples in the range of a centimeter has been developed, which avoids artifacts common with established protocols.
- Kun Song
- , Zhihui Feng
- & Moritz Helmstaedter
-
Research Briefing |
 Photoselective sequencing harnesses microscopy to guide genomic analyses
Photoselective sequencing harnesses microscopy to guide genomic analyses
Photoselective sequencing is a new method for genomic and epigenomic profiling within specific regions of a biological specimen that are chosen using light microscopy. This combination of spatial and sequencing information preserves the connections between genomic and environmental properties and deepens our understanding of structure–function relationships in cells and tissues.
-
Comment |
 Volume EM: a quiet revolution takes shape
Volume EM: a quiet revolution takes shape
Volume electron microscopy (vEM) is a group of techniques that reveal the 3D ultrastructure of cells and tissues through continuous depths of at least 1 micrometer. A burgeoning grassroots community effort is fast building the profile and revealing the impact of vEM technology in the life sciences and clinical research.
- Lucy M. Collinson
- , Carles Bosch
- & Paul Verkade
-
Article
| Open Access Virtual-scanning light-field microscopy for robust snapshot high-resolution volumetric imaging
Virtual-scanning light-field microscopy for robust snapshot high-resolution volumetric imaging
Virtual-scanning light-field microscopy (VsLFM) uses a physics-based deep learning model to improve the quality and speed of LFM, reducing motion artifacts and enabling challenging demonstrations such as fast 3D voltage imaging in Drosophila.
- Zhi Lu
- , Yu Liu
- & Qionghai Dai
-
Article |
 High-speed low-light in vivo two-photon voltage imaging of large neuronal populations
High-speed low-light in vivo two-photon voltage imaging of large neuronal populations
A suite of tools including positive-going voltage indicators, a high-speed two-photon microscope, and denoising software enables prolonged imaging of electrical activity in neurons with limited toxicity.
- Jelena Platisa
- , Xin Ye
- & Jerry L. Chen
-
Brief Communication |
 An optical design enabling lightweight and large field-of-view head-mounted microscopes
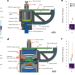
An optical design enabling lightweight and large field-of-view head-mounted microscopes
Two miniature microscopes with innovative light paths are described and applied to imaging of juvenile zebra finches and mice.
- Joseph R. Scherrer
- , Galen F. Lynch
- & Michale S. Fee
-
Review Article |
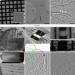 Cryo-electron tomography on focused ion beam lamellae transforms structural cell biology
Cryo-electron tomography on focused ion beam lamellae transforms structural cell biology
This Review describes advances in cryogenic electron tomography on focused ion beam lamellae, highlighting the key benefits of this technology for in situ structural biology and discussing important future directions.
- Casper Berger
- , Navya Premaraj
- & Peter J. Peters
-
Browse broader subjects
Browse narrower subjects
- Atomic force microscopy
- Ca2+ imaging
- Confocal microscopy
- Cryoelectron microscopy
- Interference microscopy
- Light-sheet microscopy
- Multiphoton microscopy
- Phase-contrast microscopy
- Polarization microscopy
- Scanning electron microscopy
- Scanning probe microscopy
- Super-resolution microscopy
- Total internal reflection microscopy
- Transmission electron microscopy
- Transmission light microscopy
- Wide-field fluorescence microscopy

 Quantification of absolute labeling efficiency at the single-protein level
Quantification of absolute labeling efficiency at the single-protein level Graphene sandwich for cryo-EM
Graphene sandwich for cryo-EM Better detection for localization microscopy
Better detection for localization microscopy Art appreciation
Art appreciation Imaging behavior with a camera array
Imaging behavior with a camera array Contrast enhancement for cryo-ET
Contrast enhancement for cryo-ET