Article
|
Open Access
Featured
-
-
Article |
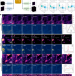 Pretraining a foundation model for generalizable fluorescence microscopy-based image restoration
Pretraining a foundation model for generalizable fluorescence microscopy-based image restoration
A pretrained foundation model (UniFMIR) enables versatile and generalizable performance across diverse fluorescence microscopy image reconstruction tasks.
- Chenxi Ma
- , Weimin Tan
- & Bo Yan
-
Article |
 Selective-plane-activation structured illumination microscopy
Selective-plane-activation structured illumination microscopy
The combination of light sheet illumination and reversibly switchable fluorophores enables improved structured illumination microscopy for fast, low-background super-resolution imaging in cells and spheroids.
- Kenta Temma
- , Ryosuke Oketani
- & Katsumasa Fujita
-
Brief Communication
| Open Access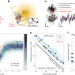 MINSTED tracking of single biomolecules
MINSTED tracking of single biomolecules
MINSTED quantifies tiny movements of individual biomolecules with high spatiotemporal precision to successfully resolve the steps of the molecular motor protein kinesin-1 labeled with a single fluorophore as it switches protofilaments.
- Lukas Scheiderer
- , Henrik von der Emde
- & Stefan W. Hell
-
Article
| Open Access An optogenetic method for the controlled release of single molecules
An optogenetic method for the controlled release of single molecules
An optogenetic system enables the controlled release of soluble and transmembrane proteins for precise exploration of cellular protein function at the single-molecule level and streamlined single-molecule imaging.
- Purba Kashyap
- , Sara Bertelli
- & Helge Ewers
-
Article
| Open Access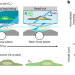 Super-sectioning with multi-sheet reversible saturable optical fluorescence transitions (RESOLFT) microscopy
Super-sectioning with multi-sheet reversible saturable optical fluorescence transitions (RESOLFT) microscopy
Multi-sheet RESOLFT combines the speed and optical sectioning of light-sheet fluorescence microscopy with reversibly photoswitchable fluorescent proteins to enable fast, volumetric super-resolution imaging in live cells.
- Andreas Bodén
- , Dirk Ollech
- & Ilaria Testa
-
-
Article
| Open Access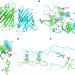 StayGold variants for molecular fusion and membrane-targeting applications
StayGold variants for molecular fusion and membrane-targeting applications
Monomeric and tandem dimer derivatives of the bright and photostable green fluorescent protein StayGold offer versatile tools for tagging target proteins and membranes in extended live-cell imaging.
- Ryoko Ando
- , Satoshi Shimozono
- & Atsushi Miyawaki
-
Article
| Open Access High-fidelity 3D live-cell nanoscopy through data-driven enhanced super-resolution radial fluctuation
High-fidelity 3D live-cell nanoscopy through data-driven enhanced super-resolution radial fluctuation
Enhanced super-resolution radial fluctuations (eSRRF) offers improved image fidelity and resolution compared to the popular SRRF method and further enables volumetric live-cell super-resolution imaging at high speeds.
- Romain F. Laine
- , Hannah S. Heil
- & Ricardo Henriques
-
Article
| Open Access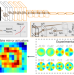 Deep learning-driven adaptive optics for single-molecule localization microscopy
Deep learning-driven adaptive optics for single-molecule localization microscopy
A deep learning approach bypasses iterative trials associated with sensorless adaptive optics to compensate for wavefront deformations when imaging biological specimens, enabling improved deep tissue localization microscopy.
- Peiyi Zhang
- , Donghan Ma
- & Fang Huang
-
Research Briefing |
 Mapping deformations and increasing quantitative accuracy in expansion microscopy
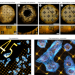
Mapping deformations and increasing quantitative accuracy in expansion microscopy
We introduce GelMap, a flexible workflow for reporting deformations and anisotropy in expansion microscopy. By intrinsically calibrating the expansion hydrogel using a fluorescent grid that scales with expansion and deforms with anisotropy, GelMap enables the reliable quantification of expansion factors and correction of deformations.
-
Article
| Open Access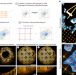 GelMap: intrinsic calibration and deformation mapping for expansion microscopy
GelMap: intrinsic calibration and deformation mapping for expansion microscopy
The GelMap workflow adds a fluorescent grid into samples before expansion, allowing for precise determination of expansion factor and subsequent deformation correction in ExM. GelMap works with diverse samples and expansion methods.
- Hugo G. J. Damstra
- , Josiah B. Passmore
- & Lukas C. Kapitein
-
Research Highlight |
 A closer look at chromatin
A closer look at chromatin
An expansion microscopy technique called ChromExM offers detailed views into the organization chromatin and associated gene expression machinery in embryos.
- Rita Strack
-
Correspondence |
 scNodes: a correlation and processing toolkit for super-resolution fluorescence and electron microscopy
scNodes: a correlation and processing toolkit for super-resolution fluorescence and electron microscopy
- Mart G. F. Last
- , Lenard M. Voortman
- & Thomas H. Sharp
-
Article |
 DBlink: dynamic localization microscopy in super spatiotemporal resolution via deep learning
DBlink: dynamic localization microscopy in super spatiotemporal resolution via deep learning
DBlink uses deep learning to capture long-term dependencies between different frames in single-molecule localization microscopy data, yielding super spatiotemporal resolution videos of fast dynamic processes in living cells.
- Alon Saguy
- , Onit Alalouf
- & Yoav Shechtman
-
Research Briefing |
 LIONESS enables 4D nanoscale reconstruction of living brain tissue
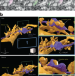
LIONESS enables 4D nanoscale reconstruction of living brain tissue
We developed LIONESS, a technology that leverages improvements to optical super-resolution microscopy and prior information on sample structure via machine learning to overcome the limitations (in 3D-resolution, signal-to-noise ratio and light exposure) of optical microscopy of living biological specimens. LIONESS enables dense reconstruction of living brain tissue and morphodynamics visualization at the nanoscale.
-
Article
| Open Access Dense 4D nanoscale reconstruction of living brain tissue
Dense 4D nanoscale reconstruction of living brain tissue
A combination of gentle stimulated emission depletion microscopy imaging and deep-learning-based improvements in signal-to-noise ratio enables high-resolution reconstruction of neuronal architecture in living tissue.
- Philipp Velicky
- , Eder Miguel
- & Johann G. Danzl
-
This Month |
Languages in the lab
Members of a lab often have a varied language background. This rich language diversity leads to lab dynamics that take mindful handling.
- Vivien Marx
-
Article |
 ERnet: a tool for the semantic segmentation and quantitative analysis of endoplasmic reticulum topology
ERnet: a tool for the semantic segmentation and quantitative analysis of endoplasmic reticulum topology
ERnet is a deep learning-based software tool for automatic segmentation and classification of structures in the endoplasmic reticulum. ERnet is compatible with many fluorescence imaging modalities and can uncover subtle phenotypic changes.
- Meng Lu
- , Charles N. Christensen
- & Clemens F. Kaminski
-
Article |
 Field-dependent deep learning enables high-throughput whole-cell 3D super-resolution imaging
Field-dependent deep learning enables high-throughput whole-cell 3D super-resolution imaging
FD-DeepLoc uses field-dependent deep learning for precise localization of spatially variant point emitters over the full chip of a modern sCMOS camera, enabling fast and high-throughput volumetric localization microscopy.
- Shuang Fu
- , Wei Shi
- & Yiming Li
-
Research Highlight |
 Next-generation expansion microscopy
Next-generation expansion microscopy
A new twist on expansion microscopy called Magnify uses a mechanically sturdy gel to simultaneously anchor and expand diverse biological samples for super-resolution imaging.
- Rita Strack
-
Article |
 A framework for evaluating the performance of SMLM cluster analysis algorithms
A framework for evaluating the performance of SMLM cluster analysis algorithms
This analysis compares the performance of seven algorithms for cluster analysis of single-molecule localization microscopy data. The results provide a framework for comparing these types of methods and point users to the best tools.
- Daniel J. Nieves
- , Jeremy A. Pike
- & Dylan M. Owen
-
Method to Watch |
 Fluorescence in structural biology
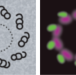
Fluorescence in structural biology
Advances in fluorescence microscopy and spectroscopy show their promise for applications that complement in situ structural biology methods like cryoelectron tomography.
- Rita Strack
-
Research Briefing |
 LocMoFit quantifies cellular structures in super-resolution data
LocMoFit quantifies cellular structures in super-resolution data
Localization Model Fit (LocMoFit) is a tool that enables fitting of super-resolution microscopy data to an arbitrary geometric model. The fit extracts quantitative parameters of individual cellular structures, which can be used to investigate dynamic and heterogenous protein assemblies and to create average protein distribution maps.
-
Matters Arising |
 Reply to: Assessment of 3D MINFLUX data for quantitative structural biology in cells
Reply to: Assessment of 3D MINFLUX data for quantitative structural biology in cells
- Klaus C. Gwosch
- , Francisco Balzarotti
- & Stefan W. Hell
-
Article
| Open Access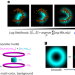 Maximum-likelihood model fitting for quantitative analysis of SMLM data
Maximum-likelihood model fitting for quantitative analysis of SMLM data
Localization Model Fit (LocMoFit) is an open-source tool for extracting meaningful parameters from individual structures in localization microscopy data. The framework was used for quantitative analysis of diverse biological structures.
- Yu-Le Wu
- , Philipp Hoess
- & Jonas Ries
-
Research Briefing |
 Robust fluorescent proteins for high-resolution microscopy and biochemical techniques
Robust fluorescent proteins for high-resolution microscopy and biochemical techniques
Hyperfolder yellow fluorescent protein (hfYFP) and its variants are fluorescent proteins with high chemical and thermal stability. They resist aggregation, withstand diverse chemical challenges and show promise in expansion and electron microscopies. The chloride resistance and uncanny stability in guanidinium of hfYFP enable fluorescence-guided protein purification under denaturing conditions.
-
Article
| Open Access Chemically stable fluorescent proteins for advanced microscopy
Chemically stable fluorescent proteins for advanced microscopy
The engineered hyperfolder YFP (hfYFP) and variants offer unprecedented chemical and thermal stability, making them versatile probes for microscopy as well as for challenging applications like correlative light and electron microscopy and expansion microscopy.
- Benjamin C. Campbell
- , Maria G. Paez-Segala
- & Ce Feng Liu
-
Research Briefing |
 Doubling the resolution of light-sheet fluorescence microscopy
Doubling the resolution of light-sheet fluorescence microscopy
A combination of light-sheet fluorescence microscopy (LSFM) with structured illumination doubles resolving power over LSFM alone. We show a practical implementation using a single objective for illumination and fluorescence detection and demonstrate its use for rapid volumetric imaging.
-
Article |
 Resolution doubling in light-sheet microscopy via oblique plane structured illumination
Resolution doubling in light-sheet microscopy via oblique plane structured illumination
The longstanding goal of combining the optical sectioning of light-sheet illumination and the resolving power of multidirectional structured illumination microscopy is realized using an oblique plane microscope for improved fast 3D subcellular imaging.
- Bingying Chen
- , Bo-Jui Chang
- & Reto P. Fiolka
-
Article
| Open Access Event-triggered STED imaging
Event-triggered STED imaging
Event-triggered STED is an automated approach that can initiate 2D or 3D STED imaging of specific regions in biological samples after detection of an event of interest. This approach can help maximize observations in live cell imaging and enable discovery.
- Jonatan Alvelid
- , Martina Damenti
- & Ilaria Testa
-
Brief Communication
| Open Access DNA-PAINT MINFLUX nanoscopy
DNA-PAINT MINFLUX nanoscopy
A systematic exploration of MINFLUX nanoscopy with DNA-PAINT labeling leads to improved nanoscopy in fixed cells and MINFLUX imaging with increased multiplexing, as exemplified by three-color imaging of mitochondria in mammalian cells.
- Lynn M. Ostersehlt
- , Daniel C. Jans
- & Stefan Jakobs
-
News & Views |
 Fluorophores’ talk turns them dark
Fluorophores’ talk turns them dark
Dipole–dipole crosstalk between fluorophores separated by a distance of less than 10 nm induces changes in their photophysics, which adds a challenge to localization microscopy in the sub-10-nm regime.
- Karim Almahayni
- , Malte Spiekermann
- & Leonhard Möckl
-
Article
| Open Access Optimal precision and accuracy in 4Pi-STORM using dynamic spline PSF models
Optimal precision and accuracy in 4Pi-STORM using dynamic spline PSF models
A dynamic model of the 4Pi point spread function enables localization microscopy with exceptional three-dimensional resolution and a simpler optical design. 4Pi-STORM images of neurons and mitochondria reveal new details of nanoscale protein and nucleic acid organization.
- Mark Bates
- , Jan Keller-Findeisen
- & Stefan W. Hell
-
Article |
 Correction of multiple-blinking artifacts in photoactivated localization microscopy
Correction of multiple-blinking artifacts in photoactivated localization microscopy
A model-based correction (MBC) algorithm offers fast and accurate correction of multiple-blinking artifacts in PALM data. MBC outperforms other algorithms in both speed and accuracy and improves quantitative downstream image analysis.
- Louis G. Jensen
- , Tjun Yee Hoh
- & Dylan M. Owen
-
Article |
 Isotropic super-resolution light-sheet microscopy of dynamic intracellular structures at subsecond timescales
Isotropic super-resolution light-sheet microscopy of dynamic intracellular structures at subsecond timescales
Combining a double-ring modulated SPIM with reduced side lobes and a sectionalized deep-learning based super-resolution algorithm enables fast, high-resolution, volumetric imaging of organelle interactions and dynamics in live cells.
- Yuxuan Zhao
- , Meng Zhang
- & Yu-Hui Zhang
-
Research Highlight |
 Three views are better than one
Three views are better than one
Researchers push the limits of confocal microscopy by combining multiple views, super-resolution and deep learning.
- Rita Strack
-
News & Views |
 Expansion microscopy opens the door to exploring more challenges
Expansion microscopy opens the door to exploring more challenges
Cryofixation-based ultrastructure-expansion microscopy (cryo-ExM) bypasses artifacts caused by chemical fixation and establishes more-native preservation of biological samples.
- Mengfei Gao
-
Article
| Open Access Visualizing the native cellular organization by coupling cryofixation with expansion microscopy (Cryo-ExM)
Visualizing the native cellular organization by coupling cryofixation with expansion microscopy (Cryo-ExM)
Cryo-ExM combines the expansion microscopy for super-resolution imaging with cryofixation for ultrastructure preservation. Cryo-ExM outperforms established fixation methods on a range of sensitive subcellular structures.
- Marine H. Laporte
- , Nikolai Klena
- & Paul Guichard
-
Article |
 Deep learning enables fast and dense single-molecule localization with high accuracy
Deep learning enables fast and dense single-molecule localization with high accuracy
DECODE uses deep learning for localizing single emitters in high-density two-dimensional and three-dimensional single-molecule localization microscopy data. DECODE outperforms available methods and enables fast live-cell SMLM of dynamic processes.
- Artur Speiser
- , Lucas-Raphael Müller
- & Srinivas C. Turaga
-
Research Highlight |
Atomic force microscopy in super-resolution
Applying principles from super-resolution fluorescence microscopy increases the resolution of atomic force microscopy.
- Nina Vogt
-
Article |
 Structured illumination microscopy with noise-controlled image reconstructions
Structured illumination microscopy with noise-controlled image reconstructions
Super-resolution structured illumination microscopy reconstruction algorithms are described that can handle structured noise artifacts in two and three dimensions. The algorithms lack adjustable parameters and enhance objective representation of imaged objects.
- Carlas S. Smith
- , Johan A. Slotman
- & Sjoerd Stallinga
-
This Month |
 Jie Xiao
Jie Xiao
Reaching ground truth in super-res microscopy, powered by a love of cooking and food.
- Vivien Marx
-
Article |
 Three-dimensional adaptive optical nanoscopy for thick specimen imaging at sub-50-nm resolution
Three-dimensional adaptive optical nanoscopy for thick specimen imaging at sub-50-nm resolution
The combination of adaptive optics with an improved isoSTED nanoscope allows imaging of cells and tissues with sub-50-nm isotropic resolution.
- Xiang Hao
- , Edward S. Allgeyer
- & Joerg Bewersdorf
-
Article |
 A pairwise distance distribution correction (DDC) algorithm to eliminate blinking-caused artifacts in SMLM
A pairwise distance distribution correction (DDC) algorithm to eliminate blinking-caused artifacts in SMLM
Distance distribution correction (DDC) eliminates repeat localizations caused by fluorophore blinking without the need for calibrations. Use of DDC yields accurate and quantifiable single-molecule localization microscopy data.
- Christopher H. Bohrer
- , Xinxing Yang
- & Jie Xiao
-
Correspondence |
 PYMEVisualize: an open-source tool for exploring 3D super-resolution data
PYMEVisualize: an open-source tool for exploring 3D super-resolution data
- Zach Marin
- , Michael Graff
- & David Baddeley
-
Research Highlight |
 Expanding views of the transcriptome
Expanding views of the transcriptome
In situ long-read sequencing combined with expansion microscopy enables precise views of transcriptomes in intact biological systems.
- Rita Strack
-
Brief Communication |
 Molecular-scale axial localization by repetitive optical selective exposure
Molecular-scale axial localization by repetitive optical selective exposure
ROSE-Z achieves axial interference through an asymmetrical optical scheme, yielding 2 nm axial localization precision with ~3,000 photons and a single objective, which offers improved multicolor three-dimensional localization microscopy for cellular structures.
- Lusheng Gu
- , Yuanyuan Li
- & Wei Ji
-
Research Highlight |
Super-resolution 3D live cell imaging
3D pRESOLFT enables volumetric live cell imaging with high spatial and temporal resolution.
- Nina Vogt
-
Research Highlight |
DNA-PAINT takes a left turn
DNA-PAINT with left-handed DNA oligomers enables imaging of nuclear structures with reduced background.
- Rita Strack

 Quantification of absolute labeling efficiency at the single-protein level
Quantification of absolute labeling efficiency at the single-protein level Better detection for localization microscopy
Better detection for localization microscopy