Featured
-
-
Article
| Open Access Compact engineered human mechanosensitive transactivation modules enable potent and versatile synthetic transcriptional control
Compact engineered human mechanosensitive transactivation modules enable potent and versatile synthetic transcriptional control
Synthetic CRISPR-based transactivation domains engineered using mechanosensitive transcription factors enable robust transcriptional control.
- Barun Mahata
- , Alan Cabrera
- & Isaac B. Hilton
-
Article
| Open Access OPUS-DSD: deep structural disentanglement for cryo-EM single-particle analysis
OPUS-DSD: deep structural disentanglement for cryo-EM single-particle analysis
OPUS-DSD is a neural network-based algorithm that reconstructs distinct conformations or continuous dynamics of the macromolecular structural landscape, starting from single-particle cryo-EM data.
- Zhenwei Luo
- , Fengyun Ni
- & Jianpeng Ma
-
-
Editorial |
Registered Reports: what we’ve learned so far
Nature Methods is proud to publish our very first Registered Report in this issue. Here, we reflect on what we have learned since introducing this article type.
-
Brief Communication |
 Molecular mechanocytometry using tension-activated cell tagging
Molecular mechanocytometry using tension-activated cell tagging
Tension-activated cell tagging (TaCT) is a new method that uses flow cytometry to sort mechanically active cells based on the forces generated by their surface adhesion receptors.
- Rong Ma
- , Sk Aysha Rashid
- & Khalid Salaita
-
News & Views |
 Real-time imaging of dynamic tissues
Real-time imaging of dynamic tissues
A combined modality enables real-time imaging of mouse lungs and spans whole-organ to cellular scales.
- Joan E. Nichols
- & Sasha R. Azar
-
Comment |
 A practitioner’s view of spectral flow cytometry
A practitioner’s view of spectral flow cytometry
Spectral flow cytometry enables the simultaneous analysis of a large number of cell surface markers at the single-cell level
- Siddhartha Sharma
- , Josh Boyer
- & Luc Teyton
-
Research Briefing |
 Evaluating long-read RNA-sequencing analysis tools with in silico mixtures
Evaluating long-read RNA-sequencing analysis tools with in silico mixtures
We conducted a comprehensive long-read RNA sequencing (RNA-seq) benchmarking experiment by combining spike-ins and in silico mixtures to establish a ground-truth dataset. We used long- and short-read RNA-seq technology to deeply sequence samples and compared the performance of a range of analysis tools on these data.
-
Article |
 CryoREAD: de novo structure modeling for nucleic acids in cryo-EM maps using deep learning
CryoREAD: de novo structure modeling for nucleic acids in cryo-EM maps using deep learning
Few methods for three-dimensional structure modeling of nucleic acids from cryo-EM data exist. CryoREAD, a fully automated DNA/RNA atomic structure modeling method based on deep learning, was developed to fill this gap.
- Xiao Wang
- , Genki Terashi
- & Daisuke Kihara
-
Article |
 Streamlined and sensitive mono- and di-ribosome profiling in yeast and human cells
Streamlined and sensitive mono- and di-ribosome profiling in yeast and human cells
A comprehensive redevelopment of the ribosome profiling workflow involves improved nuclease treatment and sequencing library preparation, enabling richer and more accurate translatome profiling with lower input and fewer technical hurdles.
- Lucas Ferguson
- , Heather E. Upton
- & Nicholas T. Ingolia
-
Article
| Open Access Spatial single-cell mass spectrometry defines zonation of the hepatocyte proteome
Spatial single-cell mass spectrometry defines zonation of the hepatocyte proteome
Single-cell Deep Visual Proteomics integrates imaging, cell segmentation, laser microdissection and multiplexed mass spectrometry for spatial single-cell proteomics measurements in complex tissues.
- Florian A. Rosenberger
- , Marvin Thielert
- & Matthias Mann
-
Resource
| Open Access Waxholm Space atlas of the rat brain: a 3D atlas supporting data analysis and integration
Waxholm Space atlas of the rat brain: a 3D atlas supporting data analysis and integration
An updated version of the Waxholm Space atlas of the rat brain includes more detailed annotations of several brain regions, including the cortex, striatopallidal region, midbrain and thalamus, expanding the previous version with 112 new and 57 revised structures.
- Heidi Kleven
- , Ingvild E. Bjerke
- & Trygve B. Leergaard
-
News & Views |
 Factoring single-cell perturbations
Factoring single-cell perturbations
GSFA is a statistical model to automatically detect latent factors (or gene modules) in single-cell CRISPR screening.
- Bicna Song
- & Wei Li
-
Article
| Open Access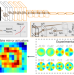 Deep learning-driven adaptive optics for single-molecule localization microscopy
Deep learning-driven adaptive optics for single-molecule localization microscopy
A deep learning approach bypasses iterative trials associated with sensorless adaptive optics to compensate for wavefront deformations when imaging biological specimens, enabling improved deep tissue localization microscopy.
- Peiyi Zhang
- , Donghan Ma
- & Fang Huang
-
Article
| Open Access D-LMBmap: a fully automated deep-learning pipeline for whole-brain profiling of neural circuitry
D-LMBmap: a fully automated deep-learning pipeline for whole-brain profiling of neural circuitry
D-LMBmap is a fully automated pipeline for mesoscale connectomics including deep-learning modules for axon segmentation, brain region segmentation and whole-brain registration. D-LMBmap works accurately across cell types and modalities.
- Zhongyu Li
- , Zengyi Shang
- & Jing Ren
-
Brief Communication |
 Rapid droplet-based mixing for single-molecule spectroscopy
Rapid droplet-based mixing for single-molecule spectroscopy
Droplet-based microfluidics enable rapid mixing with millisecond dead times and allow single-molecule measurements of non-equilibrium binding kinetics on even challenging, strongly adsorptive samples, such as intrinsically disordered proteins.
- Tianjin Yang
- , Karin J. Buholzer
- & Benjamin Schuler
-
Brief Communication |
 Flash entropy search to query all mass spectral libraries in real time
Flash entropy search to query all mass spectral libraries in real time
Flash entropy speeds up mass spectral similarity searches 10,000-fold, including for open search, hybrid search and neutral loss search.
- Yuanyue Li
- & Oliver Fiehn
-
Brief Communication
| Open Access Fast and robust metagenomic sequence comparison through sparse chaining with skani
Fast and robust metagenomic sequence comparison through sparse chaining with skani
skani achieves fast calculation of average nucleotide identity (ANI) between metagenome-assembled genomes (MAGs), with improved robustness against incomplete and fragmented MAGs.
- Jim Shaw
- & Yun William Yu
-
Article
| Open Access Deep generative modeling of transcriptional dynamics for RNA velocity analysis in single cells
Deep generative modeling of transcriptional dynamics for RNA velocity analysis in single cells
veloVI enhances RNA velocity analysis with uncertainty quantification and extensibility by deep generative modeling of gene-specific transcriptional dynamics.
- Adam Gayoso
- , Philipp Weiler
- & Nir Yosef
-
Article
| Open Access Spatiotemporal, optogenetic control of gene expression in organoids
Spatiotemporal, optogenetic control of gene expression in organoids
A workflow combining optogenetic perturbations with spatial transcriptomics to program spatiotemporal gene expression patterns in organoids.
- Ivano Legnini
- , Lisa Emmenegger
- & Nikolaus Rajewsky
-
Research Briefing |
 Capturing detailed cellular landscapes by montage cryo-electron tomography
Capturing detailed cellular landscapes by montage cryo-electron tomography
To capture expansive, seamless fields of view from frozen hydrated specimens by cryo-electron tomography, we developed methods for the collection and processing of montage data. This approach enables rapid acquisition of contiguous regions of specimens using a montaged tilt series collection scheme.
-
Technology Feature |
 Greening the lab
Greening the lab
Making biological research more sustainable requires an accurate assessment of its environmental impact, both at the bench and on the computer.
- Caroline Seydel
-
Research Briefing |
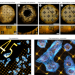 Mapping deformations and increasing quantitative accuracy in expansion microscopy
Mapping deformations and increasing quantitative accuracy in expansion microscopy
We introduce GelMap, a flexible workflow for reporting deformations and anisotropy in expansion microscopy. By intrinsically calibrating the expansion hydrogel using a fluorescent grid that scales with expansion and deforms with anisotropy, GelMap enables the reliable quantification of expansion factors and correction of deformations.
-
Article |
 Large Stokes shift fluorescent RNAs for dual-emission fluorescence and bioluminescence imaging in live cells
Large Stokes shift fluorescent RNAs for dual-emission fluorescence and bioluminescence imaging in live cells
The Clivias are a series of small, monomeric fluorescent RNAs that emit with a large Stokes shift in the orange–red. They enable multiplexed RNA imaging in live cells and BRET-based detection of protein–RNA interactions in mice.
- Li Jiang
- , Xin Xie
- & Yi Yang
-
Article
| Open Access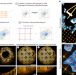 GelMap: intrinsic calibration and deformation mapping for expansion microscopy
GelMap: intrinsic calibration and deformation mapping for expansion microscopy
The GelMap workflow adds a fluorescent grid into samples before expansion, allowing for precise determination of expansion factor and subsequent deformation correction in ExM. GelMap works with diverse samples and expansion methods.
- Hugo G. J. Damstra
- , Josiah B. Passmore
- & Lukas C. Kapitein
-
Article
| Open Access Statistically unbiased prediction enables accurate denoising of voltage imaging data
Statistically unbiased prediction enables accurate denoising of voltage imaging data
Statistically unbiased prediction utilizing spatiotemporal information in imaging data (SUPPORT) is a self-supervised deep learning approach to accurately denoise voltage and calcium imaging data while preserving true dynamic signals.
- Minho Eom
- , Seungjae Han
- & Young-Gyu Yoon
-
Article
| Open Access Correlative montage parallel array cryo-tomography for in situ structural cell biology
Correlative montage parallel array cryo-tomography for in situ structural cell biology
Montage parallel array cryo-tomography adopts principles of montage tomography via regular array beam-image-shift montage acquisition and is robust for imaging large fields of view while retaining high-resolution structural information in cryo-electron tomography.
- Jie E. Yang
- , Matthew R. Larson
- & Elizabeth R. Wright
-
Perspective |
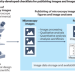 Community-developed checklists for publishing images and image analyses
Community-developed checklists for publishing images and image analyses
Community-developed checklists offer best-practice guidance for biologists preparing light microscopy images and describing image analyses for publications.
- Christopher Schmied
- , Michael S. Nelson
- & Helena Klara Jambor
-
Review Article |
 Principles and challenges of modeling temporal and spatial omics data
Principles and challenges of modeling temporal and spatial omics data
This Review discusses statistical and computational strategies for analyzing various spatial and temporal omics data types, with an emphasis on the common modeling principles.
- Britta Velten
- & Oliver Stegle
-
Article |
 Crystal ribcage: a platform for probing real-time lung function at cellular resolution
Crystal ribcage: a platform for probing real-time lung function at cellular resolution
A fabricated transparent ribcage facilitates real-time analysis of whole-lung function down to cellular resolution.
- Rohin Banerji
- , Gabrielle N. Grifno
- & Hadi T. Nia
-
Correspondence |
 Extending support for mouse data in the Molecular Signatures Database (MSigDB)
Extending support for mouse data in the Molecular Signatures Database (MSigDB)
- Anthony S. Castanza
- , Jill M. Recla
- & Jill P. Mesirov
-
Article |
 Recovery of missing single-cell RNA-sequencing data with optimized transcriptomic references
Recovery of missing single-cell RNA-sequencing data with optimized transcriptomic references
This paper presents an improved approach for mapping single-cell RNA-seq reads with optimized transcriptomic references, which markedly recovers previously missing gene expression data.
- Allan-Hermann Pool
- , Helen Poldsam
- & Yuki Oka
-
-
Editorial |
Setting standards for stem cells
New traceability and reporting standards aim to improve transparency in stem cell research.
-
Research Highlight |
 A closer look at chromatin
A closer look at chromatin
An expansion microscopy technique called ChromExM offers detailed views into the organization chromatin and associated gene expression machinery in embryos.
- Rita Strack
-
Article |
 FIOLA: an accelerated pipeline for fluorescence imaging online analysis
FIOLA: an accelerated pipeline for fluorescence imaging online analysis
FIOLA is a pipeline for processing calcium or voltage imaging data. Its advantages include the fast speed and online processing.
- Changjia Cai
- , Cynthia Dong
- & Andrea Giovannucci
-
Registered Report |
 Quantitative assessment of near-infrared fluorescent proteins
Quantitative assessment of near-infrared fluorescent proteins
This Registered Report describes an extensive comparison of 22 near-infrared fluorescent proteins in vitro, in cultured mammalian cells, and in model animals, clarifying top performers in diverse biological settings.
- Hanbin Zhang
- , Stavrini Papadaki
- & Kiryl D. Piatkevich
-
News & Views |
 Advancing multiplexed imaging for enhanced tissue complexity analysis
Advancing multiplexed imaging for enhanced tissue complexity analysis
Multiplexed spatial immunophenotyping has advanced our understanding of tissues in the context of homeostasis and disease. Two studies now provide additional tools to overcome challenges with multiplexed imaging: one procedure amplifies the detection of low-abundance antigens by integrating SABER and IMC technologies, and the other is an X-ray-based method that enables the non-destructive multiplexed detection of antigens in tissues at scalable resolution and speed.
- Marieke E. Ijsselsteijn
- & Noel F. C. C. de Miranda
-
Brief Communication
| Open Access DNA-barcoded signal amplification for imaging mass cytometry enables sensitive and highly multiplexed tissue imaging
DNA-barcoded signal amplification for imaging mass cytometry enables sensitive and highly multiplexed tissue imaging
SABER-IMC combines DNA-based signal amplification by exchange reaction (SABER) with imaging mass cytometry (IMC) to enable simultaneous and highly multiplexed marker detection, even of low-abundance markers not detectable with IMC alone.
- Tsuyoshi Hosogane
- , Ruben Casanova
- & Bernd Bodenmiller
-
Article
| Open Access Multielement Z-tag imaging by X-ray fluorescence microscopy for next-generation multiplex imaging
Multielement Z-tag imaging by X-ray fluorescence microscopy for next-generation multiplex imaging
Multielement Z-tag X-ray fluorescence (MEZ-XRF) offers a new avenue for nondestructive and highly multiplexed tissue imaging and operates from the nanometer to whole-tissue scale, unlocking new biological observations.
- Merrick Strotton
- , Tsuyoshi Hosogane
- & Bernd Bodenmiller
-
Research Briefing |
 scNanoHi-C uncovers single-cell high-order chromatin structures and gene regulation
scNanoHi-C uncovers single-cell high-order chromatin structures and gene regulation
Leveraging nanopore long-read sequencing, scNanoHi-C identifies multiway interactions between enhancers and their target promoters within a single cell. Compared with short-read-based single-cell Hi-C or population-based multiway sequencing methods, scNanoHi-C offers new opportunities to investigate the heterogeneities of single-cell gene regulation networks mediated by high-order 3D chromatin structures.
-
Article |
 A fluorogenic chemically induced dimerization technology for controlling, imaging and sensing protein proximity
A fluorogenic chemically induced dimerization technology for controlling, imaging and sensing protein proximity
CATCHFIRE (chemically assisted tethering of chimera by fluorogenic-induced recognition) tools are small tags that can chemically dimerize with turn-on fluorescence, enabling simultaneous control and visualization of protein proximity.
- Sara Bottone
- , Octave Joliot
- & Arnaud Gautier
-
Article |
 scNanoHi-C: a single-cell long-read concatemer sequencing method to reveal high-order chromatin structures within individual cells
scNanoHi-C: a single-cell long-read concatemer sequencing method to reveal high-order chromatin structures within individual cells
scNanoHi-C combines Nanopore long-read sequencing with a proximity-ligation-based Hi-C protocol to profile high-order genome structures in individual cells, enabling the capture of multiway interactions among enhancers and promoters.
- Wen Li
- , Jiansen Lu
- & Fuchou Tang
-
Technology Feature |
 A tough ask: high-efficiency, large-cargo prime editing
A tough ask: high-efficiency, large-cargo prime editing
Researchers like the gene-editing method called prime editing for its precision and versatility. As they ask the method to do more, many factors shape what happens next.
- Vivien Marx
-
Correspondence |
 scNodes: a correlation and processing toolkit for super-resolution fluorescence and electron microscopy
scNodes: a correlation and processing toolkit for super-resolution fluorescence and electron microscopy
- Mart G. F. Last
- , Lenard M. Voortman
- & Thomas H. Sharp
-
Article |
 Time-resolved cryo-EM using a combination of droplet microfluidics with on-demand jetting
Time-resolved cryo-EM using a combination of droplet microfluidics with on-demand jetting
This article describes a time-resolved cryo-plunger that combines a droplet-based microfluidic mixer with a laser-induced generator of microjets that allows rapid reaction initiation and plunge-freezing of cryo-EM grids. A time resolution of 5 ms was achieved using this approach.
- Stefania Torino
- , Mugdha Dhurandhar
- & Rouslan G. Efremov
-
Article
| Open Access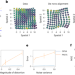 Alignment of spatial genomics data using deep Gaussian processes
Alignment of spatial genomics data using deep Gaussian processes
Gaussian Process Spatial Alignment (GPSA) aligns multiple spatially resolved genomics and histology datasets and improves downstream analysis.
- Andrew Jones
- , F. William Townes
- & Barbara E. Engelhardt
-
Article |
 UniAligner: a parameter-free framework for fast sequence alignment
UniAligner: a parameter-free framework for fast sequence alignment
Compared to other sequences, extra-long tandem repeats, such as centromeres and immunoglobulin loci, are more difficult to align. This study presents UniAligner, a computational method for efficiently and accurately aligning extra-long tandem repeats, facilitating analysis of their variation and evolution.
- Andrey V. Bzikadze
- & Pavel A. Pevzner
-
Browse broader subjects
Browse narrower subjects
- Analytical biochemistry
- Behavioural methods
- Bioinformatics
- Biological models
- Biophysical methods
- Cytological techniques
- Electrophysiology
- Epigenetics analysis
- Experimental organisms
- Gene delivery
- Gene expression analysis
- Genetic engineering
- Genetic techniques
- Genomic analysis
- High-throughput screening
- Imaging
- Immunological techniques
- Isolation, separation and purification
- Lab-on-a-chip
- Mass spectrometry
- Metabolomics
- Microbiology techniques
- Microscopy
- Molecular engineering
- Nanobiotechnology
- Optogenetics
- Proteomic analysis
- Sensors and probes
- Sequencing
- Software
- Optical spectroscopy
- Structure determination

 Population-level integration of single-cell datasets enables multi-scale analysis across samples
Population-level integration of single-cell datasets enables multi-scale analysis across samples Resolving epigenetic base ambiguity
Resolving epigenetic base ambiguity Prefabricated inks promote heart bioprinting
Prefabricated inks promote heart bioprinting