Featured
-
-
Article
| Open Access Multiscale biochemical mapping of the brain through deep-learning-enhanced high-throughput mass spectrometry
Multiscale biochemical mapping of the brain through deep-learning-enhanced high-throughput mass spectrometry
MEISTER is an integrative experimental and computational framework for mass spectrometry that integrates three-dimensional, organ-wide biomolecular mapping with single-cell analysis for multiscale profiling of spatial–biochemical organization.
- Yuxuan Richard Xie
- , Daniel C. Castro
- & Fan Lam
-
Correspondence |
 RepoRT: a comprehensive repository for small molecule retention times
RepoRT: a comprehensive repository for small molecule retention times
- Fleming Kretschmer
- , Eva-Maria Harrieder
- & Michael Witting
-
Brief Communication |
 Cardinal v.3: a versatile open-source software for mass spectrometry imaging analysis
Cardinal v.3: a versatile open-source software for mass spectrometry imaging analysis
Cardinal v.3 is an open-source software for reproducible analysis of mass spectrometry imaging experiments, and includes data processing features such as mass recalibration, statistical analyses such as single-ion segmentation and rough annotation-based classification, and analyses of large-scale multitissue experiments.
- Kylie Ariel Bemis
- , Melanie Christine Föll
- & Olga Vitek
-
Correspondence |
 METLIN-CCS: an ion mobility spectrometry collision cross section database
METLIN-CCS: an ion mobility spectrometry collision cross section database
- Erin S. Baker
- , Corey Hoang
- & Gary Siuzdak
-
Brief Communication |
 Flash entropy search to query all mass spectral libraries in real time
Flash entropy search to query all mass spectral libraries in real time
Flash entropy speeds up mass spectral similarity searches 10,000-fold, including for open search, hybrid search and neutral loss search.
- Yuanyue Li
- & Oliver Fiehn
-
Research Briefing |
 Accurate determination of molecular formulae using tandem mass spectrometry
Accurate determination of molecular formulae using tandem mass spectrometry
We highlight the BUDDY software, which was developed to accurately determine the molecular formulae of unknown chemicals in mass spectrometry data. BUDDY is a bottom-up approach that shows superior annotation performance on reference spectra and experimental datasets. Incorporation of global peak annotation could enable BUDDY to refine formula annotations and reveal feature interrelationships.
-
Article |
 BUDDY: molecular formula discovery via bottom-up MS/MS interrogation
BUDDY: molecular formula discovery via bottom-up MS/MS interrogation
BUDDY is a bottom-up tandem MS (MS/MS) interrogation method for de novo molecular formula annotation with significance estimation.
- Shipei Xing
- , Sam Shen
- & Tao Huan
-
Comment |
 Mass spectrometry imaging: the rise of spatially resolved single-cell omics
Mass spectrometry imaging: the rise of spatially resolved single-cell omics
Increasing evidence suggests that the spatial distribution of biomolecules within cells is a critical component in deciphering single-cell molecular heterogeneity. State-of-the-art single-cell MS imaging is uniquely capable of localizing biomolecules within cells, providing a dimension of information beyond what is currently available through in-depth omics investigations.
- Hua Zhang
- , Daniel G. Delafield
- & Lingjun Li
-
Article
| Open Access HyU: Hybrid Unmixing for longitudinal in vivo imaging of low signal-to-noise fluorescence
HyU: Hybrid Unmixing for longitudinal in vivo imaging of low signal-to-noise fluorescence
Hybrid Unmixing offers enhanced imaging of multiplexed fluorescence labels, enabling longitudinal imaging of multiple fluorescent signals with reduced illumination intensities.
- Hsiao Ju Chiang
- , Daniel E. S. Koo
- & Francesco Cutrale
-
Method to Watch |
 Annotating unknown metabolites
Annotating unknown metabolites
Computational approaches are pushing the limits of unknown metabolite annotation.
- Arunima Singh
-
This Month |
How to stay social
Many scientists are active on social media, especially Twitter. The social media world is changing, but these researchers want to stay socially connected.
- Vivien Marx
-
Review Article |
 Advances in measuring cancer cell metabolism with subcellular resolution
Advances in measuring cancer cell metabolism with subcellular resolution
This Review covers advances in methods for studying metabolism at the subcellular level and how they have influenced our understanding of cancer biology.
- Victor Ruiz-Rodado
- , Adrian Lita
- & Mioara Larion
-
News & Views |
 Some assembly required
Some assembly required
De novo structure generation from tandem mass spectrometry data will advance annotation of unknown metabolomic signals.
- Corey D. Broeckling
-
Article
| Open Access MSNovelist: de novo structure generation from mass spectra
MSNovelist: de novo structure generation from mass spectra
MSNovelist combines fingerprint prediction with an encoder–decoder neural network for de novo structure generation of small molecules from mass spectra.
- Michael A. Stravs
- , Kai Dührkop
- & Nicola Zamboni
-
Technology Feature |
 The chemistry of microbiome–host togetherness
The chemistry of microbiome–host togetherness
Labs are stacking techniques to disentangle the chemical influences of microbiomes.
- Vivien Marx
-
Article |
 Spatially resolved isotope tracing reveals tissue metabolic activity
Spatially resolved isotope tracing reveals tissue metabolic activity
Iso-imaging integrates stable-isotope infusions with imaging mass spectrometry to enable quantitative analysis of metabolic activity in mammalian tissues with spatial resolution. IsoScope software facilitates analysis of iso-imaging data.
- Lin Wang
- , Xi Xing
- & Shawn M. Davidson
-
Method to Watch |
 Deconstructing subcellular composition
Deconstructing subcellular composition
High-throughput and ultrasensitive mass spectrometry–based approaches are gaining ground in subcellular compositional analysis.
- Arunima Singh
-
Technology Feature |
 Single-cell metabolomics hits its stride
Single-cell metabolomics hits its stride
An array of new techniques allow researchers to catalog the chemical contents of a single cell, or even a single organelle.
- Caroline Seydel
-
Correspondence |
 GNPS Dashboard: collaborative exploration of mass spectrometry data in the web browser
GNPS Dashboard: collaborative exploration of mass spectrometry data in the web browser
- Daniel Petras
- , Vanessa V. Phelan
- & Mingxun Wang
-
Article |
 Spectral entropy outperforms MS/MS dot product similarity for small-molecule compound identification
Spectral entropy outperforms MS/MS dot product similarity for small-molecule compound identification
This manuscript proposes the use of spectral entropy similarity as a measure of similarity for small-molecule mass spectra and the use of a one-bond-difference approach for improving false discovery rates.
- Yuanyue Li
- , Tobias Kind
- & Oliver Fiehn
-
Article |
 HERMES: a molecular-formula-oriented method to target the metabolome
HERMES: a molecular-formula-oriented method to target the metabolome
HERMES is a molecular-formula-oriented and peak-detection-free method that uses LC/MS1 information to optimize MS2 acquisition for LC/MS-based metabolomic analysis.
- Roger Giné
- , Jordi Capellades
- & Oscar Yanes
-
Article |
 Metabolite discovery through global annotation of untargeted metabolomics data
Metabolite discovery through global annotation of untargeted metabolomics data
The NetID algorithm annotates untargeted LC-MS metabolomics data by combining known biochemical and metabolomic principles with a global network optimization strategy.
- Li Chen
- , Wenyun Lu
- & Joshua D. Rabinowitz
-
News & Views |
 Mass spectrometry comes of age for subcellular organelles
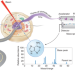
Mass spectrometry comes of age for subcellular organelles
Technological innovations in optical object recognition and high-throughput ultrasensitive mass spectrometry are enabling subcellular metabolomics and peptidomics, providing unprecedented opportunities to study small-molecule mediators of cellular function with important implications in health and disease.
- Peter Nemes
-
Article |
 SEAM is a spatial single nuclear metabolomics method for dissecting tissue microenvironment
SEAM is a spatial single nuclear metabolomics method for dissecting tissue microenvironment
SEAM is a platform for the analysis of high-resolution secondary ion mass spectrometry imaging that allows spatially resolved nuclear metabolomic profiling at the single-cell level.
- Zhiyuan Yuan
- , Qiming Zhou
- & Michael Q. Zhang
-
This Month |
 Jonathan Sweedler
Jonathan Sweedler
Scuba diving and jazz bring together chemistry, neuroscience and life inland.
- Vivien Marx
-
Article
| Open Access Image-guided MALDI mass spectrometry for high-throughput single-organelle characterization
Image-guided MALDI mass spectrometry for high-throughput single-organelle characterization
This article presents a workflow for image-guided MALDI MS characterization of peptide and lipid content in single organelles.
- Daniel C. Castro
- , Yuxuan Richard Xie
- & Jonathan V. Sweedler
-
Perspective |
 Mass spectrometry-based metabolomics: a guide for annotation, quantification and best reporting practices
Mass spectrometry-based metabolomics: a guide for annotation, quantification and best reporting practices
This Perspective, from a large group of metabolomics experts, provides best practices and simplified reporting guidelines for practitioners of liquid chromatography– and gas chromatography–mass spectrometry-based metabolomics.
- Saleh Alseekh
- , Asaph Aharoni
- & Alisdair R. Fernie
-
Article |
 DecoID improves identification rates in metabolomics through database-assisted MS/MS deconvolution
DecoID improves identification rates in metabolomics through database-assisted MS/MS deconvolution
DecoID provides acquisition-independent, database-assisted deconvolution of metabolomic MS/MS spectra for improved metabolite identification.
- Ethan Stancliffe
- , Michaela Schwaiger-Haber
- & Gary J. Patti
-
Article |
 SpaceM reveals metabolic states of single cells
SpaceM reveals metabolic states of single cells
SpaceM combines light microscopy and MALDI-imaging mass spectrometry to enable single-cell metabolic profiling of cultured cells. SpaceM reveals coexisting metabolic states of hepatocytes and aims to democratize single-cell metabolomics.
- Luca Rappez
- , Mira Stadler
- & Theodore Alexandrov
-
Article |
 Metabolomic profiling of single enlarged lysosomes
Metabolomic profiling of single enlarged lysosomes
Single-lysosome mass spectrometry (SLMS) integrates lysosomal patch-clamp recording and induced nanoESI/MS for concurrent metabolic and electrophysiological profiling of individual enlarged lysosomes.
- Hongying Zhu
- , Qianqian Li
- & Wei Xiong
-
Review Article |
 Ultra-high-performance liquid chromatography high-resolution mass spectrometry variants for metabolomics research
Ultra-high-performance liquid chromatography high-resolution mass spectrometry variants for metabolomics research
This Review surveys ultra-high-performance liquid chromatography high-resolution mass spectrometry (UHPLC–HRMS), a highly sensitive, high-throughput technique that is used for analyzing a broad range of metabolites.
- Leonardo Perez de Souza
- , Saleh Alseekh
- & Alisdair R. Fernie
-
Matters Arising |
 Reply to: Examining microbe–metabolite correlations by linear methods
Reply to: Examining microbe–metabolite correlations by linear methods
- James T. Morton
- , Daniel McDonald
- & Rob Knight
-
-
Research Highlight |
 Metabolic profiling of CD8+ T cells at the single-cell level
Metabolic profiling of CD8+ T cells at the single-cell level
A single-cell approach to metabolomics links metabolic profiles to cellular identity.
- Madhura Mukhopadhyay
-
Research Highlight |
 Label-free T cell characterization
Label-free T cell characterization
A machine learning model uses autofluorescence of key metabolites to predict T cell activation state.
- Madhura Mukhopadhyay
-
Correspondence |
 METLIN MS2 molecular standards database: a broad chemical and biological resource
METLIN MS2 molecular standards database: a broad chemical and biological resource
- Jingchuan Xue
- , Carlos Guijas
- & Gary Siuzdak
-
Brief Communication |
 Feature-based molecular networking in the GNPS analysis environment
Feature-based molecular networking in the GNPS analysis environment
Feature-based molecular networking allows the generation of molecular networks for mass spectrometry data that can recognize isomers, incorporate relative quantification and integrate ion mobility data.
- Louis-Félix Nothias
- , Daniel Petras
- & Pieter C. Dorrestein
-
Brief Communication |
 ReDU: a framework to find and reanalyze public mass spectrometry data
ReDU: a framework to find and reanalyze public mass spectrometry data
Repository-scale reanalysis of public mass spectrometry-based metabolomics data is facilitated by the Reanalysis of Data User (ReDU) interface, a system that uses consistent formatting and controlled vocabularies for metadata capture.
- Alan K. Jarmusch
- , Mingxun Wang
- & Pieter C. Dorrestein
-
Research Highlight |
Mass Spectrometry Search Tool (MASST)
Searching mass spectra against publicly available tandem MS databases just got easier.
- Arunima Singh
-
-
Technology Feature |
 Boost that metabolomic confidence
Boost that metabolomic confidence
Change is a constant in the burgeoning field of metabolomics. That includes data analysis tools and repositories.
- Vivien Marx
-
Article |
 Learning representations of microbe–metabolite interactions
Learning representations of microbe–metabolite interactions
mmvec, a neural-network-based algorithm, uses paired multiomics data (microbial sequence counts and metabolite abundances) to compute the conditional probability of observing a metabolite in the presence of a specific microorganism.
- James T. Morton
- , Alexander A. Aksenov
- & Rob Knight
-
Article |
 Multiplexed and single cell tracing of lipid metabolism
Multiplexed and single cell tracing of lipid metabolism
Cellular lipids, labeled with a charged reporter, yield characteristic MS1 and MS2 patterns during mass spectrometry. These reporters allow sample multiplexing and sensitive detection of lipid metabolism at single cell resolution.
- Christoph Thiele
- , Klaus Wunderling
- & Philipp Leyendecker
-
Brief Communication |
 A cheminformatics approach to characterize metabolomes in stable-isotope-labeled organisms
A cheminformatics approach to characterize metabolomes in stable-isotope-labeled organisms
A computational approach facilitates molecular formula, metabolite class, and structure assignment for plant metabolites on the basis of LC–MS analysis of fully 13C-labeled and unlabeled plants.
- Hiroshi Tsugawa
- , Ryo Nakabayashi
- & Kazuki Saito
-
Brief Communication |
 SIRIUS 4: a rapid tool for turning tandem mass spectra into metabolite structure information
SIRIUS 4: a rapid tool for turning tandem mass spectra into metabolite structure information
SIRIUS 4 is a fast and highly accurate tool for molecular structure interpretation from mass-spectrometry-based metabolomics data.
- Kai Dührkop
- , Markus Fleischauer
- & Sebastian Böcker
-
-
Brief Communication |
 XCMS-MRM and METLIN-MRM: a cloud library and public resource for targeted analysis of small molecules
XCMS-MRM and METLIN-MRM: a cloud library and public resource for targeted analysis of small molecules
A resource of multiple reaction monitoring–mass spectrometry transitions for quantitative analysis of biological small molecules is provided in METLIN-MRM, along with automated tools for analyzing such data in XCMS-MRM.
- Xavier Domingo-Almenara
- , J. Rafael Montenegro-Burke
- & Gary Siuzdak
-
Tools in Brief |
Metabolomes from metagenomics and modeling
-
Research Highlights |
 Tracking the protein–metabolite interactome
Tracking the protein–metabolite interactome
A method combining limited proteolysis with mass spectrometry systematically detects protein–metabolite interactions.
- Allison Doerr

 OpenMS 3 enables reproducible analysis of large-scale mass spectrometry data
OpenMS 3 enables reproducible analysis of large-scale mass spectrometry data RefMet: a reference nomenclature for metabolomics
RefMet: a reference nomenclature for metabolomics Tools for metabolomics
Tools for metabolomics