Research Highlight |
Featured
-
-
Article
| Open Access Molecular pixelation: spatial proteomics of single cells by sequencing
Molecular pixelation: spatial proteomics of single cells by sequencing
Molecular Pixelation is an optics-free method that uses DNA-tagged antibodies to enable identification of the relative location of proteins on single cells.
- Filip Karlsson
- , Tomasz Kallas
- & Simon Fredriksson
-
Article
| Open Access Quantification of absolute labeling efficiency at the single-protein level
Quantification of absolute labeling efficiency at the single-protein level
Super-resolution imaging of reference and target structures enables precise determination of the labeling efficiency of high-affinity binding proteins in cells for improved quantitative assessment of protein organization at the single-molecule level.
- Joschka Hellmeier
- , Sebastian Strauss
- & Ralf Jungmann
-
Article
| Open Access Virtual reality-empowered deep-learning analysis of brain cells
Virtual reality-empowered deep-learning analysis of brain cells
Generating training data for training deep-learning-based tools is time consuming. The DELiVR pipeline facilitates this process as demonstrated in this study on detecting c-Fos+ cells or microglia in the brain, following tissue clearing and imaging with light-sheet microscopy.
- Doris Kaltenecker
- , Rami Al-Maskari
- & Ali Ertürk
-
Article |
 Selective-plane-activation structured illumination microscopy
Selective-plane-activation structured illumination microscopy
The combination of light sheet illumination and reversibly switchable fluorophores enables improved structured illumination microscopy for fast, low-background super-resolution imaging in cells and spheroids.
- Kenta Temma
- , Ryosuke Oketani
- & Katsumasa Fujita
-
Article
| Open Access A multicolor suite for deciphering population coding of calcium and cAMP in vivo
A multicolor suite for deciphering population coding of calcium and cAMP in vivo
Improved green cAMP and red calcium sensors were developed to facilitate dual-color imaging in vivo. These sensors will allow studying the relationship between calcium and cAMP signaling.
- Tatsushi Yokoyama
- , Satoshi Manita
- & Masayuki Sakamoto
-
Brief Communication
| Open Access SpatialData: an open and universal data framework for spatial omics
SpatialData: an open and universal data framework for spatial omics
SpatialData is a user-friendly computational framework for exploring, analyzing, annotating, aligning and storing spatial omics data that can seamlessly handle large multimodal datasets.
- Luca Marconato
- , Giovanni Palla
- & Oliver Stegle
-
Analysis |
 Benchmarking spatial clustering methods with spatially resolved transcriptomics data
Benchmarking spatial clustering methods with spatially resolved transcriptomics data
A benchmark study compares 13 spatial clustering methods on spatial transcriptomics data.
- Zhiyuan Yuan
- , Fangyuan Zhao
- & Yi Zhao
-
News & Views |
 Unlocking cryo-EM’s multishot potential with square or rectangular beams
Unlocking cryo-EM’s multishot potential with square or rectangular beams
New condenser aperture designs form square or rectangular beams that match the camera dimensions, which efficiently expands the data acquisition area in cryogenic electron microscopy.
- Xiaowei Zhao
-
-
Article |
 Bright and stable monomeric green fluorescent protein derived from StayGold
Bright and stable monomeric green fluorescent protein derived from StayGold
mBaoJin is a monomeric derivative of the bright and photostable green fluorescent protein StayGold. mBaoJin offers favorable photophysical properties for use in diverse protein tagging and subcellular labeling applications.
- Hanbin Zhang
- , Gleb D. Lesnov
- & Fedor V. Subach
-
Research Briefing |
 VIBRANT: a phenotyping method for drug discovery using vibrational spectroscopy
VIBRANT: a phenotyping method for drug discovery using vibrational spectroscopy
We developed a high-content profiling method named vibrational painting (VIBRANT) for single-cell drug response measurements, combining vibrational imaging, multiplexed vibrational probes and machine learning. VIBRANT showed high performance in predicting drug mechanisms of action, discovering novel compounds and assessing drug combinations, demonstrating great promise for phenotypic drug discovery.
-
Article |
 VIBRANT: spectral profiling for single-cell drug responses
VIBRANT: spectral profiling for single-cell drug responses
Vibrational painting (VIBRANT) is a high-content single-cell phenotypic profiling method using mid-infrared imaging with vibrational probes for metabolic activity, which offers high accuracy with minimal batch effects to capture cellular responses to perturbation.
- Xinwen Liu
- , Lixue Shi
- & Wei Min
-
Editorial |
Where imaging and metrics meet
When it comes to bioimaging and image analysis, details matter. Papers in this issue offer guidance for improved robustness and reproducibility.
-
Article
| Open Access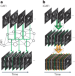 Content-aware frame interpolation (CAFI): deep learning-based temporal super-resolution for fast bioimaging
Content-aware frame interpolation (CAFI): deep learning-based temporal super-resolution for fast bioimaging
Content-aware frame interpolation (CAFI) improves the temporal resolution in time-lapse imaging by accurately predicting images in between image pairs. By allowing fewer frames to be imaged, CAFI also enables gentler live-cell imaging.
- Martin Priessner
- , David C. A. Gaboriau
- & Romain F. Laine
-
Article |
 Mapping enzyme activity in living systems by real-time mid-infrared photothermal imaging of nitrile chameleons
Mapping enzyme activity in living systems by real-time mid-infrared photothermal imaging of nitrile chameleons
Real-time mid-infrared photothermal imaging of nitrile chameleons enables simultaneous, multiplexed measurement of enzymatic activity in living systems and is poised to reveal the spatiotemporal regulation of enzymes in health and disease.
- Hongjian He
- , Jiaze Yin
- & Ji-Xin Cheng
-
Article
| Open Access Image restoration of degraded time-lapse microscopy data mediated by near-infrared imaging
Image restoration of degraded time-lapse microscopy data mediated by near-infrared imaging
InfraRed-mediated Image Restoration (IR2) uses deep learning to combine the benefits of deep-tissue imaging with NIR probes and the convenience of imaging with GFP for improved time-lapse imaging of embryogenesis.
- Nicola Gritti
- , Rory M. Power
- & Jan Huisken
-
Article |
 Smart lattice light-sheet microscopy for imaging rare and complex cellular events
Smart lattice light-sheet microscopy for imaging rare and complex cellular events
smartLLSM uses artificial intelligence-based instrument control to switch between epiflouorescence and lattice light-sheet microscopy to monitor cells at the population level while also capturing multicolor three-dimensional datasets of rare events of interest.
- Yu Shi
- , Jimmy S. Tabet
- & Wesley R. Legant
-
Article
| Open Access Thermal-plex: fluidic-free, rapid sequential multiplexed imaging with DNA-encoded thermal channels
Thermal-plex: fluidic-free, rapid sequential multiplexed imaging with DNA-encoded thermal channels
The thermal-plex method for highly multiplexed imaging uses DNA probes activated when briefly elevated to designated temperatures for rapid, fluidics-free sequential imaging in cells and tissues.
- Fan Hong
- , Jocelyn Y. Kishi
- & Peng Yin
-
Research Briefing |
 Ultra-long-working-distance multiphoton objective unlocks new possibilities for imaging
Ultra-long-working-distance multiphoton objective unlocks new possibilities for imaging
In 1858, the first standard for microscope objectives was established to encourage interchangeable components. Over the following 150 years, standards have evolved to constrain the size of objectives, which limits the parameters of working distance, field of view and resolution. A new design breaks out of this conventional envelope, offering an ultra-long working distance in air and enabling new neuroscience experiments.
-
Method to Watch |
 Human brain mapping
Human brain mapping
High-resolution connectomics of the human brain is the next frontier in neuroscience.
- Nina Vogt
-
Method to Watch |
 Imaging across scales
Imaging across scales
New twists on established methods and multimodal imaging are poised to bridge gaps between cellular and organismal imaging.
- Rita Strack
-
-
Article |
 Automated neuron tracking inside moving and deforming C. elegans using deep learning and targeted augmentation
Automated neuron tracking inside moving and deforming C. elegans using deep learning and targeted augmentation
Targettrack is a deep-learning-based pipeline for automatic tracking of neurons within freely moving C. elegans. Using targeted augmentation, the pipeline has a reduced need for manually annotated training data.
- Core Francisco Park
- , Mahsa Barzegar-Keshteli
- & Sahand Jamal Rahi
-
Article
| Open Access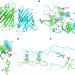 StayGold variants for molecular fusion and membrane-targeting applications
StayGold variants for molecular fusion and membrane-targeting applications
Monomeric and tandem dimer derivatives of the bright and photostable green fluorescent protein StayGold offer versatile tools for tagging target proteins and membranes in extended live-cell imaging.
- Ryoko Ando
- , Satoshi Shimozono
- & Atsushi Miyawaki
-
Article
| Open Access Next-generation MRI scanner designed for ultra-high-resolution human brain imaging at 7 Tesla
Next-generation MRI scanner designed for ultra-high-resolution human brain imaging at 7 Tesla
A combination of hardware developments has increased the achievable spatial resolution in 7 Tesla human neuroimaging to about 0.4 mm.
- David A. Feinberg
- , Alexander J. S. Beckett
- & Peter Dietz
-
Research Briefing |
 Fluorescent actinometers for fast and simple quantitative measurement of light intensity
Fluorescent actinometers for fast and simple quantitative measurement of light intensity
Fluorescent actinometers enable the measurement of light intensity even in the depths of samples and over wide ranges of wavelengths and intensities. We introduce two protocols to quantitatively characterize the spatial distribution of light of various fluorescence imaging systems and to calibrate the illumination of commercially available instruments and light sources.
-
Article
| Open Access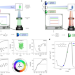 Fluorescence to measure light intensity
Fluorescence to measure light intensity
Two methods for fluorescence-based actinometry using organic dyes and photoconvertible fluorescent proteins enable rapid and precise measurement of light intensity at the sample in fluorescence microscopes.
- Aliénor Lahlou
- , Hessam Sepasi Tehrani
- & Ludovic Jullien
-
Article
| Open Access Uncovering developmental time and tempo using deep learning
Uncovering developmental time and tempo using deep learning
A deep learning-based method that uses microscopy images to stage embryos and analyze developmental time.
- Nikan Toulany
- , Hernán Morales-Navarrete
- & Patrick Müller
-
Article
| Open Access Bio-friendly long-term subcellular dynamic recording by self-supervised image enhancement microscopy
Bio-friendly long-term subcellular dynamic recording by self-supervised image enhancement microscopy
DeepSeMi is a self-supervised denoising framework that can enhance SNR over 12 dB across diverse samples and imaging modalities. DeepSeMi enables extended longitudinal imaging of subcellular dynamics with high spatiotemporal resolution.
- Guoxun Zhang
- , Xiaopeng Li
- & Qionghai Dai
-
-
Technology Feature |
 A sound solution for deep-brain imaging
A sound solution for deep-brain imaging
Ultrasound-based modalities are revealing the brain’s inner workings with steadily increasing speed, resolution and depth.
- Michael Eisenstein
-
Article
| Open Access Pulsed stimulated Brillouin microscopy enables high-sensitivity mechanical imaging of live and fragile biological specimens
Pulsed stimulated Brillouin microscopy enables high-sensitivity mechanical imaging of live and fragile biological specimens
A pulsed illumination scheme renders stimulated Brillouin microscopy less phototoxic and allows imaging of the mechanical properties of sensitive samples such as single cells, Caenorhabditis elegans embryos, zebrafish larvae and organoids.
- Fan Yang
- , Carlo Bevilacqua
- & Robert Prevedel
-
News & Views |
 Real-time imaging of dynamic tissues
Real-time imaging of dynamic tissues
A combined modality enables real-time imaging of mouse lungs and spans whole-organ to cellular scales.
- Joan E. Nichols
- & Sasha R. Azar
-
Article
| Open Access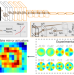 Deep learning-driven adaptive optics for single-molecule localization microscopy
Deep learning-driven adaptive optics for single-molecule localization microscopy
A deep learning approach bypasses iterative trials associated with sensorless adaptive optics to compensate for wavefront deformations when imaging biological specimens, enabling improved deep tissue localization microscopy.
- Peiyi Zhang
- , Donghan Ma
- & Fang Huang
-
Article
| Open Access D-LMBmap: a fully automated deep-learning pipeline for whole-brain profiling of neural circuitry
D-LMBmap: a fully automated deep-learning pipeline for whole-brain profiling of neural circuitry
D-LMBmap is a fully automated pipeline for mesoscale connectomics including deep-learning modules for axon segmentation, brain region segmentation and whole-brain registration. D-LMBmap works accurately across cell types and modalities.
- Zhongyu Li
- , Zengyi Shang
- & Jing Ren
-
Research Briefing |
 Mapping deformations and increasing quantitative accuracy in expansion microscopy
Mapping deformations and increasing quantitative accuracy in expansion microscopy
We introduce GelMap, a flexible workflow for reporting deformations and anisotropy in expansion microscopy. By intrinsically calibrating the expansion hydrogel using a fluorescent grid that scales with expansion and deforms with anisotropy, GelMap enables the reliable quantification of expansion factors and correction of deformations.
-
Article
| Open Access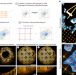 GelMap: intrinsic calibration and deformation mapping for expansion microscopy
GelMap: intrinsic calibration and deformation mapping for expansion microscopy
The GelMap workflow adds a fluorescent grid into samples before expansion, allowing for precise determination of expansion factor and subsequent deformation correction in ExM. GelMap works with diverse samples and expansion methods.
- Hugo G. J. Damstra
- , Josiah B. Passmore
- & Lukas C. Kapitein
-
Article |
 Large Stokes shift fluorescent RNAs for dual-emission fluorescence and bioluminescence imaging in live cells
Large Stokes shift fluorescent RNAs for dual-emission fluorescence and bioluminescence imaging in live cells
The Clivias are a series of small, monomeric fluorescent RNAs that emit with a large Stokes shift in the orange–red. They enable multiplexed RNA imaging in live cells and BRET-based detection of protein–RNA interactions in mice.
- Li Jiang
- , Xin Xie
- & Yi Yang
-
Article
| Open Access Statistically unbiased prediction enables accurate denoising of voltage imaging data
Statistically unbiased prediction enables accurate denoising of voltage imaging data
Statistically unbiased prediction utilizing spatiotemporal information in imaging data (SUPPORT) is a self-supervised deep learning approach to accurately denoise voltage and calcium imaging data while preserving true dynamic signals.
- Minho Eom
- , Seungjae Han
- & Young-Gyu Yoon
-
Article |
 Crystal ribcage: a platform for probing real-time lung function at cellular resolution
Crystal ribcage: a platform for probing real-time lung function at cellular resolution
A fabricated transparent ribcage facilitates real-time analysis of whole-lung function down to cellular resolution.
- Rohin Banerji
- , Gabrielle N. Grifno
- & Hadi T. Nia
-
Research Highlight |
 A closer look at chromatin
A closer look at chromatin
An expansion microscopy technique called ChromExM offers detailed views into the organization chromatin and associated gene expression machinery in embryos.
- Rita Strack
-
Registered Report |
 Quantitative assessment of near-infrared fluorescent proteins
Quantitative assessment of near-infrared fluorescent proteins
This Registered Report describes an extensive comparison of 22 near-infrared fluorescent proteins in vitro, in cultured mammalian cells, and in model animals, clarifying top performers in diverse biological settings.
- Hanbin Zhang
- , Stavrini Papadaki
- & Kiryl D. Piatkevich
-
News & Views |
 Advancing multiplexed imaging for enhanced tissue complexity analysis
Advancing multiplexed imaging for enhanced tissue complexity analysis
Multiplexed spatial immunophenotyping has advanced our understanding of tissues in the context of homeostasis and disease. Two studies now provide additional tools to overcome challenges with multiplexed imaging: one procedure amplifies the detection of low-abundance antigens by integrating SABER and IMC technologies, and the other is an X-ray-based method that enables the non-destructive multiplexed detection of antigens in tissues at scalable resolution and speed.
- Marieke E. Ijsselsteijn
- & Noel F. C. C. de Miranda
-
Brief Communication
| Open Access DNA-barcoded signal amplification for imaging mass cytometry enables sensitive and highly multiplexed tissue imaging
DNA-barcoded signal amplification for imaging mass cytometry enables sensitive and highly multiplexed tissue imaging
SABER-IMC combines DNA-based signal amplification by exchange reaction (SABER) with imaging mass cytometry (IMC) to enable simultaneous and highly multiplexed marker detection, even of low-abundance markers not detectable with IMC alone.
- Tsuyoshi Hosogane
- , Ruben Casanova
- & Bernd Bodenmiller
-
Article
| Open Access Multielement Z-tag imaging by X-ray fluorescence microscopy for next-generation multiplex imaging
Multielement Z-tag imaging by X-ray fluorescence microscopy for next-generation multiplex imaging
Multielement Z-tag X-ray fluorescence (MEZ-XRF) offers a new avenue for nondestructive and highly multiplexed tissue imaging and operates from the nanometer to whole-tissue scale, unlocking new biological observations.
- Merrick Strotton
- , Tsuyoshi Hosogane
- & Bernd Bodenmiller
-
Article |
 A fluorogenic chemically induced dimerization technology for controlling, imaging and sensing protein proximity
A fluorogenic chemically induced dimerization technology for controlling, imaging and sensing protein proximity
CATCHFIRE (chemically assisted tethering of chimera by fluorogenic-induced recognition) tools are small tags that can chemically dimerize with turn-on fluorescence, enabling simultaneous control and visualization of protein proximity.
- Sara Bottone
- , Octave Joliot
- & Arnaud Gautier
-
-
Research Highlight |
 In situ sequencing of localized translation
In situ sequencing of localized translation
Ribosome-bound mRNA mapping (RIBOmap) enables three-dimensional in situ detection of spatially resolved protein synthesis in single cells.
- Lei Tang
-
Research Briefing |
 Ultrafast nLight indicators for sensitive and specific in vivo imaging of norepinephrine
Ultrafast nLight indicators for sensitive and specific in vivo imaging of norepinephrine
We developed, characterized and validated nLight sensors, a new family of genetically encoded green and red fluorescent norepinephrine indicators based on an alpha-1 adrenergic receptor. nLight probes can detect norepinephrine in living animals with superior sensitivity, ligand specificity and temporal resolution as compared with previous tools.
Browse broader subjects
Browse narrower subjects
- Bioluminescence imaging
- 3-D reconstruction
- Diffusion tensor imaging
- Endoscopy
- Fluorescence imaging
- Functional magnetic resonance imaging
- Magnetic resonance imaging
- Magnetoencephalography
- Molecular imaging
- Optical imaging
- Positron-emission tomography
- Time-lapse imaging
- Ultrasound
- Viral tracing
- X-ray tomography

 Fast and multiplexed DNA-PAINT
Fast and multiplexed DNA-PAINT Neural networks for biomechanics
Neural networks for biomechanics Better detection for localization microscopy
Better detection for localization microscopy Art appreciation
Art appreciation