Featured
-
-
Article
| Open Access Electrospray-assisted cryo-EM sample preparation to mitigate interfacial effects
Electrospray-assisted cryo-EM sample preparation to mitigate interfacial effects
ESI-cryoPrep is a cryo-EM specimen preparation method that employs electrospray ionization techniques to deposit charged macromolecule-containing droplets on EM grids. Demonstrated across various protein samples, this approach effectively prevents biomolecule adsorption at air–water or graphene–water interfaces, addressing challenges related to protein denaturation and preferred orientation.
- Zi Yang
- , Jingjin Fan
- & Xiaoyu Zhou
-
Article
| Open Access Quantification of absolute labeling efficiency at the single-protein level
Quantification of absolute labeling efficiency at the single-protein level
Super-resolution imaging of reference and target structures enables precise determination of the labeling efficiency of high-affinity binding proteins in cells for improved quantitative assessment of protein organization at the single-molecule level.
- Joschka Hellmeier
- , Sebastian Strauss
- & Ralf Jungmann
-
Research Briefing |
 Simplifying deep learning to enhance accessibility of large-scale 3D brain imaging analysis
Simplifying deep learning to enhance accessibility of large-scale 3D brain imaging analysis
We created DELiVR, a deep-learning pipeline for 3D brain-cell mapping that is trained with virtual reality-generated reference annotations. It can be deployed via the user-friendly interface of the open-source software Fiji, which makes the analysis of large-scale 3D brain images widely accessible to scientists without computational expertise.
-
Article
| Open Access Virtual reality-empowered deep-learning analysis of brain cells
Virtual reality-empowered deep-learning analysis of brain cells
Generating training data for training deep-learning-based tools is time consuming. The DELiVR pipeline facilitates this process as demonstrated in this study on detecting c-Fos+ cells or microglia in the brain, following tissue clearing and imaging with light-sheet microscopy.
- Doris Kaltenecker
- , Rami Al-Maskari
- & Ali Ertürk
-
News & Views |
 LiMCA: Hi-C gets an RNA twist
LiMCA: Hi-C gets an RNA twist
A multiomics method measures both the cellular three-dimensional genome and transcriptome at the single-cell level.
- Jane Kawaoka
- & Stavros Lomvardas
-
This Month |
Quest: my postdoc home
For a postdoctoral fellowship, it’s advisable to be selective about lab choice and to be clear about expectations.
- Vivien Marx
-
Article
| Open Access Simultaneous single-cell three-dimensional genome and gene expression profiling uncovers dynamic enhancer connectivity underlying olfactory receptor choice
Simultaneous single-cell three-dimensional genome and gene expression profiling uncovers dynamic enhancer connectivity underlying olfactory receptor choice
LiMCA offers a tool for co-profiling 3D genome structure and gene expression at the single-cell level, enabling researchers to elucidate the olfactory receptor gene selection process.
- Honggui Wu
- , Jiankun Zhang
- & X. Sunney Xie
-
-
-
Editorial |
Peer review demystified: part 2
We continue our explanation of the peer review process at Nature Methods.
-
This Month |
 The placozoan Trichoplax
The placozoan Trichoplax
After having been largely neglected for more than a century, the tiny marine animal Trichoplax adhaerens is enjoying a revival and is now poised to become a mainstream model organism for the study of epithelial evolution, diversification and specialization.
- Marvin Leria
- , Magali Requin
- & Andrea Pasini
-
Article |
 Pretraining a foundation model for generalizable fluorescence microscopy-based image restoration
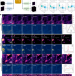
Pretraining a foundation model for generalizable fluorescence microscopy-based image restoration
A pretrained foundation model (UniFMIR) enables versatile and generalizable performance across diverse fluorescence microscopy image reconstruction tasks.
- Chenxi Ma
- , Weimin Tan
- & Bo Yan
-
Article |
 Selective-plane-activation structured illumination microscopy
Selective-plane-activation structured illumination microscopy
The combination of light sheet illumination and reversibly switchable fluorophores enables improved structured illumination microscopy for fast, low-background super-resolution imaging in cells and spheroids.
- Kenta Temma
- , Ryosuke Oketani
- & Katsumasa Fujita
-
Article
| Open Access DIP-MS: ultra-deep interaction proteomics for the deconvolution of protein complexes
DIP-MS: ultra-deep interaction proteomics for the deconvolution of protein complexes
Deep interactome profiling by mass spectrometry (DIP-MS) combines affinity purification with native BN-PAGE fractionation and mass spectrometry to resolve protein complexes sharing the same target protein. The paper also presents PPIprophet, a data-driven neural network-based protein complex deconvolution approach.
- Fabian Frommelt
- , Andrea Fossati
- & Matthias Gstaiger
-
Brief Communication
| Open Access Assessing GPT-4 for cell type annotation in single-cell RNA-seq analysis
Assessing GPT-4 for cell type annotation in single-cell RNA-seq analysis
This study evaluates the performance of GPT-4 in single-cell type annotation.
- Wenpin Hou
- & Zhicheng Ji
-
Article
| Open Access A multicolor suite for deciphering population coding of calcium and cAMP in vivo
A multicolor suite for deciphering population coding of calcium and cAMP in vivo
Improved green cAMP and red calcium sensors were developed to facilitate dual-color imaging in vivo. These sensors will allow studying the relationship between calcium and cAMP signaling.
- Tatsushi Yokoyama
- , Satoshi Manita
- & Masayuki Sakamoto
-
Article
| Open Access RoboEM: automated 3D flight tracing for synaptic-resolution connectomics
RoboEM: automated 3D flight tracing for synaptic-resolution connectomics
RoboEM enables automated proofreading of electron microscopy datasets using a strategy akin to that of self-steering cars. This decreases the need for manual proofreading of segmented datasets and facilitates connectomic analyses.
- Martin Schmidt
- , Alessandro Motta
- & Moritz Helmstaedter
-
Brief Communication
| Open Access SpatialData: an open and universal data framework for spatial omics
SpatialData: an open and universal data framework for spatial omics
SpatialData is a user-friendly computational framework for exploring, analyzing, annotating, aligning and storing spatial omics data that can seamlessly handle large multimodal datasets.
- Luca Marconato
- , Giovanni Palla
- & Oliver Stegle
-
Brief Communication
| Open Access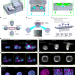 Open-top multisample dual-view light-sheet microscope for live imaging of large multicellular systems
Open-top multisample dual-view light-sheet microscope for live imaging of large multicellular systems
This work presents a highly versatile open-top, dual-view and dual-illumination light-sheet microscope for live imaging of large specimens.
- Franziska Moos
- , Simon Suppinger
- & Prisca Liberali
-
Analysis
| Open Access Multicenter integrated analysis of noncoding CRISPRi screens
Multicenter integrated analysis of noncoding CRISPRi screens
This analysis provides 108 noncoding CRISPR screens collated by the ENCODE4 consortium and establishes experimental guidelines for future CRISPRi screens characterizing functional cis-regulatory elements.
- David Yao
- , Josh Tycko
- & Steven K. Reilly
-
Analysis |
 Benchmarking spatial clustering methods with spatially resolved transcriptomics data
Benchmarking spatial clustering methods with spatially resolved transcriptomics data
A benchmark study compares 13 spatial clustering methods on spatial transcriptomics data.
- Zhiyuan Yuan
- , Fangyuan Zhao
- & Yi Zhao
-
News & Views |
 Unlocking cryo-EM’s multishot potential with square or rectangular beams
Unlocking cryo-EM’s multishot potential with square or rectangular beams
New condenser aperture designs form square or rectangular beams that match the camera dimensions, which efficiently expands the data acquisition area in cryogenic electron microscopy.
- Xiaowei Zhao
-
News & Views |
 Breaking up the StayGold dimer yields three photostable monomers
Breaking up the StayGold dimer yields three photostable monomers
The exceptionally photostable green fluorescent protein StayGold has been monomerized in different laboratories, which has generated three unique monomeric variants that will enable new imaging applications.
- Joachim Goedhart
- & Theodorus W. J. Gadella Jr
-
Brief Communication
| Open Access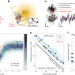 MINSTED tracking of single biomolecules
MINSTED tracking of single biomolecules
MINSTED quantifies tiny movements of individual biomolecules with high spatiotemporal precision to successfully resolve the steps of the molecular motor protein kinesin-1 labeled with a single fluorophore as it switches protofilaments.
- Lukas Scheiderer
- , Henrik von der Emde
- & Stefan W. Hell
-
-
-
-
-
Editorial |
Peer review demystified: part 1
Peer review is at the heart of publishing scientific papers. In this first installment of a two-part Editorial, we explain how we manage the process at Nature Methods.
-
Brief Communication |
 Square condenser apertures for square cameras in low-dose transmission electron microscopy
Square condenser apertures for square cameras in low-dose transmission electron microscopy
Square and rectangular beams made with new condenser apertures enable more efficient image capture in low-dose transmission electron microscopy with no compromise to image quality in single-particle cryoelectron microscopy.
- Hamish G. Brown
- , Dan Smith
- & Eric Hanssen
-
Article
| Open Access An optogenetic method for the controlled release of single molecules
An optogenetic method for the controlled release of single molecules
An optogenetic system enables the controlled release of soluble and transmembrane proteins for precise exploration of cellular protein function at the single-molecule level and streamlined single-molecule imaging.
- Purba Kashyap
- , Sara Bertelli
- & Helge Ewers
-
Article |
 Learning structural heterogeneity from cryo-electron sub-tomograms with tomoDRGN
Learning structural heterogeneity from cryo-electron sub-tomograms with tomoDRGN
TomoDRGN is a deep learning framework designed to model conformational and compositional heterogeneity from cryo-ET datasets on a per-particle basis.
- Barrett M. Powell
- & Joseph H. Davis
-
Article |
 Improved green and red GRAB sensors for monitoring spatiotemporal serotonin release in vivo
Improved green and red GRAB sensors for monitoring spatiotemporal serotonin release in vivo
Deng et al. expand the toolbox of neurotransmitter sensors with high-sensitivity green and red genetically encoded serotonin sensors. These are suitable for in vivo applications, as demonstrated in a variety of applications in mice.
- Fei Deng
- , Jinxia Wan
- & Yulong Li
-
This Month |
Early-career navigation tips
In a group interview, early-career researchers in proteomics share how they plan for the future and navigate divides over methods.
- Vivien Marx
-
Article |
 Bright and stable monomeric green fluorescent protein derived from StayGold
Bright and stable monomeric green fluorescent protein derived from StayGold
mBaoJin is a monomeric derivative of the bright and photostable green fluorescent protein StayGold. mBaoJin offers favorable photophysical properties for use in diverse protein tagging and subcellular labeling applications.
- Hanbin Zhang
- , Gleb D. Lesnov
- & Fedor V. Subach
-
Article |
 scGPT: toward building a foundation model for single-cell multi-omics using generative AI
scGPT: toward building a foundation model for single-cell multi-omics using generative AI
Pretrained using over 33 million single-cell RNA-sequencing profiles, scGPT is a foundation model facilitating a broad spectrum of downstream single-cell analysis tasks by transfer learning.
- Haotian Cui
- , Chloe Wang
- & Bo Wang
-
Article
| Open Access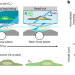 Super-sectioning with multi-sheet reversible saturable optical fluorescence transitions (RESOLFT) microscopy
Super-sectioning with multi-sheet reversible saturable optical fluorescence transitions (RESOLFT) microscopy
Multi-sheet RESOLFT combines the speed and optical sectioning of light-sheet fluorescence microscopy with reversibly photoswitchable fluorescent proteins to enable fast, volumetric super-resolution imaging in live cells.
- Andreas Bodén
- , Dirk Ollech
- & Ilaria Testa
-
Article |
 A-SOiD, an active-learning platform for expert-guided, data-efficient discovery of behavior
A-SOiD, an active-learning platform for expert-guided, data-efficient discovery of behavior
A-SOiD is a computational platform for behavioral annotation whose training includes elements of supervised and unsupervised learning. The approach is demonstrated on mouse, macaque and human datasets.
- Jens F. Tillmann
- , Alexander I. Hsu
- & Eric A. Yttri
-
Research Briefing |
 VIBRANT: a phenotyping method for drug discovery using vibrational spectroscopy
VIBRANT: a phenotyping method for drug discovery using vibrational spectroscopy
We developed a high-content profiling method named vibrational painting (VIBRANT) for single-cell drug response measurements, combining vibrational imaging, multiplexed vibrational probes and machine learning. VIBRANT showed high performance in predicting drug mechanisms of action, discovering novel compounds and assessing drug combinations, demonstrating great promise for phenotypic drug discovery.
-
Research Briefing |
 A biomimetic robotic antigen-presenting system for sensitive T cell recognition
A biomimetic robotic antigen-presenting system for sensitive T cell recognition
We introduce a biomimetic antigen-presenting system that uses hexapod heterostructures for specific T cell recognition at the single-molecule and single-cell levels. The system enables high-resolution T cell activation, uses magnetic forces to increase immune responses, and offers flexible and precise identification of antigen-specific T cell receptors, aiding the study of T cell recognition and immune cell mechanics.
-
Research Briefing |
 TREX identifies region-specific protein interactors of RNA molecules
TREX identifies region-specific protein interactors of RNA molecules
Interactions between RNA and RNA-binding proteins (RBPs) define the fate and function of every RNA molecule. We present TREX, or targeted RNase H-mediated extraction of crosslinked RBPs, an efficient and accurate method to unbiasedly reveal the protein interactors of specific regions of RNAs isolated from living cells.
-
Article |
 VIBRANT: spectral profiling for single-cell drug responses
VIBRANT: spectral profiling for single-cell drug responses
Vibrational painting (VIBRANT) is a high-content single-cell phenotypic profiling method using mid-infrared imaging with vibrational probes for metabolic activity, which offers high accuracy with minimal batch effects to capture cellular responses to perturbation.
- Xinwen Liu
- , Lixue Shi
- & Wei Min
-
Article
| Open Access TREX reveals proteins that bind to specific RNA regions in living cells
TREX reveals proteins that bind to specific RNA regions in living cells
TREX introduces an RNA-centric tool for identifying proteins binding to specific RNA regions in living cells.
- Martin Dodel
- , Giulia Guiducci
- & Faraz K. Mardakheh
-
Article
| Open Access Generation of complex bone marrow organoids from human induced pluripotent stem cells
Generation of complex bone marrow organoids from human induced pluripotent stem cells
The authors describe stem cell-derived bone marrow organoids that accurately model the structural and functional properties of the human bone marrow niche.
- Stephanie Frenz-Wiessner
- , Savannah D. Fairley
- & Christoph Klein
-
Correspondence |
 OpenMS 3 enables reproducible analysis of large-scale mass spectrometry data
OpenMS 3 enables reproducible analysis of large-scale mass spectrometry data
- Julianus Pfeuffer
- , Chris Bielow
- & Timo Sachsenberg
-
Article
| Open Access Multiscale biochemical mapping of the brain through deep-learning-enhanced high-throughput mass spectrometry
Multiscale biochemical mapping of the brain through deep-learning-enhanced high-throughput mass spectrometry
MEISTER is an integrative experimental and computational framework for mass spectrometry that integrates three-dimensional, organ-wide biomolecular mapping with single-cell analysis for multiscale profiling of spatial–biochemical organization.
- Yuxuan Richard Xie
- , Daniel C. Castro
- & Fan Lam
-
Article
| Open Access Toward universal cell embeddings: integrating single-cell RNA-seq datasets across species with SATURN
Toward universal cell embeddings: integrating single-cell RNA-seq datasets across species with SATURN
SATURN performs cross-species integration and analysis using both single-cell gene expression and protein representations generated by protein language models.
- Yanay Rosen
- , Maria Brbić
- & Jure Leskovec
-
Research Briefing |
 Deciphering protein interaction network dynamics with a machine learning-based framework
Deciphering protein interaction network dynamics with a machine learning-based framework
We developed Tapioca, an integrative ensemble machine learning-based framework, to accurately predict global protein–protein interaction network dynamics. Tapioca enabled the characterization of host regulation during reactivation from latency of an oncogenic virus. Introducing an interactome homology analysis method, we identified a proviral host factor with broad relevance for herpesviruses.
-
Article |
 Tapioca: a platform for predicting de novo protein–protein interactions in dynamic contexts
Tapioca: a platform for predicting de novo protein–protein interactions in dynamic contexts
Tapioca is an ensemble machine learning framework for studying protein–protein interactions (PPIs) that facilitates integration of curve-based dynamic PPI data from thermal proximity coaggregation, ion-based proteome-integrated solubility alteration or cofractionation mass spectrometry data with static interaction data to predict PPIs in dynamic contexts.
- Tavis. J. Reed
- , Matthew. D. Tyl
- & Ileana. M. Cristea
Browse broader subjects
Browse narrower subjects
- Analytical biochemistry
- Behavioural methods
- Bioinformatics
- Biological models
- Biophysical methods
- Cytological techniques
- Electrophysiology
- Epigenetics analysis
- Experimental organisms
- Gene delivery
- Gene expression analysis
- Genetic engineering
- Genetic techniques
- Genomic analysis
- High-throughput screening
- Imaging
- Immunological techniques
- Isolation, separation and purification
- Lab-on-a-chip
- Mass spectrometry
- Metabolomics
- Microbiology techniques
- Microscopy
- Molecular engineering
- Nanobiotechnology
- Optogenetics
- Proteomic analysis
- Sensors and probes
- Sequencing
- Software
- Optical spectroscopy
- Structure determination

 Post-translational modification-centric base editor screens to assess phosphorylation site functionality in high throughput
Post-translational modification-centric base editor screens to assess phosphorylation site functionality in high throughput Protein–ligand structure prediction
Protein–ligand structure prediction Graphene sandwich for cryo-EM
Graphene sandwich for cryo-EM Neural networks for biomechanics
Neural networks for biomechanics