Abstract
Hydrocarbon pollution is a major ecological problem facing oil-producing countries, especially in the Niger Delta region of Nigeria. In this study, a site that had been previously polluted by artisanal refining activity was investigated using 16S rRNA Illumina high-throughput sequencing technology and bioinformatics tools. These were used to investigate the bacterial diversity in soil with varying degrees of contamination, determined with a gas chromatography-flame ionization detector (GC-FID). Soil samples were collected from a heavily polluted (HP), mildly polluted (MP), and unpolluted (control sample, CS) portion of the study site. DNA was extracted using the Zymo Research (ZR) Fungi/Bacteria DNA MiniPrep kit, followed by PCR amplification and agarose gel electrophoresis. The microbiome was characterized based on the V3 and V4 hypervariable regions of the 16S rRNA gene. QIIME (Quantitative Insights Into Microbial Ecology) 2 software was used to analyse the sequence data. The final data set covered 20,640 demultiplexed high-quality reads and a total of 160 filtered bacterial OTUs. Proteobacteria dominated samples HP and CS, while Actinobacteria dominated sample MP. Denitratisoma, Pseudorhodoplanes, and Spirilospora were the leading genera in samples HP, CS, and MP respectively. Diversity analysis indicated that CS [with 25.98 ppm of total petroleum hydrocarbon (TPH)] is more diverse than HP (with 490,630 ppm of TPH) and MP (with 5398 ppm of TPH). A functional prediction study revealed that six functional modules dominated the dataset, with metabolism covering up to 70%, and 11 metabolic pathways. This study demonstrates that a higher hydrocarbon concentration in soil adversely impacts microbial diversity, creating a narrow bacterial diversity dominated by hydrocarbon-degrading species, in addition to the obvious land and ecosystem degradation caused by artisanal refining activities. Overall, the artisanal refining business is significantly driving ecosystem services losses in the Niger Delta, which calls for urgent intervention, with focus on bioremediation.
Similar content being viewed by others
Introduction
The Niger Delta, located in Nigeria, is a well-researched area in Africa, renowned for its potential for oil exploration and its diverse ecosystem. It spans 70,000 km2, making it the largest wetland in Africa and the third-largest in the world1. Furthermore, it holds the most substantial oil reserves (36.2 billion barrels) and gas reserves (185 billion cubic feet) in Africa2. The history of commercial oil and gas exploration in the Niger Delta goes back to 1958, which has led to an estimated 3.1 billion barrels of crude oil being discharged, affecting over 5000 sites from 1967 to 2014. This has resulted in the region being identified as one of the most polluted areas globally3,4,5. Studies have shown that oil pollution in the region is attributed to inadequate servicing and maintenance facilities, routine oil operations, sabotage, accidental/equipment failures, deliberate release of oil wastes into the environment, and currently, artisanal oil refining activities6,7,8. Petroleum pollution releases various types of hydrocarbons (saturates, aromatics, resins, and asphaltenes) into the environment, ultimately impacting the terrestrial ecosystem. This impact arises from their ability to disrupt the healthy ecological balance9, terminating sensitive ecological receptor10, contaminate groundwater, pose health challenges10, as well as reduce, microbial composition11,12. The shift in microbial structure favours sentinel microorganisms that metabolize those complex hydrocarbons as sources of carbon and energy for growth and other physiological processes13,14.
Petroleum hydrocarbon-degrading bacteria are abundant in contaminated habitats15,16. Autochthonous bacteria with this specialty are ubiquitous in the terrestrial ecosystem and are key to the natural attenuation and detoxification of such impacted lands17. Shen et al.18 evaluated the bacterial diversity of polluted soil by means of metagenomics and confirmed the dominance of Actinobacteria. A diversity study conducted by Obieze et al.19 demonstrated that populations of hydrocarbon-utilizing bacteria were highest when the concentration of pollutants were high. Chikere et al.20 used 16S rRNA technology to show that the shift of bacterial genera moved from a mixed group to Gram-negative bacteria with Betaproteobacteria dominance.
Bacterial diversity studies on terrestrial ecosystem impacted by artisanal crude oil refining are an emerging area in the Niger Delta. This necessitates MiSeq-driven metagenomics, which is a reliable Next-Generation Sequencing (NGS) technology for evaluating bacterial compositional structure, diversity, and function. In this study, we investigated bacterial diversity insights in polluted soil using MiSeq sequencing technology to understand how varying hydrocarbon concentrations affect the distribution of bacteria.
Results
Physicochemical analysis
The analysis of soil texture indicated that gravel constituted 5.2%, sand comprised 93.2%, clay constituted 4.4%, and silt accounted for 2.4%. Various chemical parameters, encompassing pH, electrical conductivity, total nitrogen, and total organic carbon, were evaluated for both the contaminated soil and control soil samples, with the results summarized in Table 1. The pH values for the control, mildly polluted, and heavily polluted soils were recorded as 6.50, 6.30, and 7.10, respectively. The electrical conductivity (EC) values for the control, mildly polluted, and heavily polluted soils were 30 µS/cm, 40 µS/cm, and 120 µS/cm, respectively. Similarly, the total nitrogen values were 2.67 mg/kg, 2.02 mg/kg, and 2.00 mg/kg for the control, mildly polluted, and heavily polluted soils, respectively. Additionally, the total organic carbon was 1.25%, 5.25%, and 19.61% for the control, mildly polluted, and heavily polluted soils, respectively.
TPH analysis
The values of the TPH for the heavily polluted (HP), control sample (CS) and mildly polluted (MP) soil sample are 490,630, 25.98 ppm, and 5398 ppm respectively.
Bacterial diversities from unpolluted, mildly polluted and highly polluted soils
The final dataset from the three reference samples—heavily polluted (HP), unpolluted (control sample; CS), and mildly polluted (MP)—comprised 20,640 demultiplexed high-quality reads, with an average of 6880 reads per sample. The sequences were clustered into 256 bacterial OTUs, which were further filtered to a minimum count of 4 and 20% prevalence per sample, resulting in a total of 160 retained bacterial OTUs. The dominant bacterial phyla in the heavily polluted soil (HP) samples were Proteobacteria (66%), Firmicutes (27%), Acidobacteria (4%), and Actinobacteria at less than 3%. In the unpolluted soil (CS) sample, the dominant phyla were Proteobacteria (68%), Firmicutes (30%), and Actinobacteria at less than 2%. In the mildly polluted soil (MP), the dominant phyla were Actinobacteria (39%), Proteobacteria (31%), Firmicutes (29%), Acidobacteria (0.67%), and Planctomycetes (0.33%). Figure 1 summarises the bacterial phylum distribution in the three reference points. Dominant genera in the HP sample are Denitratisoma, Clostridium, Alkaliphilus, Diplorickettsia, Methylosinus and Bacillus (in descending order: ranging from 30.56% to 1.99%). The following genera (Pseudorhodoplanes, Cohnella, Rhodovastum, Neobacillus, Neomegalonema, Acidomonas, Neobacillus, Salirhabdus) are duplicated in HP and CS samples, however, at relatively low abundance (< 2.0%) in HP but was dominant genera in CS. These genera are not observed in MP, except for Pseudorhodoplanes (2.7%). The dominant bacterial genera in MP include Spirilospora (34%), Paenibacillus, Swionibacillus, Rhizorhapis, Endobacter, Paraburkholderia, Rhodopila, Mycoavidus, Rummeliibacillus, Quasibacillus, and Phenylobacterium. Genera such as Denitratisoma, Alkaliphilus, Clostridium, and Bacillus were observed in CS but not in MP samples, while Methylosinus was observed in MP and not in CS. Figure 1a shows the dominant phyla diversity across the three samples, while Fig. 1b displays the relative abundance of the ten most dominant genera. Figures 1c–e depict that the bacterial abundance in CS is significantly different from both HP and MP (at p = 0.05). Similarly, as displayed in Table 2, CS is higher in richness and diversity as compared to the HP and MP samples.
Relative abundance of bacteria. Relative abundance at phylum (a) and genus level (b). An extended error bar plot was used to compare the top 10 genera between two samples with p < 0.05 as shown: S1 and S5 (c); S1 and S7 (d) and S7 and S5 (e). S1 Heavily polluted (HP), S5 Control sample (CS), S7 Mildly polluted (MP).
Inferred bacterial function by PICRUSt2
A total of 99 KOs (KEGG Orthologues) were predicted, and the superpathway was used for the plot. The predicted KEGG (Kyoto Encyclopedia of Genes and Genomes) orthologues, collapsed into MetaCyc meta-pathways, show abundance values for each sample using the PICRUSt methodology (Fig. 2). Six functional modules, comprising cellular processes (3%), environmental information processing (7%), genetic information processing (5%), human diseases (5%), and metabolism (70%), make up almost 90% of the complete dataset in the samples. The higher percentage of metabolism is registered in the order of MP, HP and CS respectively, indicating that hydrocarbon serves as a source of carbon or energy or both. Differentially abundant pathways (at p ≤ 0.05) showed 11 pathways (Fig. 3), with oxidative phosphorylation as the most dominant biomarker pathway. Heavily polluted (HP) sample is more pronounced with disease function, with a notable genus such as Diplorickettsia, an agent of tick-borne infection. CS leads in genetic information processing, while MP leads in cellular processes.
The link between taxonomy and predicted function. (a) Shows the abundance of predicted function and phylum. (b) Shows the relative abundance of each category of the predicted function. BURRITO, a visualization tool for exploratory data analysis of metagenomic data was used to visualize taxonomy linked to predicted function. S1 Heavily polluted (HP), S5 Control sample (CS), S7 Mildly polluted (MP).
An overall comparison of KEGG metabolic sub-pathway of the category metabolism for top three phyla (Proteobacteria, Firmicutes and Actinobacteria) as inferred by PICRUSt2. (a–c) depict significant (q-value > 0.05) features between two groups, calculated using White’s non-parametric test with Benjamini–Hochberg FDR (false discovery rate) in STAMP. S1 Heavily polluted (HP), S5 Control sample (CS), S7 Mildly polluted (MP).
Discussion
Total petroleum hydrocarbons (TPH) consist of various fractions of petroleum, including polyaromatic hydrocarbons (PAHs). TPH in soil is primarily derived from anthropogenic sources, but some fractions of PAHs are contributed by living organisms such as plants and microorganisms21. The major contribution of TPH in the study site was artisanal activities, which include spills, explosions, combustion, deposition, and the burial of the heavy fraction regarded as waste. Intrinsic factors such as soil texture, temperature, biosynthesis, topography, and erosion contribute to the overall concentration of soil TPH. The last two factors play an influential role in the unequal distribution of TPH in the soil. The concentration of the control sample (25.98 ppm) is well below the 100 ppm threshold required as a clean-up standard22. The mildly polluted soil (with value of 5398 ppm) and heavily polluted soil (with a value of 490,360 ppm) is quite significant and is comparable to that TPH concentration obtained by Martinez et al.23. The mildly polluted soil TPH concentration exceeds the Nigerian regulatory standard, 5000 ppm24. The artisanal refining operations in the Niger Delta have a profound impact on the environment as a whole. The pollution that originates from artisanal refining sites is characterized by extremely poor air quality25, contamination of surface soil and groundwater26, loss of vegetation and mangroves27 and heavy pollution of the marine ecosystem28. The air, consistently laden with hydrocarbons, has the potential to cause and exacerbate respiratory diseases29 such as reduced lung function, bronchitis, asthma, lung cancer, and chronic obstructive pulmonary disease30,31,32. The contamination of surface soil from artisanal refineries leads to a significant loss of vegetation cover and arable land, as well as a drastic alteration of microbial diversity. Contaminated groundwater poses a risk to both animals and humans, serving as a source of unsafe and toxic water. The disposal of the heavy-end fraction of hydrocarbons and other affiliated wastes (as is usually the practice) in water bodies remains a crude method of waste management. Consequently, the marine ecosystem becomes heavily polluted, resulting in a severe loss of mangrove habitats33,34. Seafood from the marine ecosystem is affected, leading to bioaccumulation and biomagnification35,36, which ultimately results in cancer and other malignant growths in humans. Overall, artisanal refining activities lead to health challenges, decreased agricultural productivity, reduction in the means of livelihood in local communities, and significant losses in biodiversity and ecosystem services, implicating microorganisms.
Microorganisms react to various situations of hydrocarbon pollution in different ways: they either develop resistance or utilize hydrocarbons as a source of carbon and energy. Those that are unable to adapt to the stress of hydrocarbon pollution are eliminated12. In all the samples analysed, there were changes in the structure of biodiversity compared to the unpolluted control. The dynamics of the phylum shift, reflecting the dominance of Proteobacteria in the heavily polluted (HP) and control sample (CS) soil samples and not in the mildly polluted (MP) is similar to those observed in the study conducted by Kim et al.37. The Firmicutes did not respond significantly to each of the samples as did Proteobacteria, Actinobacteria, and Acidobacteria. The dominance of Proteobacteria in both heavily polluted and unpolluted soils accounts for the relevant roles they play in the biogeochemical cycling of carbon, sulphur, and nitrogen38, as well as plant fitness and growth promotion39. The shift noticed in the mildly polluted sample favoured the dominance of Actinobacteria. Genera in this group thrive in hydrocarbon-polluted environment, highlighting their physiological and genomic adaptations to challenging conditions40. These bacteria play a crucial role in diverse ecological processes, such as the biodegradation of complex molecules41, involvement in biogeochemical cycles42, deterioration of artefacts, and participation in biological weathering43. Actinobacteria demonstrate proficiency in these functions owing to their exceptional capabilities in DNA repair and protection, protein synthesis, biofilm formation, as well as the synthesis of biosurfactants, secondary metabolites, and essential enzymes44,45,46,47. Notably, certain genera such as Geodermatophilus, Modestobacter, and Kocuria, within the Actinobacteria phylum, exhibit resistance to desiccation, heavy metal toxicity, and ionization, as reported by Sayed et al.48, Shivlata and Satyanarayana49 and Guesmi et al.50 Consequently, Actinobacteria represent a valuable resource for the bioremediation of highly contaminated environments. The trend shows that Acidobacteria diversities decreased significantly in the mildly polluted soil. This study demonstrates that hydrocarbon contamination gradients influence Proteobacteria, Acidobacteria and Actinobacteria as Abena et al.51 showed.
The prevalent genera identified in the heavily polluted (HP) sample include Clostridium (Firmicutes), Methylosinus (Alphaproteobacterial methanotroph), and Bacillus (Firmicutes) among other genera including Denitratisoma (Betaproteobacteria) and Daegula (Alphaproteobacteria). The last two genera are the two most dominant in the HP sample. Clostridium species have the requisite enzyme system52, genetic repertoire9,53 and cell surface properties54, to access and degrade hydrocarbon (especially halogenated species) under anaerobic condition. Methylosinus is a methanotrophic bacteria that has proven proficiency in degrading methane and chlorinated hydrocarbon55,56 in consortium with other hydrocarbon-degrading bacteria and communities57. Bacillus spp. is a widely distributed prolific bacteria known for their ability to metabolise hydrocarbon taking advantage of their ability to form biofilm58, genomic capacity59, biosurfactant production60, metabolic diversity61, and favourable redox potentials62. Some species are hydrocarbonoclastic63, thus effecting an increased population density as they utilise hydrocarbon as source of energy and carbon. While Denitratisoma and Daegula exhibit the highest population density in the heavily polluted soil, there is limited information available about them. However, it is established that Denitratisoma functions as aerobic denitrifiers64 and plays a crucial role in rhizoremediation65. Their reduced presence in the mildly and unpolluted polluted soil suggest that they may have affinity for highly polluted environment. Genera like Neomegalonema, Neobacillus, Acidomonas, Pseudorhodoplanes, Cohnella, Rhodovastum, and Salirhabdus, each comprising less than 2% in the highly polluted sample, exhibited high abundance in the unpolluted sample, indicating their sensitivity to hydrocarbon pollution. However, these genera did not show in the mildly polluted soil, except Pseudorhodoplanes. This suggests that Pseudorhodoplanes is an excellent hydrocarbon-degrading bacteria or an emerging hydrocarbonoclastic bacteria. This bacteria genus has been implicated in hydrocarbon degradation12. Tirandaz et al.66 illustrated that P. sinuspersici exhibits optimal activity at a temperature of 30 °C and a pH of 7. However, it demonstrates tolerance within a pH range of 5.5 to 8 and a temperature range of 15 to 35 °C. It is interesting to note that the study site reflects these optimal parameters, including suitable soil (sandy loamy) type. Other genera that appeared prominent in the unpolluted control sample that has been reported to affiliate with hydrocarbon polluted soil are Cohnella67, Rhodovastum68 and Salirhabdus69. Genera that shifted from being rare in the unpolluted sample to becoming more prominent in the polluted samples can be considered as emerging hydrocarbon-degrading bacteria (or tolerant taxa), and they include Spirilospora, Swionibacillus, and Paenibacillus.
The catalogue of hydrocarbon-degrading bacteria is abundant with genera like Paenibacillus, Paraburkholderia, Methylosinus, and Phenylobacterium. In the mildly polluted soil examined in this study, these four genera are present, along with the identification of less common bacterial genera, including Spirilospora, which appears to be the most prevalent bacterial genus. Paenibacillus spp. (also abundant in the heavily polluted sample) has been shown to degrade hydrocarbon in consortium with Gordonia, Cupriavidus spp70., is associated with rhizoremediation, produce biosurfactants in contaminated soil, harbour hydrocarbon-degrading pahE genes and other requisite genes71. These key biomarkers and other biological factors have positioned Paenibacillus spp. to degrade PAH72 and transform heavy crude to light oil73. Kanwal et al.74 had indicated that sporogeneisis permits Paenibacillus, Bacillus and other related bacteria to survive inhospitable environments. Some species of Paenibacillus has been described as hydrocarbonoclastic75 and at the same time diazotrophic76 in the total environment, highlighting their dual relevance in bioremediation and plant growth promotion. Diazotrophic Paenibacillus polymyxa has been implicated in hydrocarbon degradation77 specifically polyaromatic hydrocarbon78 and biodegradation of mixed pesticides79. However, there are more diazostrophic Paenibacillus spp. than hydrocarbon-degrading diazotrophic Paenibacillus spp. confirmed by publicly available literature80,81,82. From the key hydrocarbon-degrading bacteria examined, it is worthy of note that hydrocarbon degradation is linked to biological factors such as adaptation, metabolic competence, genetics, enzyme system, biomass, biosurfactant production, cell surface property, microbial interaction, biofilm formation and cell’s redox potential. Activation of a considerable number of these attributes has defined Paraburkholderia, Methylosinus, and Phenylobacterium as hydrocarbon-degrading bacteria and possibly Spirilospora and Pseudorhodoplanes, noted as emerging hydrocarbon-degrading bacteria83,84,85. Paraburkholderia aromaticivorans BN5 has been reported to degrade aliphatic hydrocarbons, naphthalene and BTEX83, while Methylosinus spp. is an obligate methane metabolizer86 apart from degrading hydrocarbon through cometabolic pathway57. In addition, Methylosinus has species that are diazotrophic87, and heavy metal detoxifiers. For its part Phenylobacterium spp. has been reported in PAH degradation88. Signature sequences in the mildly polluted soil is an indication of hydrocarbon metabolism, some of which may represent emerging hydrocarbon-degraders while some may be affiliated with diazotrophism. The diversity of these signature sequences is critical to the understanding of their structure in relation to the heavily polluted (HP) and control (CS) soil samples.
The OTU’s diversity index shows that CS is higher in richness (by observed OTUs), while the Shannon diversity index shows that the CS sample is higher in diversity. The higher diversity index in CS reflects the unpolluted nature of the control sample and the toxicity effect of obvious pollution in HP and MP samples. The consequence of hydrocarbon impact include the shift of broad microbial diversity characterised with broad ecological functions to a narrow microbial diversity with prominent hydrocarbon-degrading and hydrocarbon-tolerant phylotypes. Examination of the mildly and heavily polluted samples reflect a few number of notable and emerging hydrocarbon-degrading bacteria including Paenibacillus, Paraburkholderia, Methylosinus Phenylobacterium, Bacillus, Burkholderia, Alkaliphilus, Pseudorhodoplanes and others. The reason behind this “shift to the left’ phenomenon is the adaptation of keystone species to survive in a stressed ecosystem either as resistant or utilizers of hydrocarbon as source of carbon and energy. Bacteria that can tolerate and degrade moderate hydrocarbons, in their mixed form, can initiate biodegradation process, as much as those that can tolerate and benefit from the process’ metabolites89. This phenomenon becomes prominent in ecosystem with long-term history of pollution90. Results from most studies in pollution ecology and bioremediation align with the concept of broad-to-narrow concept, confirmed in this study. The trend observed in this study counters that of Benedek et al.91 which observed positive correlation between TPH (147,000 ppm) and diversity. The reasons, according to Benedek et al.91, for this negative results are long-term exposure to hydrocarbon, significant rise of a particular hydrocarbon-degrading bacterial genus, need for alternative carbon source and lack of humus. Recent studies that support positive correlation between TPH and microbial diversity in the Niger Delta are Iturbe-Espinoza et al.90, Edet and Antai92 and in other regional settings are Lee et al.93, Mukherjee et al.94 and Yerulker et al.95. These two contradicting research outcome suggests that besides, ecotoxicity effects, other factors play influential roles in diversity profiling96. These factors may include soil chemistry, soil’s trophic status, and genera composition in higher taxa. These factors condition a non-uniform trend and responses of microbial taxa to contamination in soil.
In pollution-affiliated microbial diversity dynamics, dominance of hydrocarbon-degrading bacteria is a common phenomenon, which is underscored by biological functions such as cellular processes, genetic information and degradation. Hydrocarbons are utilised as an energy source through oxidative phosphorylation, in the inner section of bacteria cytoplasm with the release of ATP for cellular processes: growth, replication, quorum sensing, chemotaxis, and catabolism. The latter is achieved through the use of key signatory enzymes. Though an enzyme profile was not conducted in this study, Obieze et al.19 reported hydrocarbon-degrading enzymes (through functional prediction) in the same study site, which include (3S,4R)-3,4-dichloroxycyclohexa-1,5-diene-1,4-dicarboxylate dehydrogenase, 2,4-diclorophenol-6-monooxygenase, 3-carboxyethylcatechol-2,3-dioxygenase. Bidja-Abena et al.51 identified a few functional enzymes (in diesel-polluted soil) specific for xenobiotic metabolisms, chlorobenzene degradation, and polyaromatic hydrocarbon. Protein export and gene repair (indices of genetic information processing) are protective against hydrocarbon toxicity to cells97,98. Another KEGG functional profile that is connected to metabolism is environmental information processing such as the ABC transporters for mineral/organic ions, amino acids, and lipid transportation38. Protein transportation is necessary for uptake of hydrocarbons by bacteria for degradation and metabolism. These functional module is dominated by metabolism with 11 pathways, which support active engagement of hydrocarbon degradation in this study.
Concluding remarks
This work was undertaken to understand the impact of artisanal refining activities on soil bacterial diversity through metagenomics. Consequently, Ngia Ama was chosen because of its hydrocarbon pollution history of more than six years. Composite samples of heavily (HP) polluted, mildly (MP) polluted and unpolluted (CS) soil were used for the analysis and the results showed a broad-to-narrow bacterial diversity with known and emerging hydrocarbon-degrading bacteria found abundantly in polluted samples. This paradigm shifts in bacteria diversity points to distortion of ecological service at the detriment of the total environmental and its receptors in addition to the vicious impact of artisanal refining activities on vegetation and marine ecosystem in the Niger Delta. However, the study area features as a hub of activated soil critical for ex situ bioremediation programme.
Methodology
Site description
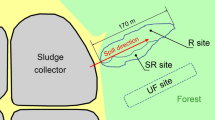
The study was carried out on contaminated fallow land at Ngia Ama (4°47ʹ42ʺ N, 6°51ʹ45ʺE), a community in Tombia Kingdom where illegal refining activities had occurred for over six years, with respect to the sampling year, 2018. The study site is enveloped by mangroves and creeks, featuring moderate lowlands and an average temperature of 25–37 °C.
Soil sample collection
The heavily (HP) and moderately (MP) polluted portions (7.5 ft away from each other) of the study site were spotted and soil sampling was carried out using a soil auger. Three sub-samples were collected at each point from a depth of ≤ 30 cm and mixed thoroughly to create a composite sample. A similar approach was used to collect a control sample (CS, non-polluted), 23 ft away from the polluted field. The samples were aseptically transferred into sterile plastic containers and preserved at 4 °C and − 20 °C for downstream analysis.
Soil’s physicochemical analysis
The soil samples collected were initially dried at 25 °C, ground, and then sieved through a 2 mm mesh before analysis, following the method outlined by Durak et al.99. The Bouyoucos Hydrometer method was employed for soil texture analysis, following the protocol adopted by Babalola et al.100, to determine the content of sand, silt, and clay. In summary, a beaker was filled with 50 g of pre-treated soil and 125 ml of sodium hexametaphosphate (40 g/L) was added. The mixture was stirred until the soil was fully saturated and then left to rest for ten minutes. The resulting soil slurry was moved to a mixer, and distilled water was added until the mixing cup was half full. The solution was then mixed for two minutes. Subsequently, the soil slurry was quickly transferred to an unoccupied sedimentation cylinder, and distilled water was added up to the reference mark. The cylinder was flipped upside down and back 30 times. After placing the cylinder down, the time was noted. The stopper was taken off the cylinder, and a hydrometer was gently inserted. The initial hydrometer reading was taken immediately, followed by a second reading after 15 s. Further hydrometer readings were taken in a doubling pattern until 48 h, resulting in a total of 16 readings. The collected data was then analysed to determine the composition of soil particles. The pH level of the soil was assessed using the method outlined by Adekiya et al.101. To measure the soil’s pH, 10 g of soil was placed into a clean 100 mL beaker, to which 20 mL of deionized water was added. A pH tester 20 was then inserted into the resulting suspension in the beaker, with the aim of determining the average pH from three repeated measurements. The analysis of the electrical conductivity (EC) was carried out as previously detailed Oyem and Oyem102. This involved adding 10 g of soil to 20 mL of deionized water and allowing it to stand for 30 min. The resulting slurry was then filtered, and the EC was measured using a Hanna digital conductivity meter. The organic carbon content was ascertained using the method reported by Mrayyan and Battikhi103 introducing 1 g of soil into 10 mL of 1.0 M of K2Cr4O7 and the mixture was shaken for homogeneity. Later, 20 mL of 98% H2SO4 was rapidly added using a burette and shaken with vigour for 1 min and left standing on a white tile for 30 min. The mixture was then added with 200 mL of deionized water and later with 10 mL of 85% H3PO4, 0.2 g NaF and 15 drops of diphenylamine indicator. The ensuing solution was back-titrated with 0.5 N iron(II) sulphate and organic carbon calculated104.
The Kjeldahl method was used to determine the total nitrogen content. Initially, 10 g of the sample was weighed and placed into a 500 mL Kjeldahl flask. Then, 20 mL of deionized water was added to the flask, which was shaken for a few minutes and left to stand for 30 min. Copper and sodium sulphate (1.5 g each), along with 30 mL of concentrated sulphuric acid, were added and mixed until homogeneous. The contents of the flask were heated until no froth was visible. The mixture was then boiled for 5 h, cooled, and 100 mL of distilled water was added to the flask. A boric acid indicator (50 mL) was used to rinse the sandy residue, which was then added to a conical flask positioned under the condenser of the distillation setup. The Kjeldahl flask, including the digest, was connected to the distillation unit. Sodium hydroxide (150 mL of 10 M NaOH) was added to the distillation flask and distilled until 150 mL of distillate was collected. The nitrogen content/concentration was calculated using a titration technique with a 0.01 M sulphuric acid distillate. The endpoint was indicated by a colour change from green to pink. A blank titration was also performed to obtain the blank titre104. The total nitrogen was calculated using Eq. (1).
where n represent normality of acid.
Determination of total petroleum hydrocarbons (TPH)
Two grams of soil sample were heated at 50 °C and crushed well afterwards. Ten millilitres (10 ml) of dichloromethane (Sigma Aldrich, USA) was then added to the finely crushed soil and shaken firmly. To precipitate the soil, it was centrifuged at 3000×g for 10 min105. The solvent phase was removed. The TPH analysis was carried out following steps earlier prescribed by106. In summary, the hydrocarbon portion was stirred for 5 mins and separated using a Whatman filter paper No. 42. The extracted hydrocarbon was concentrated to 1 mL after being evaporated in a water bath. The TPH was determined using a GC spectrometer (Thermo Scientific™ Nicolet iCS). The samples were run in triplicate. The procedural blank was determined by going through the extraction and clean-up procedures using glass beads instead of a soil sample.
Next-generation sequencing for metagenomic analyses
Metagenomic DNA extraction
The DNA extraction was carried out on the samples using Zymo Research (ZR) Fungi/Bacteria DNA MiniPrep™ (California, USA) supplied by Inqaba Biotec, South Africa according to the manufacturer’s instructions. The summary of the extraction process is illustrated in Fig. 4. In summary, 0.25 g of soil is added to a ZR BashingBead™ Lysis Tube along with 750 μl of Lysis Solution. The tube is then processed in a bead beater at maximum speed for at least 5 min. Following centrifugation at 10,000×g for 1 min, up to 400 μl of the supernatant is transferred to a Zymo-Spin™ IV Spin Filter, and after centrifuging at 8000×g for 1 min, the filtrate is combined with 1200 μl of Fungal/Bacterial DNA Binding Buffer. Subsequently, 800 μl of this mixture is loaded onto a Zymo-Spin™ IIC Column and centrifuged at 10,000×g for 1 min, with a repeat of the step. The Zymo-Spin™ IIC Column is then treated with 200 μl of DNA Pre-Wash Buffer and centrifuged for 1 min at 10,000×g, followed by the addition of 500 μl Fungal/Bacterial DNA Wash Buffer and another round of centrifugation. The Zymo-Spin™ IIC Column is transferred to a clean 1.5 ml microcentrifuge tube, and 100 μl of DNA Elution Buffer is added directly to the column matrix. The elution is achieved by centrifuging at 10,000×g for 30 s, resulting in the extraction of DNA suitable for downstream analysis107.
Polymerase chain reaction (PCR)
For the PCR analysis, the hypervariable region (V3-V4) of the 16S rRNA was targeted using bacteria-specific primers, namely 341F and 806R. The primers were tagged according to108. All the PCRs were done in triplicate (n = 3). Polymerase chain reaction master mix aliquot was dispensed into PCR tubes and the different DNA samples were introduced into each tube alongside a negative control. The PCR reagents in each tube amounted to 50 μl containing: buffer (5 μl: 100 mM), MgCl2 (1.5 μl: 25 mM), universal primer1 (2 μl forward: 20 μM), primer2 (2 μl reverse: 20 μM), dNTP mix (1 μl: 200 μM), Dream Taq DNA Polymerase (0.25 μl: 1.25 units/50 μl), sterile water (35.25 μl) and DNA samples (3 μl)109,110,111. The PCR condition was set at 3 min at 94 °C, followed by 35 cycles comprising 30 s at 94 °C, 30 s at 58 °C, and 1 min at 72 °C61. The process concluded with a final extension step of 10 min at 72 °C. The PCRs were performed using an MJ Mini thermal cycler (Bio-Rad, Hercules, CA, USA).
The resulting amplicons were separated electrophoretically with 1% agarose gel stained with 0.1 μg/ml ethidium bromide running at 80 V for 60 min, using TAE electrophoresis buffer. The PCR amplicons were visualized by UV fluorescence to determine the amplicon sizes. The PCR products (20 μl each) were later cleaned up using 160 μl of 13% polyethene glycol (PEG) 8000, 20 μl of 5 M NaCl solution and 200 μl of 70% ethanol.
MiSeq sequencing and sequence analysis
The PCR products (after purification using Omega, Bio-Tek and quantification with Agilent Bioanalyzer 2100) were sequenced with the Miseq platform at the University of South Africa (UNISA), Science Campus, Florida, Roodepoort. This process involved 600 cycles (300 cycles for each paired read and 12 cycles for the barcode sequence) as per the manufacturer’s guidelines. This also involved 600 cycles (300 cycles for each paired read and 12 cycles for the barcode sequence) following the manufacturer’s instructions. The sequence data was analysed using the 16S-based metagenomics workflow provided by MiSeq Reporter v2.3 (Illumina). The 16S rRNA gene, a frequently targeted region, was used for microbial identification, thereby eliminating the need to sequence the entire genome. The Illumina workflow began with purified genomic DNA, where primers were extended with sequences that included indexing barcodes. The samples were then merged into a single library and sequenced on the Illumina MiSeq platform, resulting in paired 230 bp reads112,113.
Bioinformatic analyses
Demultiplexed paired-end reads obtained from the sequencing facility were quality-checked using FastQC software version 0.11.5 (Babraham Institute, United Kingdom). Subsequently, Trimmomatic software (version 0.38)114 was used to quality-trim paired reads, including clipping off any Illumina barcodes and eliminating reads with an average quality score (Phred Q score) lower than 20. Quality-filtered paired reads were then analyzed in the Quantitative Insights into Microbial Ecology (version 2) (QIIME2) software115. DADA2 denoiser116 was used to merge pair-end sequences into full-length sequences as well as remove chimaras. USEARCH version 7 was used to cluster similar sequences into operational taxonomic units (OTUs) at 97% similarity117. Taxonomic classification of the clustered OTUs was performed against the RDP classifier118. The obtained OTU table was further rarefied to even depths of 7544 sequences. The OTU and sequences of clustered OTUs were used as an input to PICRUSt2 software (installed as a QIIME2 plugin) to predict metabolic functions119 based on 16S rRNA. PICRUSt2 was developed in 2020 as an improvement over the 2013 version. It is more accurate and features a larger database. PICRUSt2 is a promising tool with the potential for various research applications. For instance, it could be employed to investigate the functional potential of microbial communities in different environments, as demonstrated in this study. The bacterial communities’ relative abundance was visualized at the phylum and genus level to better convey the biological information in these samples. The OTU table with assigned taxonomy was taxonomy was normalized (relative abundance) using MicrobiomeAnalyst120; and used to plot 100% stacked bar graph.
Statistical analysis
QIIME2 output—OTU table was in text and biom format. OTU table in text format was imported into Rstudio and ranacapa (ranacapa::runRanacapa()) package was used for rarefication curve, Shannon index and Observed OTUs calculations121 The biom format of the OTU table was uploaded to MicrobiomeDB—A data-mining platform for interrogating microbiome experiments was used to determine the top 10 abundant genera. These genera were then compared between the samples where significant (q-value > 0.05) features between two groups (HP-MP, HP-CS and CS-MP) calculated using White’s non-parametric test with Benjamini–Hochberg FDR (false discovery rate) in STAMP. The same test was used for PICRUSt2 predicted function.
Data availability
The metagenomic datasets generated and analysed during the current study are available in the National Centre of Biotechnology Information (NCBI) Sequence Read Archive (SRA) database under the accession number PRJNA563161 for public access.
References
Izah, S. Ecosystem of the Niger-Delta region of Nigeria: Potentials and threats. Biodivers. Int. J. 2, 338–345 (2018).
Kadafa, A. A. Oil exploration and spillage in the Niger Delta of Nigeria. Civ. Environ. Res. 2, 38–51 (2012).
Enyoghasim, M. O. Oil exploration and exploitation in Nigeria and the challenge of sustainable development: An assessment of the Niger Delta. 670216917 (2019).
Nwozor, A., Audu, J. & Adama, I. J. The political economy of hydrocarbon pollution: Assessing socio-ecological sustainability of Nigeria’s Niger Delta region. Int. J. Energy Econ. Policy 9, 1–8 (2018).
Obida, C. B., Blackburn, G. A., Whyatt, J. D. & Semple, K. T. Quantifying the exposure of humans and the environment to oil pollution in the Niger Delta using advanced geostatistical techniques. Environ. Int. 111, 32–42 (2018).
Albert, O., Amaratunga, D. & Haigh, R. An investigation into root causes of sabotage and vandalism of pipes: A major environmental hazard in Niger Delta, Nigeria. in ASCENT Festival 2019: International Conference on Capacity Building for Research and Innovation in Disaster Resilience 22–37 (National Science Foundation of Sri Lanka, 2019).
Elum, Z. A., Mopipi, K. & Henri-Ukoha, A. Oil exploitation and its socioeconomic effects on the Niger Delta region of Nigeria. Environ. Sci. Pollut. Res. 23, 12880–12889 (2016).
Osuji, L. Some environmental hazards of oil pollution in Niger Delta, Nigeria. Afr. J. Interdisc. Stud. 3, 11–17 (2002).
Ramírez, D. et al. Bacterial diversity in surface sediments from the continental shelf and slope of the North West gulf of Mexico and the presence of hydrocarbon degrading bacteria. Mar. Pollut. Bull. 150, 110590 (2020).
Cocârţă, D. M., Stoian, M. A. & Karademir, A. Crude oil contaminated sites: evaluation by using risk assessment approach. Sustainability 9, 1365 (2017).
Neethu, C. S., Saravanakumar, C., Purvaja, R., Robin, R. S. & Ramesh, R. Oil-spill triggered shift in indigenous microbial structure and functional dynamics in different marine environmental matrices. Sci. Rep. 9, 1–13 (2019).
Xu, X. et al. Petroleum hydrocarbon-degrading bacteria for the remediation of oil pollution under aerobic conditions: A perspective analysis. Front. Microbiol. 9, 2885 (2018).
Das, N. & Chandran, P. Microbial degradation of petroleum hydrocarbon contaminants: An overview. Biotechnol. Re. Int. 2011, 1–13 (2011).
Nzila, A. Current status of the degradation of aliphatic and aromatic petroleum hydrocarbons by thermophilic microbes and future perspectives. Int. J. Environ. Res. Public Health 15, 2782 (2018).
Ali, S. S. et al. Recent advances in the life cycle assessment of biodiesel production linked to azo dye degradation using yeast symbionts of termite guts: A critical review. Energy Rep. 8, 7557–7581 (2022).
Mironov, O. G. Black Sea microorganisms growing on hydrocarbons. Mikrobiologiia 38, 728–731 (1969).
Koshlaf, E. & Ball, A. S. Soil bioremediation approaches for petroleum hydrocarbon polluted environments. AIMS Microbiol. 3, 25 (2017).
Shen, Z. et al. Bacterial diversity in surface sediments of collapsed lakes in Huaibei, China. Sci. Rep. 12, 1–12 (2022).
Obieze, C. C. et al. Field-scale biostimulation shifts microbial community composition and improves soil pollution recovery at an artisanal crude oil refining site. Int. J. Environ. Stud. 198, 1–20 (2022).
Chikere, C. B., Azubuike, C. C. & Fubara, E. M. Shift in microbial group during remediation by enhanced natural attenuation (RENA) of a crude oil-impacted soil: a case study of Ikarama Community, Bayelsa, Nigeria. 3 Biotech 7, 1–11 (2017).
Krauss, M. et al. Atmospheric versus biological sources of polycyclic aromatic hydrocarbons (PAHs) in a tropical rain forest environment. Environ. Pollut. 135, 143–154 (2005).
Kuppusamy, S., Maddela, N. R., Megharaj, M. & Venkateswarlu, K. Impact of total petroleum hydrocarbons on human health. in Total Petroleum Hydrocarbons, 139–165 (Springer, 2020).
Gallegos Martínez, M. et al. Diagnostic and resulting approaches to restore petroleum-contaminated soil in a Mexican tropical swamp. Water Sci. Technol. 42, 377–384 (2000).
Olawuyi, D. S. & Tubodenyefa, Z. Review of the Environmental Guidelines and Standards for the Petroleum Industry in Nigeria (EGASPIN). (OGEES Institute, 2018).
Onakpohor, A., Fakinle, B. S., Sonibare, J. A., Oke, M. A. & Akeredolu, F. A. Investigation of air emissions from artisanal petroleum refineries in the Niger-Delta Nigeria. Heliyon 6, 05608 (2020).
Azuamah, K. O., Appiah-Effah, E. & Akodwaa-Boadi, K. Water quality index, ecotoxicology and human health risk modelling of surface and groundwater along illegal crude oil refining sites in a developing economy. Heliyon 9, 20631 (2023).
Onuh, P. A. et al. Artisanal refining of crude oil in the Niger Delta: A challenge to clean-up and remediation in Ogoniland. Local Econ. 36, 468–486 (2021).
Ite, A. E., Harry, T. A., Obadimu, C. O., Asuaiko, E. R. & Inim, I. J. Petroleum hydrocarbons contamination of surface water and groundwater in the Niger Delta region of Nigeria. J. Environ. Pollut. Hum. Health 6, 51–61 (2018).
Xue, Y., Chu, J., Li, Y. & Kong, X. The influence of air pollution on respiratory microbiome: A link to respiratory disease. Toxicol. Lett. 334, 14–20 (2020).
Viegi, G., Maio, S., Pistelli, F., Baldacci, S. & Carrozzi, L. Epidemiology of chronic obstructive pulmonary disease: Health effects of air pollution. Respirology 11, 523–532 (2006).
Ko, F. W. S. & Hui, D. S. C. Air pollution and chronic obstructive pulmonary disease. Respirology 17, 395–401 (2012).
Duan, R.-R., Hao, K. & Yang, T. Air pollution and chronic obstructive pulmonary disease. Chronic Dis. Transl. Med. 6, 260–269 (2020).
Onyena, A. P. & Sam, K. A review of the threat of oil exploitation to mangrove ecosystem: Insights from Niger Delta, Nigeria. Glob. Ecol. Conserv. 22, e00961 (2020).
Das, S. Ecological restoration and livelihood: Contribution of planted mangroves as nursery and habitat for artisanal and commercial fishery. World Dev. 94, 492–502 (2017).
D’Costa, A., Shyama, S. K. & Kumar, M. P. Bioaccumulation of trace metals and total petroleum and genotoxicity responses in an edible fish population as indicators of marine pollution. Ecotoxicol. Environ. Saf. 142, 22–28 (2017).
Akinsanya, B., Adebusoye, S. A., Alinson, T. & Ukwa, U. D. Bioaccumulation of polycyclic aromatic hydrocarbons, histopathological alterations and parasito-fauna in bentho-pelagic host from Snake Island, Lagos, Nigeria. JoBAZ 79, 40 (2018).
Kim, D., Choi, K. Y., Yoo, M., Zylstra, G. J. & Kim, E. Biotechnological Potential of Rhodococcus Biodegradative Pathways. (2018).
Zhou, Z., Tran, P. Q., Kieft, K. & Anantharaman, K. Genome diversification in globally distributed novel marine Proteobacteria is linked to environmental adaptation. ISME J. 14, 2060–2077 (2020).
Dube, J. P., Valverde, A., Steyn, J. M., Cowan, D. A. & Van der Waals, J. E. Differences in bacterial diversity, composition and function due to long-term agriculture in soils in the eastern free State of South Africa. Diversity 11, 61 (2019).
Saadouli, I. et al. Diversity and adaptation properties of actinobacteria associated with Tunisian stone ruins. Front. Microbiol. 13, 7832 (2022).
Baoune, H. et al. Petroleum degradation by endophytic Streptomyces spp. isolated from plants grown in contaminated soil of southern Algeria. Ecotoxicol. Environ. Saf. 147, 602–609 (2018).
Sun, S., Ma, B., Wang, G. & Tan, X. Linking microbial biogeochemical cycling genes to the rhizosphere of pioneering plants in a glacier foreland. Sci. Total Environ. 872, 161944 (2023).
Cockell, C. S., Kelly, L. C. & Marteinsson, V. Actinobacteria: An ancient phylum active in volcanic rock weathering. Geomicrobiol. J. 30, 706–720 (2013).
Behera, S. & Das, S. Potential and prospects of Actinobacteria in the bioremediation of environmental pollutants: Cellular mechanisms and genetic regulations. Microbiol. Res. 273, 127399 (2023).
Farda, B., Djebaili, R., Vaccarelli, I., Del Gallo, M. & Pellegrini, M. Actinomycetes from caves: An overview of their diversity, biotechnological properties, and insights for their use in soil environments. Microorganisms 10, 453 (2022).
Alvarez, A. et al. Actinobacteria: Current research and perspectives for bioremediation of pesticides and heavy metals. Chemosphere 166, 41–62 (2017).
El-Naggar, N. E.-A. Streptomyces-based cell factories for production of biomolecules and bioactive metabolites. in Microbial Cell Factories Engineering for Production of Biomolecules, 183–234 (Elsevier, 2021).
Sayed, A. M. et al. Extreme environments: Microbiology leading to specialized metabolites. J. Appl. Microbiol. 128, 630–657 (2020).
Shivlata, L. & Satyanarayana, T. Thermophilic and alkaliphilic Actinobacteria: Biology and potential applications. Front. Microbiol. 6, 1014 (2015).
Guesmi, S. et al. Roots of the xerophyte Panicum turgidum host a cohort of ionizing-radiation-resistant biotechnologically-valuable bacteria. Saudi J. Biol. Sci. 29, 1260–1268 (2022).
Bidja Abena, M. T. et al. Microbial diversity changes and enrichment of potential petroleum hydrocarbon degraders in crude oil-, diesel-, and gasoline-contaminated soil. 3 Biotech 10, 1–15 (2020).
Sethunathan, N. & Yoshida, T. Degradation of chlorinated hydrocarbons by Clostridium sp. isolated from lindane-amended, flooded soil. Plant Soil 38, 663–666 (1973).
Tan, B. et al. Comparative analysis of metagenomes from three methanogenic hydrocarbon-degrading enrichment cultures with 41 environmental samples. ISME J. 9, 2028–2045 (2015).
Carrié, M., Gabelle, J.-C., Lopes-Ferreira, N. & Velly, H. Enzymatic breakdown of biofilm matrix to allow flow cytometry viability analysis of Clostridium beijerinckii cells. J. Appl. Microbiol. 134, 062 (2023).
Fenibo, E. O., Selvarajan, R., Wang, H., Wang, Y. & Abia, A. L. K. Untapped talents: Insight into the ecological significance of methanotrophs and its prospects. Sci. Total Environ. 903, 166145 (2023).
Arinageswaran, S., Balaji, E., Srimuthu, G. & Uma, S. Methanotrophs and its metabolic cascade of biotransformation of methane into valuable products. Agric. Allied Sci. (2023).
Lee, S.-W., Keeney, D. R., Lim, D.-H., Dispirito, A. A. & Semrau, J. D. Mixed pollutant degradation by Methylosinus trichosporium OB3b expressing either soluble or particulate methane monooxygenase: Can the tortoise beat the hare?. Appl. Environ. Microbiol. 72, 7503–7509 (2006).
Elumalai, P. et al. Characterization of crude oil degrading bacterial communities and their impact on biofilm formation. Environ. Pollut. 286, 117556 (2021).
Saeed, M. et al. Biodegradation of PAHs by Bacillus marsiflavi, genome analysis and its plant growth promoting potential. Environ. Pollut. 292, 118343 (2022).
Thirumurugan, D. et al. Impact of biosurfactant produced by Bacillus spp. on biodegradation efficiency of crude oil and anthracene. Chemosphere 344, 140340 (2023).
Wu, Y. et al. The metagenomics of soil bacteria and fungi and the release of mechanical dormancy in hard seeds. Front. Plant Sci. 14, 1187416 (2023).
Zeng, J., Zhu, Q., Wu, Y. & Lin, X. Oxidation of polycyclic aromatic hydrocarbons using Bacillus subtilis CotA with high laccase activity and copper independence. Chemosphere 148, 1–7 (2016).
Das, S., Das, N., Choure, K. & Pandey, P. Biodegradation of asphaltene by lipopeptide-biosurfactant producing hydrocarbonoclastic, crude oil degrading Bacillus spp. Bioresour. Technol. 382, 129198 (2023).
Xia, Z. et al. Nitrogen removal pathway and dynamics of microbial community with the increase of salinity in simultaneous nitrification and denitrification process. Sci. Total Environ. 697, 134047 (2019).
Kawasaki, A., Watson, E. R. & Kertesz, M. A. Indirect effects of polycyclic aromatic hydrocarbon contamination on microbial communities in legume and grass rhizospheres. Plant Soil 358, 169–182 (2012).
Tirandaz, H. et al. Pseudorhodoplanes sinuspersici gen. nov., sp. nov., isolated from oil-contaminated soil. Int. J. Syst. Evol. Microbiol. 65, 4743–4748 (2015).
Xu, S. et al. Total petroleum hydrocarbons and influencing factors in co-composting of rural sewage sludge and organic solid wastes. Environ. Pollut. 319, 120911 (2023).
Ulrich, N. et al. Response of aquatic bacterial communities to hydraulic fracturing in northwestern Pennsylvania: A five-year study. Sci. Rep. 8, 5683 (2018).
Al-Kindi, S. & Abed, R. M. M. Effect of biostimulation using sewage sludge, soybean meal, and wheat straw on oil degradation and bacterial community composition in a contaminated desert soil. Front. Microbiol. 7, 240 (2016).
Construction of a hydrocarbon-degrading consortium and characterization of two new lipopeptides biosurfactants. ScienceDirect. https://www.sciencedirect.com/science/article/abs/pii/S004896971936396X.
pahE, a functional marker gene for polycyclic aromatic hydrocarbon-degrading bacteria. Appl. Environ. Microbiol. https://doi.org/10.1128/aem.02399-18.
Al-Thukair, A. A., Malik, K. & Nzila, A. Biodegradation of selected hydrocarbons by novel bacterial strains isolated from contaminated Arabian Gulf sediment. Sci. Rep. 10, 21846 (2020).
Shibulal, B. et al. The potential of indigenous Paenibacillus ehimensis BHP for recovering heavy crude oil by biotransformation to light fractions. PLoS ONE 12, e0171432 (2017).
Kanwal, M. et al. Biodegradation of petroleum hydrocarbons and the factors effecting rate of biodegradation. Am. J. Biomed. Sci. Res 16, 6 (2022).
Mesbaiah, F. Z. et al. Preliminary characterization of biosurfactant produced by a PAH-degrading Paenibacillus sp. under thermophilic conditions. Environ. Sci. Pollut. Res. 23, 14221–14230 (2016).
Dashti, N. et al. Most hydrocarbonoclastic bacteria in the total environment are diazotrophic, which highlights their value in the bioremediation of hydrocarbon contaminants. Microb. Environ. 30, 70–75 (2015).
Langendries, S. & Goormachtig, S. Paenibacillus polymyxa, a Jack of all trades. Environ. Microbiol. 23, 5659–5669 (2021).
Timmusk, S., Teder, T. & Behers, L. Paenibacillus polymyxa A26 and its surfactant-deficient mutant degradation of polycyclic aromatic hydrocarbons. Stresses 1, 266–276 (2021).
Zhang, X. et al. Study on the simultaneous degradation of five pesticides by Paenibacillus polymyxa from Panax ginseng and the characteristics of their products. Ecotoxicol. Environ. Saf. 168, 415–422 (2019).
Li, Y., Li, Y., Zhang, H., Wang, M. & Chen, S. Diazotrophic Paenibacillus beijingensis BJ-18 provides nitrogen for plant and promotes plant growth, nitrogen uptake and metabolism. Front. Microbiol. 10, 1119 (2019).
Tang, Q., Puri, A., Padda, K. P. & Chanway, C. P. Biological nitrogen fixation and plant growth promotion of lodgepole pine by an endophytic diazotroph Paenibacillus polymyxa and its GFP-tagged derivative. Botany 95, 611–619 (2017).
Beneduzi, A. et al. Genome sequence of the diazotrophic gram-positive rhizobacterium Paenibacillus riograndensis SBR5 T. J. Bacteriol. 193, 6391–6392 (2011).
Lee, Y., Lee, Y. & Jeon, C. O. Biodegradation of naphthalene, BTEX, and aliphatic hydrocarbons by Paraburkholderia aromaticivorans BN5 isolated from petroleum-contaminated soil. Sci. Rep. 9, 860 (2019).
Margesin, R., Volgger, G., Wagner, A. O., Zhang, D. & Poyntner, C. Biodegradation of lignin monomers and bioconversion of ferulic acid to vanillic acid by Paraburkholderia aromaticivorans AR20-38 isolated from Alpine forest soil. Appl. Microbiol. Biotechnol. 105, 2967–2977 (2021).
Hwangbo, M., Shao, Y., Hatzinger, P. B. & Chu, K. Acidophilic methanotrophs: Occurrence, diversity, and possible bioremediation applications. Environ. Microbiol. Rep. 15, 265–281 (2023).
Sahoo, K. K., Goswami, G. & Das, D. Biotransformation of methane and carbon dioxide into high-value products by methanotrophs: Current state of art and future prospects. Front. Microbiol. 12, 636486 (2021).
Minamisawa, K. Mitigation of greenhouse gas emission by nitrogen-fixing bacteria. Biosci. Biotechnol. Biochem. 87, 7–12 (2023).
Chicca, I., Becarelli, S. & Di Gregorio, S. Microbial involvement in the bioremediation of total petroleum hydrocarbon polluted soils: Challenges and perspectives. Environments 9, 52 (2022).
Shibulal, B. et al. Heavy crude oil biodegradation: Catechol dioxygenase gene copy number variation determination by droplet digital polymerase chain reaction. Geomicrobiol. J. 40, 295–306 (2023).
Iturbe-Espinoza, P. et al. Microbial communities associated with landfarming amendments during bioremediation of crude oil in Niger Delta soils. Appl. Soil Ecol. 191, 105058 (2023).
Remarkable impact of PAHs and TPHs on the richness. Google Scholar. https://scholar.google.com/scholar?hl=en&as_sdt=0%2C5&q=Remarkable+impact+of+PAHs+and+TPHs+on+the+richness+and+diversity+of+bacterial+species+in+surface+soils+exposed+to+long-term+hydrocarbon+pollution&btnG=.
Edet, U. O. & Antai, S. P. Correlation and distribution of xenobiotics genes and metabolic activities with level of total petroleum hydrocarbon in soil, sediment and estuary water in the Niger Delta Region of Nigeria. Asian J. Biotechnol. Genet. Eng. 1, 1–11 (2018).
Lee, E.-H., Lee, S. H. & Cho, K.-S. Bacterial diversity dynamics in a long-term petroleum-contaminated soil. J. Environ. Sci. Health A 46, 281–290 (2011).
Mukherjee, S. et al. Spatial patterns of microbial diversity and activity in an aged creosote-contaminated site. ISME J. 8, 2131–2142 (2014).
Yerulker, G. et al. Comparative assessment of soil microbial community in crude oil contaminated sites. Environ. Pollut. 328, 121578 (2023).
Ramirez, D., Shaw, L. J. & Collins, C. D. Ecotoxicity of oil sludges and residuals from their washing with surfactants: Soil dehydrogenase and ryegrass germination tests. Environ. Sci. Pollut. Res. 28, 13312–13322 (2021).
Guérin, J. & Buchanan, S. K. Protein import and export across the bacterial outer membrane. Curr. Opin. Struct. Biol. 69, 55–62 (2021).
Molan, K. & Žgur Bertok, D. Small prokaryotic DNA-binding proteins protect genome integrity throughout the life cycle. Int. J. Mol. Sci. 23, 4008 (2022).
Durak, A., Buyukguner, E. & Dogan, H. M. Determination of physical and chemical properties of the soils under different land managements. Asian J. Chem. 22, 6375–6386 (2010).
Babalola, E., Asad, M. H. & Bais, A. Soil surface texture classification using RGB images acquired under uncontrolled field conditions. IEEE Access 11, 67140 (2023).
Adekiya, A. O. Different organic manure sources and NPK fertilizer on soil chemical properties, growth, yield and quality of okra. Sci. Rep. 4, 208 (2020).
Oyem, H. H., Oyem, I. M. & Ezeweali, D. Temperature, pH, electrical conductivity, total dissolved solids and chemical oxygen demand of groundwater in Boji-BojiAgbor/Owa area and immediate suburbs. Res. J. Environ. Sci. 8, 444 (2014).
Mrayyan, B. & Battikhi, M. N. Biodegradation of total organic carbons (TOC) in Jordanian petroleum sludge. J. Hazard. Mater. 120, 127–134 (2005).
De Varennes, A., Torres, M. O., Cunha-Queda, C., Goss, M. J. & Carranca, C. Nitrogen conservation in soil and crop residues as affected by crop rotation and soil disturbance under Mediterranean conditions. Biol. Fertil. Soils 44, 49–58 (2007).
Minai-Tehrani, D. & Herfatmanesh, A. Biodegradation of aliphatic and aromatic fractions of heavy crude oil–contaminated soil: A pilot study. Bioremed. J. 11, 71–76 (2007).
Sari, G. L., Trihadiningrum, Y., Suci, F. C. & FashanahHadining, A. Identification of total petroleum hydrocarbon and heavy metals levels in crude oil contaminated soil at Wonocolo public mining. Environ. Asia 11, 109–117 (2018).
dos Reis, C. B. L. et al. First report of the production of a potent biosurfactant with α, β-trehalose by Fusarium fujikuroi under optimized conditions of submerged fermentation. Braz. J. Microbiol. 49, 185–192 (2018).
Caporaso, J. G. et al. Ultra-high-throughput microbial community analysis on the Illumina HiSeq and MiSeq platforms. ISME J. 6, 1621–1624 (2012).
Uyaguari-Diaz, M. I. et al. A comprehensive method for amplicon-based and metagenomic characterization of viruses, bacteria, and eukaryotes in freshwater samples. Microbiome 4, 1–19 (2016).
Gao, J. et al. Metagenome analysis of bacterial diversity in Tibetan kefir grains. Eur. Food Res. Technol. 236, 549–556 (2013).
Hemmat-Jou, M. H., Safari-Sinegani, A. A., Mirzaie-Asl, A. & Tahmourespour, A. Analysis of microbial communities in heavy metals-contaminated soils using the metagenomic approach. Ecotoxicology 27, 1281–1291 (2018).
Qiu, Z. et al. Characterization of microbial community structure and metabolic potential using Illumina MiSeq platform during the black garlic processing. Food Res. Int. 106, 428–438 (2018).
Tawfik, S. A., Azab, M. M., Ahmed, A. A. A. & Fayyad, D. M. Illumina MiSeq sequencing for preliminary analysis of microbiome causing primary endodontic infections in Egypt. Int. J. Microbiol. 2018, 1–15 (2018).
Bolger, A. M., Lohse, M. & Usadel, B. Trimmomatic: A flexible trimmer for Illumina sequence data. Bioinformatics 30, 2114–2120 (2014).
Bolyen, E. et al. Reproducible, interactive, scalable and extensible microbiome data science using QIIME 2. Nat. Biotechnol. 37, 852–857 (2019).
Callahan, B. J. et al. DADA2: High-resolution sample inference from Illumina amplicon data. Nat. Methods 13, 581–583 (2016).
Rognes, T., Flouri, T., Nichols, B., Quince, C. & Mahé, F. VSEARCH: A versatile open source tool for metagenomics. PeerJ 4, e2584 (2016).
Wang, Q., Garrity, G. M., Tiedje, J. M. & Cole, J. R. Naïve Bayesian classifier for rapid assignment of rRNA sequences into the new bacterial taxonomy. Appl. Environ. Microbiol. 73, 5261–5267 (2007).
Douglas, G. M. et al. PICRUSt2 for prediction of metagenome functions. Nat. Biotechnol. 38, 685–688 (2020).
Dhariwal, A. et al. MicrobiomeAnalyst: A web-based tool for comprehensive statistical, visual and meta-analysis of microbiome data. Nucleic Acids Res. 45, W180–W188 (2017).
Kandlikar, G. S. et al. ranacapa: An R package and Shiny web app to explore environmental DNA data with exploratory statistics and interactive visualizations. F1000 Res. 7, 1734 (2018).
Author information
Authors and Affiliations
Contributions
E.O.F. and T.M.: Development of original draft: E.O.F., R.N., and M.T.: Review and editing: Visualization: R.N. and E.O.F.; Supervision and funding acquisition: M.T. All authors agree to publish this manuscript with Scientific Reports after reading the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Fenibo, E.O., Nkuna, R. & Matambo, T. Impact of artisanal refining activities on bacterial diversity in a Niger Delta fallow land. Sci Rep 14, 3866 (2024). https://doi.org/10.1038/s41598-024-53147-4
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-024-53147-4
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.