Featured
-
-
News |
 Ferns communicate to decide their sexes
Ferns communicate to decide their sexes
Older generations release pheromones to balance the sex ratio in youngsters.
- Mark Zastrow
-
Letter |
 In vivo engineering of oncogenic chromosomal rearrangements with the CRISPR/Cas9 system
In vivo engineering of oncogenic chromosomal rearrangements with the CRISPR/Cas9 system
The CRISPR/Cas system has been used to induce the Eml4–Alk chromosomal inversion in mice, a characteristic chromosomal rearrangement seen in human non-small cell lung cancers; the mice developed lung cancer and responded to the ALK inhibitor crizotinib, which is used to treat lung cancer patients with the EML4–ALK rearrangement; this general strategy can be used to engineer other disease-associated chromosomal rearrangements in mice and potentially in other organisms.
- Danilo Maddalo
- , Eusebio Manchado
- & Andrea Ventura
-
Feature |
 Molecular biology: Genetic touch-ups
Molecular biology: Genetic touch-ups
Simplified techniques have made the field of gene editing much more accessible to non-specialists.
- Jeffrey M. Perkel
-
Outlook |
 Q&A: Brian Kobilka
Q&A: Brian Kobilka
Brian Kobilka shared the 2012 Nobel Prize in Chemistry with Robert Lefkowitz for their studies of G protein-coupled receptors. He is professor of molecular and cellular physiology at the Stanford University School of Medicine in California. Haya Jamal Azouz asks Kobilka what it takes to spend 30 years answering a single research question.
- Haya Jamal Azouz
-
Letter |
 Cohesin-dependent globules and heterochromatin shape 3D genome architecture in S. pombe
Cohesin-dependent globules and heterochromatin shape 3D genome architecture in S. pombe
Genome-wide chromatin conformation capture (Hi-C) is used to investigate three-dimensional genome organization in Schizosaccharomyces pombe; small domains of chromatin interact locally on chromosome arms to form globules, which depend on cohesin but not heterochromatin for formation, and heterochromatin at centromeres and telomeres provides crucial structural constraints to shape genome architecture.
- Takeshi Mizuguchi
- , Geoffrey Fudenberg
- & Shiv I. S. Grewal
-
Brief Communications Arising |
 POLRMT does not transcribe nuclear genes
POLRMT does not transcribe nuclear genes
- Inge Kühl
- , Christian Kukat
- & Nils-Göran Larsson
-
Letter |
 R-loops induce repressive chromatin marks over mammalian gene terminators
R-loops induce repressive chromatin marks over mammalian gene terminators
R-loops, which have been considered to be rare and potentially harmful transcriptional by-products, are now shown to be needed for antisense transcription and to induce repressive chromatin marks that reinforce pausing of transcription and thereby enhance its termination.
- Konstantina Skourti-Stathaki
- , Kinga Kamieniarz-Gdula
- & Nicholas J. Proudfoot
-
Letter |
 The complete structure of the large subunit of the mammalian mitochondrial ribosome
The complete structure of the large subunit of the mammalian mitochondrial ribosome
The structure of the 39S large mitoribosome subunit is solved by cryo-electron microscopy at an impressive 3.4 Å resolution, revealing the location of 50 ribosomal proteins, the peptidyl transferase centre, the tRNAs within this active site, and the nascent peptide chain within the exit tunnel.
- Basil J. Greber
- , Daniel Boehringer
- & Nenad Ban
-
Letter |
 Programmable RNA recognition and cleavage by CRISPR/Cas9
Programmable RNA recognition and cleavage by CRISPR/Cas9
In the presence of a short DNA oligonucleotide containing a protospacer adjacent motif, a guide-RNA-programmed Cas9 is able to specifically bind and/or cleave single-stranded RNA—this system can be used to isolate specific endogenous RNA transcripts from a cell lysate without any tag or modification.
- Mitchell R. O’Connell
- , Benjamin L. Oakes
- & Jennifer A. Doudna
-
Letter |
 An evolutionary arms race between KRAB zinc-finger genes ZNF91/93 and SVA/L1 retrotransposons
An evolutionary arms race between KRAB zinc-finger genes ZNF91/93 and SVA/L1 retrotransposons
The authors show that two primate-specific genes encoding KRAB domain containing zinc finger proteins, ZNF91 and ZNF93, have evolved during the last 25 million years to repress retrotransposon families that emerged during this time period; according to the new data KZNF gene expansion limits the activity of newly emerged retrotransposons, which subsequently mutate to evade repression.
- Frank M. J. Jacobs
- , David Greenberg
- & David Haussler
-
News & Views |
 Lariat lessons
Lariat lessons
The spliceosome enzyme complex removes intron sequences from RNA transcripts to form messenger RNA. The crystal structure of a lasso-shaped RNA suggests a mechanism for this splicing process. See Article p.193
- Robert T. Batey
-
Letter |
 Tyrosine phosphorylation of histone H2A by CK2 regulates transcriptional elongation
Tyrosine phosphorylation of histone H2A by CK2 regulates transcriptional elongation
A conserved tyrosine residue, Tyr 57, of histone H2A is phosphorylated by an unsuspected tyrosine kinase activity of casein kinase 2, influencing a series of histone marks associated with active transcription and regulating transcription elongation.
- Harihar Basnet
- , Xue B. Su
- & Michael G. Rosenfeld
-
Article |
 Crystal structure of a eukaryotic group II intron lariat
Crystal structure of a eukaryotic group II intron lariat
This study determines the structure of a branched lariat RNA, providing insights into rearrangement of the intron between the two steps of RNA splicing.
- Aaron R. Robart
- , Russell T. Chan
- & Navtej Toor
-
Letter |
 Regulation of RNA polymerase II activation by histone acetylation in single living cells
Regulation of RNA polymerase II activation by histone acetylation in single living cells
The interplay of histone acetylation and RNA polymerase II activity is investigated using fluorescence microscopy; acetylation of H3 at Lys 27 enhances the recruitment of a transcriptional activator and accelerates the transition of RNA polymerase II from initiation to elongation, thus indicating that histone acetylation has a causal effect on two distinct steps in transcription activation.
- Timothy J. Stasevich
- , Yoko Hayashi-Takanaka
- & Hiroshi Kimura
-
News & Views |
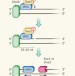 Making the cut
Making the cut
Analysis of the first step in repairing double-stranded-DNA breaks reveals that the Mre11 enzyme makes a DNA nick at a point separate from the break ends, creating an entry site for further processing by exonuclease enzymes. See Letter p.122
- Lorraine S. Symington
-
Letter |
 Sae2 promotes dsDNA endonuclease activity within Mre11–Rad50–Xrs2 to resect DNA breaks
Sae2 promotes dsDNA endonuclease activity within Mre11–Rad50–Xrs2 to resect DNA breaks
The MRX complex, required for double-strand break (DSB) repair by homologous recombination, has 3′ to 5′ exonuclease activity, but homologous recombination at a DSB uses a 3′-tailed molecule, which requires resection of the 5′ strand; here it is shown that in yeast, Sae2 nuclease promotes MRX to make an initial endonucleolytic cut on the 5′ strand that may allow MRX to digest the 5′ strand back to the end in a 3′ to 5′ fashion.
- Elda Cannavo
- & Petr Cejka
-
Letter |
 Histone H2A.Z subunit exchange controls consolidation of recent and remote memory
Histone H2A.Z subunit exchange controls consolidation of recent and remote memory
The authors identify a specific histone variant as a memory-suppressor that is initially reduced in expression within the hippocampus during memory formation; as a memory is consolidated to the cortex, reduced histone association with specific plasticity genes is observed, promoting stabilization of the memory.
- Iva B. Zovkic
- , Brynna S. Paulukaitis
- & J. David Sweatt
-
News & Views |
 Ribosome revelations
Ribosome revelations
Proteins are synthesized in cells by the ribosome apparatus. A report of 16 yeast ribosome structures, each bound by a different inhibitor, broadens our understanding of how drugs affect ribosome activity. See Article p.517
- Nelson B. Olivier
-
Article |
 Structural basis for the inhibition of the eukaryotic ribosome
Structural basis for the inhibition of the eukaryotic ribosome
Whereas previous structural investigation of ribosome inhibitors has been done using the prokaryotic ribosome, this work presents X-ray crystal structures of the yeast ribosome in complex with 16 inhibitors including eukaryotic-specific inhibitors; the inhibitors all bind the mRNA or tRNA binding sites, larger molecules appear to target specifically the first elongation cycle.
- Nicolas Garreau de Loubresse
- , Irina Prokhorova
- & Marat Yusupov
-
Letter |
 Structural basis for the assembly of the Sxl–Unr translation regulatory complex
Structural basis for the assembly of the Sxl–Unr translation regulatory complex
The crystal structure of the RNA binding domains of Sxl and Unr with msl2 RNA shows that interwoven interactions establish cooperative assembly of the ternary complex, highlighting how binding of relatively general RNA binding domains to RNA can result in a unique and specific protein–RNA architecture.
- Janosch Hennig
- , Cristina Militti
- & Michael Sattler
-
Letter |
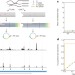 Pseudouridine profiling reveals regulated mRNA pseudouridylation in yeast and human cells
Pseudouridine profiling reveals regulated mRNA pseudouridylation in yeast and human cells
The modification of uridine to pseudouridine is widespread in transfer and ribosomal RNAs but not observed so far in a coding RNA; here a new technique is used to detect this modification on a genome-wide scale, leading to the identification of pseudouridylation in messenger RNAs as well as almost 100 new sites in non-coding RNAs.
- Thomas M. Carlile
- , Maria F. Rojas-Duran
- & Wendy V. Gilbert
-
Letter |
 Transcript-RNA-templated DNA recombination and repair
Transcript-RNA-templated DNA recombination and repair
Endogenous RNA transcripts are shown to mediate recombination with yeast chromosomal DNA; as the level of RNAs in the nucleus is quite high, these results may open up new understanding of the plasticity of repair and genome instability mechanisms.
- Havva Keskin
- , Ying Shen
- & Francesca Storici
-
Letter
| Open Access Comparative analysis of metazoan chromatin organization
Comparative analysis of metazoan chromatin organization
A large collection of new modENCODE and ENCODE genome-wide chromatin data sets from cell lines and developmental stages in worm, fly and human are analysed; this reveals many conserved features of chromatin organization among the three organisms, as well as notable differences in the composition and locations of repressive chromatin.
- Joshua W. K. Ho
- , Youngsook L. Jung
- & Peter J. Park
-
Letter |
 Transcriptional interference by antisense RNA is required for circadian clock function
Transcriptional interference by antisense RNA is required for circadian clock function
The transcriptions of frq sense and antisense RNAs are mutually inhibitory and form a double negative feedback loop required for robust and sustained circadian rhythmicity: antisense transcription inhibits sense expression by causing chromatin modifications and premature transcription termination.
- Zhihong Xue
- , Qiaohong Ye
- & Yi Liu
-
Letter |
 Contrasting roles of histone 3 lysine 27 demethylases in acute lymphoblastic leukaemia
Contrasting roles of histone 3 lysine 27 demethylases in acute lymphoblastic leukaemia
T-cell acute lymphoblastic leukaemia (T-ALL) is a haematological malignancy with a poor prognosis and no available targeted therapies; now two histone H3 lysine 27 demethylases, JMJD3 and UTX, are shown to have contrasting roles in human T-ALL cells and a mouse model of the disease, and a small molecule demethylase inhibitor is found to inhibit the growth of T-ALL cell lines, introducing a potential therapeutic avenue for acute leukaemia.
- Panagiotis Ntziachristos
- , Aristotelis Tsirigos
- & Iannis Aifantis
-
Letter |
 Crystal structure of the RNA-guided immune surveillance Cascade complex in Escherichia coli
Crystal structure of the RNA-guided immune surveillance Cascade complex in Escherichia coli
The CRISPR/Cas system is an RNA-guided bacterial protection system against foreign nucleic acids of bacterial and archaeal origin; here a high-resolution crystal structure of the CRIPSR RNA–Cas complex shows that the CRIPSR RNA plays an essential role not only in target recognition but also in complex assembly.
- Hongtu Zhao
- , Gang Sheng
- & Yanli Wang
-
Letter |
 A long noncoding RNA protects the heart from pathological hypertrophy
A long noncoding RNA protects the heart from pathological hypertrophy
Here, a long noncoding RNA, termed Mhrt, is identified in the loci of myosin heavy chain (Myh) genes in mice and shown to be capable of suppressing cardiomyopathy in the animals, as well as being repressed in diseased human hearts.
- Pei Han
- , Wei Li
- & Ching-Pin Chang
-
Letter |
 Noncoding RNA transcription targets AID to divergently transcribed loci in B cells
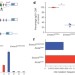
Noncoding RNA transcription targets AID to divergently transcribed loci in B cells
The 11-subunit RNA exosome is thought to regulate the mammalian noncoding transcriptome; here, a mouse model is generated in which the essential Exosc3 subunit of the RNA exosome in B cells is conditionally deleted, revealing a link between sites of genomic RNA exosome function and AID-mediated chromosomal translocations.
- Evangelos Pefanis
- , Jiguang Wang
- & Uttiya Basu
-
Letter |
 Promoter sequences direct cytoplasmic localization and translation of mRNAs during starvation in yeast
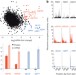
Promoter sequences direct cytoplasmic localization and translation of mRNAs during starvation in yeast
Transcription and translation are generally thought of as disconnected processes in eukaryotes; however, under starvation conditions in yeast, the promoter sequence influences not only messenger RNA levels but also several processes downstream of transcription, including the localization of mRNA within the cytoplasm and the translation rate of mRNA.
- Brian M. Zid
- & Erin K. O’Shea
-
Letter |
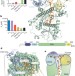 Mechanism of Dis3l2 substrate recognition in the Lin28–let-7 pathway
Mechanism of Dis3l2 substrate recognition in the Lin28–let-7 pathway
The structure of mouse Dis3l2 bound to an oligoU substrate shows a funnel-like substrate-binding site with the RNA being fed into the active site along a path that is distinct from that seen in the related catalytic subunit of the exosome — 12 uracils of the oligoU-tailed RNA are recognized in a complex network of interactions, suggesting the basis for target specificity.
- Christopher R. Faehnle
- , Jack Walleshauser
- & Leemor Joshua-Tor
-
Letter |
 Required enhancer–matrin-3 network interactions for a homeodomain transcription program
Required enhancer–matrin-3 network interactions for a homeodomain transcription program
The POU homeodomain transcription factor Pit1 is required for pituitary development; here Pit1-occupied enhancers are shown to interact with the nuclear architecture components matrin-3 and Satb1, and this association is required for activation of Pit1-regulated enhancers and coding target genes.
- Dorota Skowronska-Krawczyk
- , Qi Ma
- & Michael G. Rosenfeld
-
Letter |
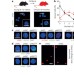 Replication stress is a potent driver of functional decline in ageing haematopoietic stem cells
Replication stress is a potent driver of functional decline in ageing haematopoietic stem cells
Haematopoietic stem cell (HSC) function is known to degrade with age; here, replication stress is shown to be a potent driver of the functional decline of HSCs during physiological ageing in mice due to decreased expression of mini-chromosome maintenance helicase components and reduced activity of the DNA replication machinery.
- Johanna Flach
- , Sietske T. Bakker
- & Emmanuelle Passegué
-
Letter |
 Structural basis of PAM-dependent target DNA recognition by the Cas9 endonuclease
Structural basis of PAM-dependent target DNA recognition by the Cas9 endonuclease
Crystal structure of the RNA-guided endonuclease Cas9 bound to a guide RNA and a target DNA duplex reveals how base-specific recognition of a short motif known as PAM in the DNA target results in localized strand separation in the DNA immediately upstream of the PAM, allowing the target DNA strand to hybridize to the guide RNA.
- Carolin Anders
- , Ole Niewoehner
- & Martin Jinek
-
Brief Communications Arising |
 Universality of core promoter elements?
Universality of core promoter elements?
- Matthias Siebert
- & Johannes Söding
-
Letter |
 The DNA methylation landscape of human early embryos
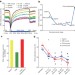
The DNA methylation landscape of human early embryos
Base-resolution maps of DNA methylation in human gametes and early embryos offer novel insights into human methylation dynamics and the functional relationship between DNA methylation and gene expression.
- Hongshan Guo
- , Ping Zhu
- & Jie Qiao
-
Letter |
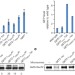 The Get1/2 transmembrane complex is an endoplasmic-reticulum membrane protein insertase
The Get1/2 transmembrane complex is an endoplasmic-reticulum membrane protein insertase
The receptor for the cytoplasmic factor that targets tail-anchored proteins to the endoplasmic reticulum is an enzyme that enables a facilitated insertion path into the lipid bilayer.
- Fei Wang
- , Charlene Chan
- & Vladimir Denic
-
Comment |
 History: Fifty years of EMBO
History: Fifty years of EMBO
Georgina Ferry reflects on the evolution of the European Molecular Biology Organization, founded to help Europe to compete with the United States.
- Georgina Ferry
-
News & Views |
 Fine-tuned amplification in cells
Fine-tuned amplification in cells
The transcription factor Myc has been posited to cause a cell-wide increase in gene expression. But two studies show that Myc, when modulated by other transcription factors, can amplify select targets. See Letters p.483 and p.488
- Chi V. Dang
-
Letter |
 Selective transcriptional regulation by Myc in cellular growth control and lymphomagenesis
Selective transcriptional regulation by Myc in cellular growth control and lymphomagenesis
Global transcriptional and epigenomic analyses in diverse cell types reveal that the primary action of Myc is to up- and downregulate transcription of distinct groups of genes, rather than to amplify transcription of all active genes; general RNA amplification, when observed, is better explained as an indirect consequence of Myc’s action on cellular physiology.
- Arianna Sabò
- , Theresia R. Kress
- & Bruno Amati
-
Article |
 Ribosomal frameshifting in the CCR5 mRNA is regulated by miRNAs and the NMD pathway
Ribosomal frameshifting in the CCR5 mRNA is regulated by miRNAs and the NMD pathway
Programmed −1 ribosomal frameshifting (−1 PRF) is a process by which a signal in a messenger RNA causes a translating ribosome to shift by one nucleotide, thus changing the reading frame; here −1 PRF in the mRNA for the co-receptor for HIV-1, CCR5, is stimulated by two microRNAs and leads to degradation of the transcript by nonsense-mediated decay and at least one other decay pathway.
- Ashton Trey Belew
- , Arturas Meskauskas
- & Jonathan D. Dinman
-
Letter |
 Activation and repression by oncogenic MYC shape tumour-specific gene expression profiles
Activation and repression by oncogenic MYC shape tumour-specific gene expression profiles
Inducing changes in the levels of the MYC oncoprotein is shown to activate and repress specific sets of target genes that are characteristic of tumour cells, providing an insight into the mechanism by which MYC can stimulate tumorigenesis in contrast to its physiological role.
- Susanne Walz
- , Francesca Lorenzin
- & Martin Eilers
-
Letter |
 DENR–MCT-1 promotes translation re-initiation downstream of uORFs to control tissue growth
DENR–MCT-1 promotes translation re-initiation downstream of uORFs to control tissue growth
This study identifies the DENR–MCT-1 complex as the first factors in animals specific for translation re-initiation downstream of upstream Open Reading Frames (uORFs).
- Sibylle Schleich
- , Katrin Strassburger
- & Aurelio A. Teleman
-
Research Highlights |
Halting inheritance of genetic disease
-
Letter |
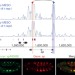 Enhancer loops appear stable during development and are associated with paused polymerase
Enhancer loops appear stable during development and are associated with paused polymerase
A high-resolution map of enhancer three-dimensional contacts during Drosophila embryogenesis shows that although local regulatory interactions are frequent, long-range interactions are also very common; unexpectedly, most interactions appear unchanged between tissues and across development and are formed prior to gene expression, indicating that transcription initiates from preformed enhancer–promoter loops, which are associated with paused polymerase.
- Yad Ghavi-Helm
- , Felix A. Klein
- & Eileen E. M. Furlong
-
News |
 Elite labs hire more men than women
Elite labs hire more men than women
Women account for half of qualified candidates but are outnumbered in labs led by top male academics.
- Elizabeth Gibney
-
Letter |
 Histone H4 tail mediates allosteric regulation of nucleosome remodelling by linker DNA
Histone H4 tail mediates allosteric regulation of nucleosome remodelling by linker DNA
A nucleosome-spacing mechanism for human ATP-dependent chromatin assembly and remodelling factor (ACF).
- William L. Hwang
- , Sebastian Deindl
- & Xiaowei Zhuang
-
Letter |
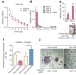 RETRACTED ARTICLE: miR-34a blocks osteoporosis and bone metastasis by inhibiting osteoclastogenesis and Tgif2
RETRACTED ARTICLE: miR-34a blocks osteoporosis and bone metastasis by inhibiting osteoclastogenesis and Tgif2
A microRNA, miR-34a, is a novel and critical suppressor of osteoclastogenesis, bone resorption and the bone metastatic niche.
- Jing Y. Krzeszinski
- , Wei Wei
- & Yihong Wan
-
Letter |
 PVT1 dependence in cancer with MYC copy-number increase
PVT1 dependence in cancer with MYC copy-number increase
Pvt1 overexpression in mice contributes to high Myc levels due to 8q24.21 gain and to MYC-driven tumorigenesis.
- Yuen-Yi Tseng
- , Branden S. Moriarity
- & Anindya Bagchi
-
Research Highlights |
Genome editing of stem cells
Browse broader subjects
Browse narrower subjects
- Cell division
- Chromatin
- Chromosomes
- CRISPR-Cas systems
- DNA damage and repair
- DNA metabolism
- DNA recombination
- DNA replication
- Epigenetics
- Non-coding RNAs
- Nuclear organization
- Post-translational modifications
- Protein folding
- Proteolysis
- Proteomics
- Riboswitches
- Ribozymes
- RNA metabolism
- RNAi
- Single-molecule biophysics
- Transcription
- Transcriptomics
- Translation
- Transposition

 Cancer cells can ‘infect’ normal neighbours
Cancer cells can ‘infect’ normal neighbours