Review Article |
Featured
-
-
Review Article |
 Engineering 3D genome organization
Engineering 3D genome organization
There is a rapidly growing appreciation of the complexities of 3D genome organization, as well as associations with gene expression and wider cellular and organismal phenotypes, including diseases. In this Review, the authors describe diverse experimental methods for manipulating 3D genome organization — from fine-scale control of DNA contacts to large-scale nuclear repositioning — which are facilitating detailed testing of the biological functions of 3D genome organization.
- Haifeng Wang
- , Mengting Han
- & Lei S. Qi
-
Research Highlight |
 Linking newly occurring mutations to autism
Linking newly occurring mutations to autism
Three new studies identify different types of de novo and somatic mutations associated with autism spectrum disorder (ASD), with potential insights into underlying molecular mechanisms.
- Darren J. Burgess
-
In Brief |
Addressing admixture with Tractor
In this paper in Nature Genetics, Atkinson et al. describe Tractor, a statistical framework and software package that enables admixed populations to be included in large-scale genomics studies.
- Dorothy Clyde
-
In Brief |
Cascading CRISPR–Cas9 genome edits
In this study in Molecular Cell, Clarke et al. describe a system that enables multiple Cas9-mediated genome edits to be introduced into cells in a defined, sequential order.
- Dorothy Clyde
-
Research Highlight |
 A new view of genome organization
A new view of genome organization
A report in Science describes in situ genome sequencing (IGS), a method that enables genomes to be simultaneously sequenced and imaged in intact samples, including early mouse embryos. IGS will facilitate insight into the relationship between genome sequence and organization.
- Dorothy Clyde
-
Review Article |
 The roles of microRNAs in mouse development
The roles of microRNAs in mouse development
MicroRNAs (miRNA) exert essential functions in mammalian development and physiology. The authors review recent insights from the phenotypic analysis of miRNA knockouts in mice that emphasize roles for these non-coding RNAs at different developmental stages and in adults, and illustrate the importance of functional miRNA targets, miRNA dosage, miRNA interactions and cellular context.
- Brian DeVeale
- , Jennifer Swindlehurst-Chan
- & Robert Blelloch
-
Research Highlight |
 Fixing an ageing mutation
Fixing an ageing mutation
A study in Nature reports that adenine base editors can correct the mutation that causes Hutchinson–Gilford progeria syndrome in a mouse model of this disease, extending lifespan.
- Katharine H. Wrighton
-
Research Highlight |
 Complex targeted sequencing in real time
Complex targeted sequencing in real time
Two new papers in Nature Biotechnology report methods for targeted sequencing of complex DNA samples, achieved in real time during nanopore sequencing runs.
- Darren J. Burgess
-
In Brief |
Spaceflight causes mitochondrial stress
An article in Cell describes a multi-omic analysis of health risks from spaceflights that implicates mitochondrial stress and dysregulation as key drivers.
- Dorothy Clyde
-
In Brief |
Uncovering distinct roles for H3K9me3
A study in Genome Biology uses EpiGo-KRAB to analyse the roles of H3K9me3 in genome organization and transcriptional repression and reveals the two functions may be distinct.
- Dorothy Clyde
-
In Brief |
Mapping metastasis
Jin et al. describe a barcoding approach for analysing metastasis, which they used to generate an organ-specific metastasis map for 500 cancer cell lines.
- Dorothy Clyde
-
In Brief |
SARS-CoV-2 detection goes mobile
Fozouni et al. describe a CRISPR–Cas13a-based approach for rapid, accurate and portable detection of SARS-CoV-2 in infected individuals.
- Dorothy Clyde
-
Review Article |
 Reprogramming the genetic code
Reprogramming the genetic code
The ability to reprogramme cellular translation and genomes to produce non-canonical biopolymers has wide-ranging applications, including in therapeutics, but has yet to be fully realized. In this Review, de la Torre and Chin discuss recent advances towards achieving this goal.
- Daniel de la Torre
- & Jason W. Chin
-
In Brief |
CRISPR–Cas13 targets circRNAs
The CRISPR–Cas13 system can be used to knock down circular RNAs (circRNAs) without any impact on related mRNAs, reports a study in Nature Methods.
- Linda Koch
-
In Brief |
Probing the RNA–protein interface
Corley et al. report fSHAPE, a simplified strategy for in vivo transcriptome-wide footprinting with SHAPE structure-probing techniques.
- Linda Koch
-
Research Highlight |
 Host genetics of coronavirus infection
Host genetics of coronavirus infection
Two new reports in Cell use genome-wide CRISPR screens to uncover host determinants of coronavirus infection, identifying potential leads for antiviral therapeutics.
- Darren J. Burgess
-
Research Highlight |
 SHARE-seq reveals chromatin potential
SHARE-seq reveals chromatin potential
SHARE-seq, a new high-throughput, high-resolution multi-omics method described in Cell, measures chromatin accessibility and gene expression in the same cell and enables future potential gene expression (and therefore lineage choices) to be inferred from chromatin profiles.
- Dorothy Clyde
-
Perspective |
 The epitranscriptome beyond m6A
The epitranscriptome beyond m6A
This Perspective reviews efforts to map six different RNA modifications — pseudouridine (Ψ), 5-methylcytidine (m5C), N 1-methyladenosine (m1A), N 4-acetylcytidine (ac4C), ribose methylations (Nm) and N 7-methylguanosine (m7G) — and how they differ from N 6-methyladenosine (m6A). The authors discuss the technical and analytical challenges of characterizing the epitranscriptome and provide their own conclusions on the abundance and distribution of these modifications.
- David Wiener
- & Schraga Schwartz
-
Review Article |
 Deciphering cell–cell interactions and communication from gene expression
Deciphering cell–cell interactions and communication from gene expression
Cell–cell interactions and communication can be inferred from RNA sequencing data of, for example, ligand–receptor pairs. The authors review insights gained and the methods and tools used in studies of cell–cell interactions based on transcriptomic data.
- Erick Armingol
- , Adam Officer
- & Nathan E. Lewis
-
Research Highlight |
 Nucleic-acid sequencing of proteomes
Nucleic-acid sequencing of proteomes
A recent study re-casts proteomic analyses as a DNA sequencing problem; by fusing in vivo-expressed proteins to their encoding mRNA, molecular interactions can be identified and quantified through high-throughput nucleic-acid sequencing.
- Linda Koch
-
Research Highlight |
 Reaching completion for GTEx
Reaching completion for GTEx
The GTEx consortium reports results from its third and final phase in several new papers. They provide unprecedented detail of human gene expression regulation across tissues.
- Darren J. Burgess
-
Comment |
African ancient DNA research requires robust ethics and permission protocols
In Africa, there is a disparity in ethics and permission requirements for molecular research on samples from living people versus ancient DNA. At the precipice of the archaeogenomics revolution, heritage agencies require updated policies and procedures for genetic and genomic research on African ancient DNA.
- Victoria E. Gibbon
-
Research Highlight |
 Tracing chromatin architecture
Tracing chromatin architecture
A study in Cell presents a new approach that increases resolution and throughput compared with existing imaging methods and provides insights into the relationship between transcription and the 3D genome.
- Linda Koch
-
Review Article |
 Genetics meets proteomics: perspectives for large population-based studies
Genetics meets proteomics: perspectives for large population-based studies
In this Review, Suhre, McCarthy and Schwenk describe how combining genetics with plasma proteomics is providing notable insights into human disease. As changes in the circulating proteome are often an intermediate molecular readout between a genetic variant and its organismal effect, proteomics can enable a deeper understanding of disease mechanisms, clinical biomarkers and therapeutic opportunities.
- Karsten Suhre
- , Mark I. McCarthy
- & Jochen M. Schwenk
-
Viewpoint |
The road ahead in genetics and genomics
To celebrate the first 20 years of Nature Reviews Genetics, we asked 12 leading scientists to reflect on the key challenges and opportunities faced by the field of genetics and genomics.
- Amy L. McGuire
- , Stacey Gabriel
- & Jin-Soo Kim
-
Research Highlight |
 Testing the developing epigenome
Testing the developing epigenome
A recent study combines CRISPR-based perturbation with single-cell RNA sequencing to characterize the roles of epigenome regulator proteins in controlling cell fate and identity during embryonic development.
- Darren J. Burgess
-
Research Highlight |
 Accessible disease insights
Accessible disease insights
A new study in Science uses chromatin accessibility profiles to reveal gene regulatory alterations associated with genetic variants in neuropsychiatric disease.
- Darren J. Burgess
-
Review Article |
 Integrating genetic and non-genetic determinants of cancer evolution by single-cell multi-omics
Integrating genetic and non-genetic determinants of cancer evolution by single-cell multi-omics
Both genetic and non-genetic factors underlie the intratumoural heterogeneity that fuels cancer evolution. This Review discusses the application of single-cell multi-omics technologies to the study of cancer evolution, which capture and integrate the different layers of heritable information and reveal their complex interplay.
- Anna S. Nam
- , Ronan Chaligne
- & Dan A. Landau
-
Comment |
The Human Genome Project changed everything
Thirty years on from the launch of the Human Genome Project, Richard Gibbs reflects on the promisesthat this voyage of discovery bore. Its success should be measured by how this project transformed the rules of research, the way of practising biological discovery and the ubiquitous digitization of biological science.
- Richard A. Gibbs
-
Research Highlight |
 A long read of the human genome
A long read of the human genome
A study in Nature describes the assembly of a human genome with greater continuity than the current reference genome, as well as the assembly of a complete human X chromosome. These assemblies were achieved by combining data generated by different long-read sequencing technologies.
- Katharine H. Wrighton
-
Research Highlight |
 An odyssey to Oceania
An odyssey to Oceania
A study in Nature analysing genome-wide variation in individuals from islands across Polynesia reports evidence of admixture with Native Americans related to Indigenous inhabitants of northern South America.
- Linda Koch
-
Research Highlight |
 A zipcode for transcriptomes
A zipcode for transcriptomes
A new study in Nature Methods presents the ‘ZipSeq’ spatial transcriptomics approach, whereby patterned illumination is used to print barcodes onto chosen tissue regions.
- Darren J. Burgess
-
Research Highlight |
 Resolving the roles of structural variants
Resolving the roles of structural variants
Structural variants have proved difficult to characterize using traditional sequencing approaches. In two new studies in Cell, the authors demonstrate the use of pan-genome approaches to identify and explore the impact of structural variants in crop genomes and reveal variants linked to specific agronomic traits.
- Joseph Willson
-
Research Highlight |
 Multitasking for base editors
Multitasking for base editors
Three new studies in Nature Biotechnology combine the adenine and cytosine deaminase activities of single base editors to generate dual base editor systems for combinatorial editing in human cells.
- Darren J. Burgess
-
Review Article |
 Discovering and validating cancer genetic dependencies: approaches and pitfalls
Discovering and validating cancer genetic dependencies: approaches and pitfalls
The development of successful anticancer therapies relies on identifying drug targets that are genuine cancer-specific vulnerabilities. In this article, Lin and Sheltzer discuss how the different genetic and pharmacological methods for identifying and characterizing cancer dependencies each have important strengths and limitations. Responsible and orthogonal use of these methods holds promise for maximizing the ability of preclinical research to translate into clinical benefit.
- Ann Lin
- & Jason M. Sheltzer
-
Review Article |
 A decade of advances in transposon-insertion sequencing
A decade of advances in transposon-insertion sequencing
In this Review, several experts discuss progress in the decade since the development of transposon-based approaches for bacterial genetic screens. They describe how advances in both experimental technologies and analytical strategies are resulting in insights into diverse biological processes.
- Amy K. Cain
- , Lars Barquist
- & Tim van Opijnen
-
Research Highlight |
 Exploring human genomic diversity with gnomAD
Exploring human genomic diversity with gnomAD
A collection of seven articles from the gnomAD consortium, published in Nature, Nature Medicine and Nature Communications, showcases analyses of global human genetic variation in coding and non-coding genomic regions across this data set.
- Linda Koch
-
Review Article |
 From molecules to populations: appreciating and estimating recombination rate variation
From molecules to populations: appreciating and estimating recombination rate variation
Genetic recombination is a fundamental biological process generating genetic variation by shuffling combinations of alleles. In this Review, Peñalba and Wolf focus on how sequencing-based approaches are providing diverse insights into recombination rate variation across levels of biological organization and timescales, from individual gametes of single individuals to populations through evolutionary history.
- Joshua V. Peñalba
- & Jochen B. W. Wolf
-
Research Highlight |
 A mouse with history
A mouse with history
A new study in Cell describes the CRISPR array repair lineage tracing (CARLIN) engineered mouse line that genomically encodes all the components for CRISPR-based lineage tracking at single-cell resolution.
- Darren J. Burgess
-
Research Highlight |
 A platform for RNA virus cloning
A platform for RNA virus cloning
A study reports on the suitability of the yeast Saccharomyces cerevisiae as a platform for the assembly and maintenance of diverse RNA virus genomes, including SARS-CoV-2.
- Linda Koch
-
Research Highlight |
 Chromosome structure at micro-scale
Chromosome structure at micro-scale
Two studies in Molecular Cell report fine-scale structural profiles of mammalian genomes using Micro-C, indicating that fine chromosomal structure is regulated by diverse transcription-related features.
- Darren J. Burgess
-
Research Highlight |
 Early chromatin dynamics
Early chromatin dynamics
A study in Nature provides new insights into chromatin dynamics and allele-specific gene expression regulation during early mouse embryogenesis.
- Linda Koch
-
Review Article |
 Animal domestication in the era of ancient genomics
Animal domestication in the era of ancient genomics
Improvements in DNA extraction methods and sequencing technologies have led to the successful sequencing of numerous whole ancient genomes. In this Review, the authors provide an overview of how ancient DNA has informed our understanding of the domestication of various animal species, including dogs, pigs, cattle, goats and chickens.
- Laurent A. F. Frantz
- , Daniel G. Bradley
- & Ludovic Orlando
-
Perspective |
 Rights, interests and expectations: Indigenous perspectives on unrestricted access to genomic data
Rights, interests and expectations: Indigenous perspectives on unrestricted access to genomic data
In this Perspective article, the authors discuss how Indigenous Peoples' desires for greater involvement and oversight when participating in genomic research projects can be balanced against calls for unrestricted data access. They provide practical recommendations for the handling and sharing of Indigenous genomic data, with the aim of achieving mutual benefit for the research community and participating Indigenous communities.
- Maui Hudson
- , Nanibaa’ A. Garrison
- & Stephanie Russo Carroll
-
Research Highlight |
 CRISPR screens beyond Cas9
CRISPR screens beyond Cas9
Two new studies in Nature Biotechnology demonstrate the feasibility of functional genomics systems beyond Cas9: a combinatorial DNA editing system involving both Cas9 and Cas12a, and an RNA-targeted system based on Cas13.
- Darren J. Burgess
-
Review Article |
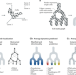 Lineage tracing meets single-cell omics: opportunities and challenges
Lineage tracing meets single-cell omics: opportunities and challenges
Understanding developmental trajectories has recently been enabled by progress in modern lineage-tracing methods that combine genetic lineage analysis with omics-based characterization of cell states (particularly transcriptomes). In this Review, Wagner and Klein discuss the conceptual underpinnings, experimental strategies and analytical considerations of these approaches, as well as the biological insights gained.
- Daniel E. Wagner
- & Allon M. Klein
-
Review Article |
 Electronic health records and polygenic risk scores for predicting disease risk
Electronic health records and polygenic risk scores for predicting disease risk
Electronic health records (EHRs) linked to biobanks provide new opportunities for developing and applying polygenic risk scores in the clinic. The authors review the opportunities and challenges that arise when using EHR data for the systematic evaluation of patient disease susceptibilities.
- Ruowang Li
- , Yong Chen
- & Jason H. Moore
-
Research Highlight |
 Repressors of healthy ageing
Repressors of healthy ageing
RNA interference screening in Caenorhabditis elegans has identified two repressive epigenetic regulators of age-related behavioural performance that are conserved in mammals.
- Linda Koch
-
Research Highlight |
 Transcriptomics of developmental fate
Transcriptomics of developmental fate
Two new studies in Science combine single-cell RNA sequencing with either lineage tracing or a computational framework to link transcriptomes to future developmental trajectories.
- Darren J. Burgess
Browse broader subjects
Browse narrower subjects
- Analytical biochemistry
- Behavioural methods
- Bioinformatics
- Biological models
- Biophysical methods
- Cytological techniques
- Electrophysiology
- Epigenetics analysis
- Experimental organisms
- Gene delivery
- Gene expression analysis
- Genetic engineering
- Genetic techniques
- Genomic analysis
- High-throughput screening
- Imaging
- Immunological techniques
- Isolation, separation and purification
- Lab-on-a-chip
- Mass spectrometry
- Metabolomics
- Microbiology techniques
- Microscopy
- Molecular engineering
- Nanobiotechnology
- Optogenetics
- Proteomic analysis
- Sensors and probes
- Sequencing
- Software
- Optical spectroscopy
- Structure determination

 Cross-species RNA-seq for deciphering host–microbe interactions
Cross-species RNA-seq for deciphering host–microbe interactions