Research Highlight |
Featured
-
-
Journal Club |
Decoding human genetic variation using a synthetic paradigm
Aashiq Kachroo highlights a recent paper by van Loggerenberg et al. that demonstrates the experimental power of ‘humanized yeast’ to gain insight into the genetic variants underlying disease.
- Aashiq H. Kachroo
-
Research Highlight |
Predicting the effects of multigene perturbations
Roohani et al. have developed a deep learning tool called GEARS that predicts the effects of multigene perturbations.
- Henry Ertl
-
Research Highlight |
Assigning phenotypes to essential human genes
A microscopy-based pooled CRISPR screening approach described in Cell enables the cellular functions of thousands of genes to be assessed at remarkable phenotypic depth.
- Dorothy Clyde
-
Journal Club |
Tightening the (neural) net for protein structure prediction
In this Journal Club article, Yana Bromberg discusses an early application of machine learning for protein structure prediction — a paper that shaped her career. It illustrates the value of ensuring that machine learning approaches are rooted in known biological principles.
- Yana Bromberg
-
Review Article |
 From systems to structure — using genetic data to model protein structures
From systems to structure — using genetic data to model protein structures
Large-scale genetic datasets and deep learning approaches are being used to model the structures of proteins or protein complexes. This Review describes approaches based on coevolution, deep mutational scanning and genome-scale genetic or chemical–genetic interaction mapping and their application and integration to inform structural modelling.
- Hannes Braberg
- , Ignacia Echeverria
- & Nevan J. Krogan
-
Review Article |
 Engineering synthetic RNA devices for cell control
Engineering synthetic RNA devices for cell control
Synthetic RNA devices integrate sensing, processing and actuation of signals into defined, programmable functions to control cell behaviour. This Review discusses the emerging applications of RNA devices in biomedical research and biomanufacturing, as well as progress in creating new ligand sensors and new mechanisms of action with engineered RNAs.
- Peter B. Dykstra
- , Matias Kaplan
- & Christina D. Smolke
-
Research Highlight |
 Host genetics of coronavirus infection
Host genetics of coronavirus infection
Two new reports in Cell use genome-wide CRISPR screens to uncover host determinants of coronavirus infection, identifying potential leads for antiviral therapeutics.
- Darren J. Burgess
-
Review Article |
 Discovering and validating cancer genetic dependencies: approaches and pitfalls
Discovering and validating cancer genetic dependencies: approaches and pitfalls
The development of successful anticancer therapies relies on identifying drug targets that are genuine cancer-specific vulnerabilities. In this article, Lin and Sheltzer discuss how the different genetic and pharmacological methods for identifying and characterizing cancer dependencies each have important strengths and limitations. Responsible and orthogonal use of these methods holds promise for maximizing the ability of preclinical research to translate into clinical benefit.
- Ann Lin
- & Jason M. Sheltzer
-
Review Article |
 A decade of advances in transposon-insertion sequencing
A decade of advances in transposon-insertion sequencing
In this Review, several experts discuss progress in the decade since the development of transposon-based approaches for bacterial genetic screens. They describe how advances in both experimental technologies and analytical strategies are resulting in insights into diverse biological processes.
- Amy K. Cain
- , Lars Barquist
- & Tim van Opijnen
-
Research Highlight |
 CRISPR screens beyond Cas9
CRISPR screens beyond Cas9
Two new studies in Nature Biotechnology demonstrate the feasibility of functional genomics systems beyond Cas9: a combinatorial DNA editing system involving both Cas9 and Cas12a, and an RNA-targeted system based on Cas13.
- Darren J. Burgess
-
Research Highlight |
 Repressors of healthy ageing
Repressors of healthy ageing
RNA interference screening in Caenorhabditis elegans has identified two repressive epigenetic regulators of age-related behavioural performance that are conserved in mammals.
- Linda Koch
-
Review Article |
 Towards a comprehensive catalogue of validated and target-linked human enhancers
Towards a comprehensive catalogue of validated and target-linked human enhancers
In this Review, Gasperini, Tome and Shendure discuss the evolving definitions of transcriptional enhancers, as well as diverse modern experimental tools to identify them. The authors describe how these diverse mindsets and methods provide differing but complementary insights into enhancers, each with notable strengths and caveats. They discuss how such views and approaches might be combined in a comprehensive catalogue of functional enhancers.
- Molly Gasperini
- , Jacob M. Tome
- & Jay Shendure
-
Research Highlight |
 CRISPR screens come into sight
CRISPR screens come into sight
A new study in Cell reports a mammalian genetic screening strategy that combines CRISPR libraries with in situ sequencing to read out both complex cellular phenotypes and genetic perturbations using microscopy.
- Darren J. Burgess
-
Research Highlight |
 Genotype–phenotype mapping in another dimension
Genotype–phenotype mapping in another dimension
A study in Science shows that it is now possible to create high-resolution, non-linear maps of mammalian genetic interactions.
- Linda Koch
-
Review Article |
 The biogenesis, biology and characterization of circular RNAs
The biogenesis, biology and characterization of circular RNAs
In eukaryotes, circular RNAs (circRNAs) carry out important biological roles by acting as microRNA or protein sponges, regulating protein function or through cap-independent translation. New technologies for identifying and characterizing circRNAs will increase our knowledge of their biogenesis and function in health and disease.
- Lasse S. Kristensen
- , Maria S. Andersen
- & Jørgen Kjems
-
Review Article |
 High-throughput mouse phenomics for characterizing mammalian gene function
High-throughput mouse phenomics for characterizing mammalian gene function
Although the field of functional genomics is increasingly adopting genome-scale approaches, a comprehensive understanding of gene functions requires the parallel development of deep phenotyping platforms. This Review discusses strategies for broad-based mouse phenomics, applied both to gene knockout collections and to diverse strains harbouring natural genetic variation. The authors discuss technical challenges, analysis pipelines and insights into human disease genetics.
- Steve D. M. Brown
- , Chris C. Holmes
- & Sara Wells
-
-
Review Article |
 Am I ready for CRISPR? A user's guide to genetic screens
Am I ready for CRISPR? A user's guide to genetic screens
The rapid development of CRISPR-based gene manipulation has enabled various approaches for high-throughput functional genomics. This Review guides users through the practicalities of CRISPR-based functional genomics screens, including study design options, best-practice approaches, pitfalls to avoid and data analysis strategies.
- John G. Doench
-
-
Review Article |
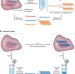 Scaling by shrinking: empowering single-cell 'omics' with microfluidic devices
Scaling by shrinking: empowering single-cell 'omics' with microfluidic devices
As the genetic and phenotypic heterogeneities among cells become more appreciated, there is increasing demand for technologies that facilitate high-throughput and high-efficiency single-cell 'omic' analyses in miniaturized and automated formats. This Review discusses the diverse microfluidic methodologies — with a primary focus on valve-, droplet- and nanowell-based platforms — for characterization of the genomes, epigenomes, transcriptomes and proteomes of single cells, and addresses technical considerations and future opportunities.
- Sanjay M. Prakadan
- , Alex K. Shalek
- & David A. Weitz
-
-
-
-
-
Review Article |
 Loss-of-function genetic tools for animal models: cross-species and cross-platform differences
Loss-of-function genetic tools for animal models: cross-species and cross-platform differences
Loss-of-function (LOF) approaches are powerful experimental tools for characterizing gene functions. However, emerging discrepancies when genes are investigated using different tools or organisms has triggered debate about how such LOF results should be biologically interpreted. In this Review, experts from varied fields discuss how understanding the underlying features of each LOF approach can provide explanations for different experimental outcomes and can guide their optimal and reliable application.
- Benjamin E. Housden
- , Matthias Muhar
- & Norbert Perrimon
-
-
-
-
Review Article |
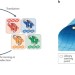 Methods for the directed evolution of proteins
Methods for the directed evolution of proteins
Directed evolution uses laboratory-based evolution to enhance the properties of biomolecules, primarily to generate proteins with optimized or novel activities. This Review discusses the diverse range of technologies for the directed evolution of proteins, particularly methods for generating diversity in the gene library and approaches for screening and selecting for variants with desired properties. The relative strengths and limitations of these approaches are highlighted to guide readers to appropriate strategies.
- Michael S. Packer
- & David R. Liu
-
Review Article |
 High-throughput functional genomics using CRISPR–Cas9
High-throughput functional genomics using CRISPR–Cas9
CRISPR–Cas9 has been adopted as a powerful genome-editing technology in various species. By generating libraries of thousands of guide RNAs — which direct the Cas9 nuclease to chosen genomic loci — high-throughput genetic perturbations are now possible. This Review discusses the latest applications of CRISPR–Cas9 in mammalian functional genomics screens. It covers related genome-scale applications of Cas9 for either gene knockout or transcriptional modulation, and provides comparisons with complementary RNA interference (RNAi)-based approaches.
- Ophir Shalem
- , Neville E. Sanjana
- & Feng Zhang
-
-
-
-
Review Article |
 Investigating human disease using stem cell models
Investigating human disease using stem cell models
Understanding disease pathogenesis and developing potential therapies require accurate and genetically tractable models. This Review discusses how human stem cells — including embryonic stem cells, adult stem cells and induced pluripotent stem cells — can provide informative models of diverse human diseases. Such methods can also be extended through gene editing, co-culture or infectious agent approaches.
- Jared L. Sterneckert
- , Peter Reinhardt
- & Hans R. Schöler
-
Review Article |
 In pursuit of design principles of regulatory sequences
In pursuit of design principles of regulatory sequences
Gene-regulatory DNA elements control complex spatiotemporal patterns of gene expression, and alterations to these sequences are commonly associated with inter-individual phenotypic variation and human disease. This Review discusses our latest understanding of how different layers of information in these sequences control the binding of regulators and influence gene expression outcomes.
- Michal Levo
- & Eran Segal
-
-

 The regulatory landscape of chromatin accessibility
The regulatory landscape of chromatin accessibility A drop in an ocean of gene variants
A drop in an ocean of gene variants Shining a light on genetic screen strategies
Shining a light on genetic screen strategies CRISPR-based mapping of genetic interactions
CRISPR-based mapping of genetic interactions Combining CRISPR perturbations and RNA-seq
Combining CRISPR perturbations and RNA-seq CREATE-ing genome-wide designed mutations
CREATE-ing genome-wide designed mutations A global map of genetic interactions
A global map of genetic interactions A network to guide precision cancer therapy
A network to guide precision cancer therapy CRISPR screens go in vivo
CRISPR screens go in vivo CRISPR screening from both ways
CRISPR screening from both ways Testing all possibilities
Testing all possibilities