« Prev Next »
Personalized medicine is based on using an individual's genetic profile to make the best therapeutic choice by facilitating predictions about whether that person will benefit from a particular medicine or suffer serious side effects. Drugs are generally tested on a large population of people and the average response is reported. This sort of evidence-based medicine (that is, medical decision making based on empirical data) relies on the law of averages; personalized medicine, on the other hand, recognizes that no two patients are alike.
Basics of Pharmacogenomics
In pharmacogenomics, genomic information is used to study individual responses to drugs. When a gene variant is associated with a particular drug response in a patient, there is the potential for making clinical decisions based on genetics by adjusting the dosage or choosing a different drug, for example. Scientists assess gene variants affecting an individual's drug response the same way they assess gene variants associated with diseases: by identifying genetic loci associated with known drug responses, and then testing individuals whose response is unknown. Modern approaches include multigene analysis or whole-genome single nucleotide polymorphism (SNP) profiles, and these approaches are just coming into clinical use for drug discovery and development.
When studying drug action in individuals, researchers focus on two major determinants: (1) how much of a drug is needed to reach its target in the body, and (2) how well the target cells, such as heart tissue or neurons, respond to the drug. The scientific terms for these two determinants are pharmacokinetics and pharmacodynamics, and both are critical considerations in the field of pharmacogenomics.
Pharmacokinetics
Pharmacokinetics encompasses four processes: absorption, distribution, metabolism, and excretion, which are often abbreviated as ADME (Figure 1). Absorption usually refers to how a drug enters the bloodstream after a person takes a pill or uses an inhalant; intravenous injection circumvents absorption by putting a drug directly into the blood. Distribution describes where the drug travels after absorption and how much of the drug reaches the target site. Many drugs, for example, cannot get past the blood-brain barrier. Metabolism refers to how the drug gets broken down in the body, which can happen immediately by way of enzyme action in the stomach and sometimes involves end products with their own pharmacologic action. Finally, excretion describes how drugs leave the body, whether by urine, bile, or, in some cases, exhalation.
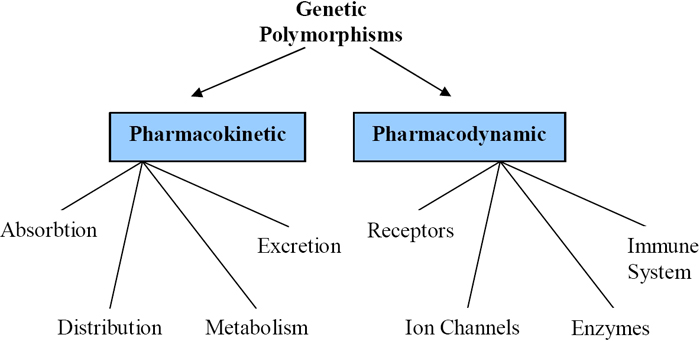
Pharmacodynamics
As previously mentioned, pharmacodynamics is the molecular action of a drug on its target, whether this is a cell surface target (e.g., a receptor), an ion channel, or an intracellular target (e.g., an enzyme or regulatory protein). For example, the beta-2 adrenergic receptor is the target of both beta-agonists in the treatment of asthma and beta-blockers in the treatment of hypertension, and this receptor has polymorphisms that have been associated with response to these drugs (Goldstein et al., 2003).
Drug Safety
Within the United States, serious side effects from pharmaceutical drugs occur in 2 million people each year and may cause as many as 100,000 deaths, according to the Food and Drug Administration. Costs associated with adverse drug reactions (ADRs) are estimated at $136 billion annually. ADRs come in two forms. One form results from misuse, such as taking too much of a medication or taking the medication too often or for too long. The second form involves the mysterious, idiosyncratic effects of various drugs. The term "idiosyncratic" is used because these (often serious) side effects are not related to drug dose and are thought to be unpredictable. Scientists believe many idiosyncratic effects result from individual variation that is encoded in the genome. Thus, genetic variation in genes for drug-metabolizing enzymes, drug receptors, and drug transporters have been associated with individual variability in the efficacy and toxicity of drugs. Genetics also underlies hypersensitivity reactions in patients who are allergic to certain drugs, such as penicillin, wherein the body mounts a rapid, aggressive immune response that can cause not only a rash, but can also hinder breathing and cause edema to the point of cardiovascular collapse.
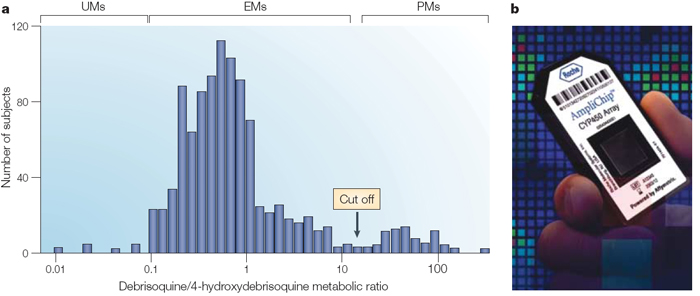
Predicting serious ADRs is a priority for pharmacogenomic research. For example, the enzyme CYP2D6, one of a class of drug-metabolizing enzymes found in the liver, breaks down and terminates the action of certain antidepressant, antiarrhythmic, and antipsychotic drugs. Molecular cloning and characterization studies of the gene that codes for this enzyme have described more than 70 variant alleles (Meyer, 2000). These alleles contain one or more point mutations, only some of which affect enzyme activity; however, some of these alleles involve gene deletions and duplications that can lead to increased enzyme activity. Individuals who are homozygous or heterozygous for the wild-type or normal activity enzymes (75%–85% of the population) are called extensive metabolizers; intermediate (10%–15%) or poor (5%–10%) metabolizers are carriers of two alleles that decrease enzyme activity (Ingelman-Sundberg, 1999); and ultrarapid metabolizers (1%–10%) are carriers of duplicated genes. The most common alleles can be detected by DNA chip microarrays, allowing most patients to be assigned to a particular phenotype group.
Genotype, Phenotype, and ADRs
If there are multiple mutations that can lead to similar susceptibilities to ADRs, diagnostic tests that directly assess the phenotype rather than a genetic mutation may be more reliable (and may remain necessary even if genetic diagnostic tests do become available). Although DNA-based technology is potentially faster and requires only a single blood sample from a patient, for many genetic variants, the correlation between genotype and phenotype has not been well described. As Weinshilboum and Wang (2004) write:
The phenotype is what the physician wants to know and, unfortunately, present DNA-based tests can fail to reflect the full range of phenotypic variation. As a result, a major challenge for companies designing DNA-based tests is to develop dependable, economical, high-throughput genotyping platforms, and a major challenge for pharmacogenomic science is to determine comprehensive, clinically useful genotype-phenotype correlations.
Differential responses to thiopurine drugs provide a concrete example of why this is true. Thiopurine drugs are used to treat acute lymphoblastic leukemia, inflammatory bowel disease, and organ transplant recipients (Weinshilboum & Wang, 2004). These drugs are useful, but they are also toxic, and the window for dosing to induce the desired therapeutic effect before causing toxicity is very narrow. The major toxicity related to thiopurines is life-threatening bone-marrow suppression.
Thiopurines are inactivated by the metabolic enzyme thiopurine S-methyltransferase (TPMT), which is encoded by a polymorphic gene. In Asian populations, TPMT*3C is a common gene variant with a cytosine at position 3, but in Caucasians, TPMT*3A is more common. TPMT*3A has two different SNPs that result in alterations in the encoded amino acids. The gene product encoded by TPMT*3A is degraded rapidly, such that individuals homozygous for this allele have little or no detectable TPMT protein in their tissues. Thus, individuals homozygous for TPMT*3A are at greatly increased risk for life-threatening myelosuppression when treated with standard doses of thiopurine drugs. However, such people can be treated with these drugs at approximately one-tenth the standard dose, but only with careful monitoring (Figure 2).
Variation in the TPMT gene is used diagnostically to evaluate dosing of thiopurines in the treatment of acute lymphoblastic leukemia. As an alternative to genotyping, TPMT activity can also be determined by a laboratory test of red blood cells. The test can identify those rare variants that might be missed at the DNA level. Interestingly, a recent study (in a sample of healthy Europeans) showed a high concordance between TMPT genotype (DNA) and phenotype (enzyme activity) (Schaeffeler et al., 2004). The authors concluded, "Because all these parameters show values higher than 90% in our large-scale study, genetic testing for TPMT is worthy of adoption into clinical practice."
Improving Cancer Outcomes
Both drugs and chemotherapy are used for the treatment of breast cancer, and diagnostic tests have allowed some limited degree of disease typing. For patients with estrogen receptor-sensitive cancer that has not yet spread to the lymph nodes, for example, tamoxifen is the drug of choice, but chemotherapy is frequently offered as an adjunct. However, chemotherapy is known to help only a small number of patients. In fact, a long-term study called the National Surgical Adjuvant Breast and Bowel Project (NSABP) found that only 4% of patients who received chemotherapy had improved outcomes (Paik et al., 2004).
Could patients receive individualized treatment based on their specific type of cancer? It appears that the answer to this question is "yes." For instance, using real-time PCR methods to study gene expression in breast cancer and taking advantage of a large collection of paraffin-preserved tissue samples, a team of NSABP researchers developed a diagnostic kit that assays 21 genes (Paik et al., 2004). This project has resulted in a diagnostic tool that identifies key genetic components of particular patients' breast cancer and can improve outcomes. This diagnostic kit predicts the likelihood of cancer recurring for patients who are categorized into one of three risk groups: low, intermediate, and high. In a clinical validation study, the diagnostic data did indeed predict long-term recurrence (Paik et al., 2006). Moreover, in studies to determine treatment benefit, the researchers analyzed tissue samples from the original NSABP study. There was no benefit to adding chemotherapy to tamoxifen in low-risk patients, and only a tiny benefit (2%) in intermediate-risk patients. In high-risk patients, though, the benefit of chemotherapy was clear, with a 28% decrease in recurrence of cancer.
Therapeutic Promise of the Human Genome Project
With the completion of the Human Genome Project, anticipation was high that genetic information would radically improve medicine, that side effects would be more predictable, and that patients could be screened for likely drug responses. But thus far, progress has been much slower than what the initial excitement suggested.
A great deal of this delay relates to the fact that an individual's response to drugs is multifactorial, resulting from multiple gene and environmental interactions (Haga & Burke, 2004). Scientists also recognize that even as the knowledge base continues to expand, the clinical translation of that knowledge still requires empirical evidence, generated for a particular disease and drug combination, before treatment can be customized to a patient's genotype. Thus, much work remains to be done before personalized medicine can reach its fullest potential.
References and Recommended Reading
Haga, S. B., & Burke, W. Using pharmacogenetics to improve drug safety and efficacy. Journal of the American Medical Association 291, 2869–2871 (2004)
Ingelman-Sundberg, M., et al. Polymorphic human cytochrome P450 enzymes: An opportunity for individualized drug treatment. Trends in Pharmacological Sciences 20, 342–349 (1999)
Meyer, U. A. Pharmacogenetics and adverse drug reactions. Lancet 356, 1667–1671 (2000) doi:10.1016/S0140-6736(00)03167-6
Paik, S., et al. A multigene assay to predict recurrence of tamoxifen-treated, node-negative breast cancer. New England Journal of Medicine 351, 2817–2826 (2004)
———. Gene expression and benefit of chemotherapy in women with node-negative, estrogen receptor-positive breast cancer. Journal of Clinical Oncology 24, 3726–3734 (2006)
Pirmohamed, M., & Park, B. K. Genetic susceptibility to adverse drug reactions. Trends in Pharmacological Sciences 22, 298–305 (2001) doi:10.1016/S0165-6147(00)01717-X (link to article)
Schaeffeler, E., et al. Comprehensive analysis of thiopurine S-methyltransferase phenotype-genotype correlation in a large population of German-Caucasians and identification of novel TPMT variants. Pharmacogenetics 14, 407–417 (2004)
Scripture, C. D., & Figg, W. D. Drug interactions in cancer therapy. Nature Reviews Cancer 6, 546–558 (2006) doi:10.1038/nrc1887 (link to article)
Weinshilboum, R., & Wang, L. Pharmacogenomics: Bench to bedside. Nature Reviews Drug Discovery 3, 739–748 (2004) doi:10.1038/nrd1497 (link to article)






























