Article
|
Open Access
Featured
-
-
Research Briefing |
 Inferring how animals deform improves cell tracking
Inferring how animals deform improves cell tracking
Tracking cells is a time-consuming part of biological image analysis, and traditional manual annotation methods are prohibitively laborious for tracking neurons in the deforming and moving Caenorhabditis elegans brain. By leveraging machine learning to develop a ‘targeted augmentation’ method, we substantially reduced the number of labeled images required for tracking.
-
Article |
 Automated neuron tracking inside moving and deforming C. elegans using deep learning and targeted augmentation
Automated neuron tracking inside moving and deforming C. elegans using deep learning and targeted augmentation
Targettrack is a deep-learning-based pipeline for automatic tracking of neurons within freely moving C. elegans. Using targeted augmentation, the pipeline has a reduced need for manually annotated training data.
- Core Francisco Park
- , Mahsa Barzegar-Keshteli
- & Sahand Jamal Rahi
-
Brief Communication
| Open Access Uniform quantification of single-nucleus ATAC-seq data with Paired-Insertion Counting (PIC) and a model-based insertion rate estimator
Uniform quantification of single-nucleus ATAC-seq data with Paired-Insertion Counting (PIC) and a model-based insertion rate estimator
This study demonstrates the need and advantage of uniformly quantifying single-nucleus ATAC-seq data using Paired-Insertion Counting.
- Zhen Miao
- & Junhyong Kim
-
Article
| Open Access Combinatorial single-cell profiling of major chromatin types with MAbID
Combinatorial single-cell profiling of major chromatin types with MAbID
MAbID offers a multiplexing approach to uncover the genomic distributions of various epigenetic markers, enabling the study of how these markers jointly direct gene expression.
- Silke J. A. Lochs
- , Robin H. van der Weide
- & Jop Kind
-
Article
| Open Access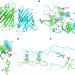 StayGold variants for molecular fusion and membrane-targeting applications
StayGold variants for molecular fusion and membrane-targeting applications
Monomeric and tandem dimer derivatives of the bright and photostable green fluorescent protein StayGold offer versatile tools for tagging target proteins and membranes in extended live-cell imaging.
- Ryoko Ando
- , Satoshi Shimozono
- & Atsushi Miyawaki
-
Article |
 Improved green and red GRAB sensors for monitoring dopaminergic activity in vivo
Improved green and red GRAB sensors for monitoring dopaminergic activity in vivo
Next-generation red and green G-protein-coupled receptor-based dopamine sensors with improved properties have been developed. Their performance is demonstrated in cell culture, in brain slices and in vivo in the mouse.
- Yizhou Zhuo
- , Bin Luo
- & Yulong Li
-
Article
| Open Access AlphaFold predictions are valuable hypotheses and accelerate but do not replace experimental structure determination
AlphaFold predictions are valuable hypotheses and accelerate but do not replace experimental structure determination
An analysis of AlphaFold protein structure predictions shows that while in many cases the predictions are highly accurate, there are also many instances where the predicted structures or parts of predicted structures do not agree with experimentally resolved data. Therefore, care must be taken when using these predictions for informing structural hypotheses.
- Thomas C. Terwilliger
- , Dorothee Liebschner
- & Paul D. Adams
-
Article |
 Improved sequence mapping using a complete reference genome and lift-over
Improved sequence mapping using a complete reference genome and lift-over
By combining fast lift-over and selective re-mapping, levioSAM2 enables efficient and accurate read mapping and variant calling leveraging complete reference genomes.
- Nae-Chyun Chen
- , Luis F. Paulin
- & Ben Langmead
-
Article
| Open Access Next-generation MRI scanner designed for ultra-high-resolution human brain imaging at 7 Tesla
Next-generation MRI scanner designed for ultra-high-resolution human brain imaging at 7 Tesla
A combination of hardware developments has increased the achievable spatial resolution in 7 Tesla human neuroimaging to about 0.4 mm.
- David A. Feinberg
- , Alexander J. S. Beckett
- & Peter Dietz
-
Research Briefing |
 Fluorescent actinometers for fast and simple quantitative measurement of light intensity
Fluorescent actinometers for fast and simple quantitative measurement of light intensity
Fluorescent actinometers enable the measurement of light intensity even in the depths of samples and over wide ranges of wavelengths and intensities. We introduce two protocols to quantitatively characterize the spatial distribution of light of various fluorescence imaging systems and to calibrate the illumination of commercially available instruments and light sources.
-
Article
| Open Access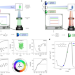 Fluorescence to measure light intensity
Fluorescence to measure light intensity
Two methods for fluorescence-based actinometry using organic dyes and photoconvertible fluorescent proteins enable rapid and precise measurement of light intensity at the sample in fluorescence microscopes.
- Aliénor Lahlou
- , Hessam Sepasi Tehrani
- & Ludovic Jullien
-
Article
| Open Access Uncovering developmental time and tempo using deep learning
Uncovering developmental time and tempo using deep learning
A deep learning-based method that uses microscopy images to stage embryos and analyze developmental time.
- Nikan Toulany
- , Hernán Morales-Navarrete
- & Patrick Müller
-
Brief Communication |
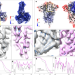 Improving resolution and resolvability of single-particle cryoEM structures using Gaussian mixture models
Improving resolution and resolvability of single-particle cryoEM structures using Gaussian mixture models
This manuscript describes a refinement protocol that extends the e2gmm method to optimize both the orientation and conformation estimation of particles to improve the alignment for flexible domains of proteins.
- Muyuan Chen
- , Michael F. Schmid
- & Wah Chiu
-
Article
| Open Access Bio-friendly long-term subcellular dynamic recording by self-supervised image enhancement microscopy
Bio-friendly long-term subcellular dynamic recording by self-supervised image enhancement microscopy
DeepSeMi is a self-supervised denoising framework that can enhance SNR over 12 dB across diverse samples and imaging modalities. DeepSeMi enables extended longitudinal imaging of subcellular dynamics with high spatiotemporal resolution.
- Guoxun Zhang
- , Xiaopeng Li
- & Qionghai Dai
-
Article
| Open Access High-fidelity 3D live-cell nanoscopy through data-driven enhanced super-resolution radial fluctuation
High-fidelity 3D live-cell nanoscopy through data-driven enhanced super-resolution radial fluctuation
Enhanced super-resolution radial fluctuations (eSRRF) offers improved image fidelity and resolution compared to the popular SRRF method and further enables volumetric live-cell super-resolution imaging at high speeds.
- Romain F. Laine
- , Hannah S. Heil
- & Ricardo Henriques
-
Article |
 Peptide sequencing based on host–guest interaction-assisted nanopore sensing
Peptide sequencing based on host–guest interaction-assisted nanopore sensing
A phenylalanine-containing peptide probe can be used for discriminating all 20 amino acids via current blockage during translocation through an α-hemolysin (αHL) nanopore. The paper provides proof-of-concept peptide sequencing demonstrations.
- Yun Zhang
- , Yakun Yi
- & Hai-Chen Wu
-
Article |
 Adaptable, turn-on maturation (ATOM) fluorescent biosensors for multiplexed detection in cells
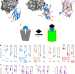
Adaptable, turn-on maturation (ATOM) fluorescent biosensors for multiplexed detection in cells
Adaptable, turn-on maturation (ATOM) biosensors use monobody or nanobody targeting to control fluorescent protein maturation for fluorescence in the presence of target biomolecules, enabling bright and specific cellular biosensing.
- Harsimranjit Sekhon
- , Jeung-Hoi Ha
- & Stewart N. Loh
-
-
-
-
Research Highlight |
 Capturing subcellular metal ion dynamics
Capturing subcellular metal ion dynamics
Newly developed DNA devices can be used to study intracellular sodium and potassium ion levels with organellar resolution.
- Rita Strack
-
Correspondence |
 METLIN-CCS: an ion mobility spectrometry collision cross section database
METLIN-CCS: an ion mobility spectrometry collision cross section database
- Erin S. Baker
- , Corey Hoang
- & Gary Siuzdak
-
Article
| Open Access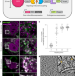 Genetically encoded multimeric tags for subcellular protein localization in cryo-EM
Genetically encoded multimeric tags for subcellular protein localization in cryo-EM
Genetically encoded multimeric particles (GEMs) are 25-nm tags with recognizable structural signatures, which can be used to label specific proteins in mammalian cells to facilitate their subcellular localization in cryo-ET.
- Herman K. H. Fung
- , Yuki Hayashi
- & Julia Mahamid
-
News & Views |
 Modeling the complexity of mammary gland in vitro
Modeling the complexity of mammary gland in vitro
A new method models mammary gland in vitro during physiological and pathological conditions.
- Marco Fioramonti
- & Cédric Blanpain
-
Article
| Open Access Causal identification of single-cell experimental perturbation effects with CINEMA-OT
Causal identification of single-cell experimental perturbation effects with CINEMA-OT
CINEMA-OT is a framework that enables accurate causal inference of the effects of perturbation experiments at the single-cell level.
- Mingze Dong
- , Bao Wang
- & David van Dijk
-
Article |
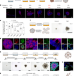 Reconstruction of dynamic mammary mini gland in vitro for normal physiology and oncogenesis
Reconstruction of dynamic mammary mini gland in vitro for normal physiology and oncogenesis
A dynamic, hormone-responsive mammary gland organoid system that can mimic complex physiological processes in vitro.
- Lei Yuan
- , Shaofang Xie
- & Shang Cai
-
Correspondence |
 Chromoscope: interactive multiscale visualization for structural variation in human genomes
Chromoscope: interactive multiscale visualization for structural variation in human genomes
- Sehi L’Yi
- , Dominika Maziec
- & Nils Gehlenborg
-
Technology Feature |
 A sound solution for deep-brain imaging
A sound solution for deep-brain imaging
Ultrasound-based modalities are revealing the brain’s inner workings with steadily increasing speed, resolution and depth.
- Michael Eisenstein
-
Article
| Open Access Pulsed stimulated Brillouin microscopy enables high-sensitivity mechanical imaging of live and fragile biological specimens
Pulsed stimulated Brillouin microscopy enables high-sensitivity mechanical imaging of live and fragile biological specimens
A pulsed illumination scheme renders stimulated Brillouin microscopy less phototoxic and allows imaging of the mechanical properties of sensitive samples such as single cells, Caenorhabditis elegans embryos, zebrafish larvae and organoids.
- Fan Yang
- , Carlo Bevilacqua
- & Robert Prevedel
-
Article
| Open Access nextPYP: a comprehensive and scalable platform for characterizing protein variability in situ using single-particle cryo-electron tomography
nextPYP: a comprehensive and scalable platform for characterizing protein variability in situ using single-particle cryo-electron tomography
nextPYP is a turn-key framework for single-particle cryo-electron tomography that streamlines complex data analysis pipelines, from pre-processing of tilt series to high-resolution refinement, for efficient analysis and visualization of large datasets.
- Hsuan-Fu Liu
- , Ye Zhou
- & Alberto Bartesaghi
-
Perspective |
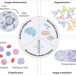 Artificial intelligence-enabled quantitative phase imaging methods for life sciences
Artificial intelligence-enabled quantitative phase imaging methods for life sciences
This Perspective introduces advances in quantitative phase imaging and artificial intelligence-based image analysis and further describes how the two technologies intersect and synergize to enable biomedical research.
- Juyeon Park
- , Bijie Bai
- & YongKeun Park
-
Research Briefing |
 Hydrogel fibers that enable optogenetic pain inhibition during locomotion
Hydrogel fibers that enable optogenetic pain inhibition during locomotion
We developed fatigue-resistant hydrogel optical fibers through the controlled growth of polymeric nanocrystalline domains to enable light delivery to peripheral nerves during locomotion. The hydrogel fibers withstand locomotion strain across more than 30,000 fiber stretch cycles and enable the optogenetic inhibition of pain hypersensitivity in naturally behaving mice.
-
Article |
 Fatigue-resistant hydrogel optical fibers enable peripheral nerve optogenetics during locomotion
Fatigue-resistant hydrogel optical fibers enable peripheral nerve optogenetics during locomotion
Flexible and fatigue-resistant optical fibers made from hydrogel allow optogenetic manipulations in the periphery in freely behaving mice.
- Xinyue Liu
- , Siyuan Rao
- & Xuanhe Zhao
-
Article |
 Microbial-enrichment method enables high-throughput metagenomic characterization from host-rich samples
Microbial-enrichment method enables high-throughput metagenomic characterization from host-rich samples
This work introduces microbial-enrichment methodology (MEM) that enables removal of host DNA in human intestinal biopsies and characterization of low-abundance microbial taxa down to 1%.
- Natalie J. Wu-Woods
- , Jacob T. Barlow
- & Rustem F. Ismagilov
-
Article
| Open Access Population-level integration of single-cell datasets enables multi-scale analysis across samples
Population-level integration of single-cell datasets enables multi-scale analysis across samples
By learning representations for both cells and various condition covariates, scPoli facilitates atlas-level integration and analysis of single-cell genomics datasets with improved interpretability.
- Carlo De Donno
- , Soroor Hediyeh-Zadeh
- & Fabian J. Theis
-
Article
| Open Access Compact engineered human mechanosensitive transactivation modules enable potent and versatile synthetic transcriptional control
Compact engineered human mechanosensitive transactivation modules enable potent and versatile synthetic transcriptional control
Synthetic CRISPR-based transactivation domains engineered using mechanosensitive transcription factors enable robust transcriptional control.
- Barun Mahata
- , Alan Cabrera
- & Isaac B. Hilton
-
Article
| Open Access OPUS-DSD: deep structural disentanglement for cryo-EM single-particle analysis
OPUS-DSD: deep structural disentanglement for cryo-EM single-particle analysis
OPUS-DSD is a neural network-based algorithm that reconstructs distinct conformations or continuous dynamics of the macromolecular structural landscape, starting from single-particle cryo-EM data.
- Zhenwei Luo
- , Fengyun Ni
- & Jianpeng Ma
-
-
Editorial |
Registered Reports: what we’ve learned so far
Nature Methods is proud to publish our very first Registered Report in this issue. Here, we reflect on what we have learned since introducing this article type.
-
Brief Communication |
 Molecular mechanocytometry using tension-activated cell tagging
Molecular mechanocytometry using tension-activated cell tagging
Tension-activated cell tagging (TaCT) is a new method that uses flow cytometry to sort mechanically active cells based on the forces generated by their surface adhesion receptors.
- Rong Ma
- , Sk Aysha Rashid
- & Khalid Salaita
-
News & Views |
 Real-time imaging of dynamic tissues
Real-time imaging of dynamic tissues
A combined modality enables real-time imaging of mouse lungs and spans whole-organ to cellular scales.
- Joan E. Nichols
- & Sasha R. Azar
-
Comment |
 A practitioner’s view of spectral flow cytometry
A practitioner’s view of spectral flow cytometry
Spectral flow cytometry enables the simultaneous analysis of a large number of cell surface markers at the single-cell level
- Siddhartha Sharma
- , Josh Boyer
- & Luc Teyton
-
Research Briefing |
 Evaluating long-read RNA-sequencing analysis tools with in silico mixtures
Evaluating long-read RNA-sequencing analysis tools with in silico mixtures
We conducted a comprehensive long-read RNA sequencing (RNA-seq) benchmarking experiment by combining spike-ins and in silico mixtures to establish a ground-truth dataset. We used long- and short-read RNA-seq technology to deeply sequence samples and compared the performance of a range of analysis tools on these data.
-
Article |
 CryoREAD: de novo structure modeling for nucleic acids in cryo-EM maps using deep learning
CryoREAD: de novo structure modeling for nucleic acids in cryo-EM maps using deep learning
Few methods for three-dimensional structure modeling of nucleic acids from cryo-EM data exist. CryoREAD, a fully automated DNA/RNA atomic structure modeling method based on deep learning, was developed to fill this gap.
- Xiao Wang
- , Genki Terashi
- & Daisuke Kihara
-
Article |
 Streamlined and sensitive mono- and di-ribosome profiling in yeast and human cells
Streamlined and sensitive mono- and di-ribosome profiling in yeast and human cells
A comprehensive redevelopment of the ribosome profiling workflow involves improved nuclease treatment and sequencing library preparation, enabling richer and more accurate translatome profiling with lower input and fewer technical hurdles.
- Lucas Ferguson
- , Heather E. Upton
- & Nicholas T. Ingolia
-
Article
| Open Access Spatial single-cell mass spectrometry defines zonation of the hepatocyte proteome
Spatial single-cell mass spectrometry defines zonation of the hepatocyte proteome
Single-cell Deep Visual Proteomics integrates imaging, cell segmentation, laser microdissection and multiplexed mass spectrometry for spatial single-cell proteomics measurements in complex tissues.
- Florian A. Rosenberger
- , Marvin Thielert
- & Matthias Mann
-
Resource
| Open Access Waxholm Space atlas of the rat brain: a 3D atlas supporting data analysis and integration
Waxholm Space atlas of the rat brain: a 3D atlas supporting data analysis and integration
An updated version of the Waxholm Space atlas of the rat brain includes more detailed annotations of several brain regions, including the cortex, striatopallidal region, midbrain and thalamus, expanding the previous version with 112 new and 57 revised structures.
- Heidi Kleven
- , Ingvild E. Bjerke
- & Trygve B. Leergaard
-
News & Views |
 Factoring single-cell perturbations
Factoring single-cell perturbations
GSFA is a statistical model to automatically detect latent factors (or gene modules) in single-cell CRISPR screening.
- Bicna Song
- & Wei Li
-
Article
| Open Access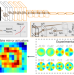 Deep learning-driven adaptive optics for single-molecule localization microscopy
Deep learning-driven adaptive optics for single-molecule localization microscopy
A deep learning approach bypasses iterative trials associated with sensorless adaptive optics to compensate for wavefront deformations when imaging biological specimens, enabling improved deep tissue localization microscopy.
- Peiyi Zhang
- , Donghan Ma
- & Fang Huang
Browse broader subjects
Browse narrower subjects
- Analytical biochemistry
- Behavioural methods
- Bioinformatics
- Biological models
- Biophysical methods
- Cytological techniques
- Electrophysiology
- Epigenetics analysis
- Experimental organisms
- Gene delivery
- Gene expression analysis
- Genetic engineering
- Genetic techniques
- Genomic analysis
- High-throughput screening
- Imaging
- Immunological techniques
- Isolation, separation and purification
- Lab-on-a-chip
- Mass spectrometry
- Metabolomics
- Microbiology techniques
- Microscopy
- Molecular engineering
- Nanobiotechnology
- Optogenetics
- Proteomic analysis
- Sensors and probes
- Sequencing
- Software
- Optical spectroscopy
- Structure determination

 Embryo mechanics cartography: inference of 3D force atlases from fluorescence microscopy
Embryo mechanics cartography: inference of 3D force atlases from fluorescence microscopy Art appreciation
Art appreciation Modeling morphogenesis
Modeling morphogenesis Resolving epigenetic base ambiguity
Resolving epigenetic base ambiguity