Research Briefing |
Featured
-
-
Article |
 VIBRANT: spectral profiling for single-cell drug responses
VIBRANT: spectral profiling for single-cell drug responses
Vibrational painting (VIBRANT) is a high-content single-cell phenotypic profiling method using mid-infrared imaging with vibrational probes for metabolic activity, which offers high accuracy with minimal batch effects to capture cellular responses to perturbation.
- Xinwen Liu
- , Lixue Shi
- & Wei Min
-
Article
| Open Access TREX reveals proteins that bind to specific RNA regions in living cells
TREX reveals proteins that bind to specific RNA regions in living cells
TREX introduces an RNA-centric tool for identifying proteins binding to specific RNA regions in living cells.
- Martin Dodel
- , Giulia Guiducci
- & Faraz K. Mardakheh
-
Article
| Open Access Generation of complex bone marrow organoids from human induced pluripotent stem cells
Generation of complex bone marrow organoids from human induced pluripotent stem cells
The authors describe stem cell-derived bone marrow organoids that accurately model the structural and functional properties of the human bone marrow niche.
- Stephanie Frenz-Wiessner
- , Savannah D. Fairley
- & Christoph Klein
-
Correspondence |
 OpenMS 3 enables reproducible analysis of large-scale mass spectrometry data
OpenMS 3 enables reproducible analysis of large-scale mass spectrometry data
- Julianus Pfeuffer
- , Chris Bielow
- & Timo Sachsenberg
-
Article
| Open Access Multiscale biochemical mapping of the brain through deep-learning-enhanced high-throughput mass spectrometry
Multiscale biochemical mapping of the brain through deep-learning-enhanced high-throughput mass spectrometry
MEISTER is an integrative experimental and computational framework for mass spectrometry that integrates three-dimensional, organ-wide biomolecular mapping with single-cell analysis for multiscale profiling of spatial–biochemical organization.
- Yuxuan Richard Xie
- , Daniel C. Castro
- & Fan Lam
-
Article
| Open Access Toward universal cell embeddings: integrating single-cell RNA-seq datasets across species with SATURN
Toward universal cell embeddings: integrating single-cell RNA-seq datasets across species with SATURN
SATURN performs cross-species integration and analysis using both single-cell gene expression and protein representations generated by protein language models.
- Yanay Rosen
- , Maria Brbić
- & Jure Leskovec
-
Research Briefing |
 Deciphering protein interaction network dynamics with a machine learning-based framework
Deciphering protein interaction network dynamics with a machine learning-based framework
We developed Tapioca, an integrative ensemble machine learning-based framework, to accurately predict global protein–protein interaction network dynamics. Tapioca enabled the characterization of host regulation during reactivation from latency of an oncogenic virus. Introducing an interactome homology analysis method, we identified a proviral host factor with broad relevance for herpesviruses.
-
Article |
 Tapioca: a platform for predicting de novo protein–protein interactions in dynamic contexts
Tapioca: a platform for predicting de novo protein–protein interactions in dynamic contexts
Tapioca is an ensemble machine learning framework for studying protein–protein interactions (PPIs) that facilitates integration of curve-based dynamic PPI data from thermal proximity coaggregation, ion-based proteome-integrated solubility alteration or cofractionation mass spectrometry data with static interaction data to predict PPIs in dynamic contexts.
- Tavis. J. Reed
- , Matthew. D. Tyl
- & Ileana. M. Cristea
-
-
-
-
Editorial |
Where imaging and metrics meet
When it comes to bioimaging and image analysis, details matter. Papers in this issue offer guidance for improved robustness and reproducibility.
-
Article
| Open Access A paintbrush for delivery of nanoparticles and molecules to live cells with precise spatiotemporal control
A paintbrush for delivery of nanoparticles and molecules to live cells with precise spatiotemporal control
Micro-kiss (μkiss) is a micropipette-based approach for delivering very small amounts of nanoparticles and small molecules to the cell surface with exquisite spatiotemporal control, enabling a wide range of biological investigations.
- Cornelia Holler
- , Richard William Taylor
- & Vahid Sandoghdar
-
This Month |
From zero to hero in budget-making
The grant deadline approaches but budget details are still missing. Experienced budget-makers share how they manage budgets and the ‘boom–bust’ cycle.
- Vivien Marx
-
Article
| Open Access Exonuclease-enhanced prime editors
Exonuclease-enhanced prime editors
Exo-PE is an approach to improve prime editing efficacy for insertions while maintaining precision.
- Dong-Jiunn Jeffery Truong
- , Julian Geilenkeuser
- & Gil Gregor Westmeyer
-
Brief Communication
| Open Access An improved pathway for autonomous bioluminescence imaging in eukaryotes
An improved pathway for autonomous bioluminescence imaging in eukaryotes
Improvements to the fully genetically encoded Neonothopanus nambi bioluminescence pathway enhance autobioluminescence by up to two orders of magnitude in plants and other species, enabling novel applications of bioluminescence imaging in biology.
- Ekaterina S. Shakhova
- , Tatiana A. Karataeva
- & Alexander S. Mishin
-
Brief Communication
| Open Access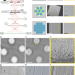 Square beams for optimal tiling in transmission electron microscopy
Square beams for optimal tiling in transmission electron microscopy
A square electron beam improves imaging of large fields of view in transmission electron microscopes by facilitating montage tomography of vitrified specimens with no loss in data quality relative to conventional round beams.
- Eugene Y. D. Chua
- , Lambertus M. Alink
- & Alex de Marco
-
Article |
 Deep generative design of RNA family sequences
Deep generative design of RNA family sequences
RNA family sequence generator (RfamGen) is a deep generative model for designing novel, functional RNA sequences. RfamGen is applicable to diverse RNA families and can yield ribozymes with higher enzymatic activity.
- Shunsuke Sumi
- , Michiaki Hamada
- & Hirohide Saito
-
Article
| Open Access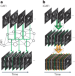 Content-aware frame interpolation (CAFI): deep learning-based temporal super-resolution for fast bioimaging
Content-aware frame interpolation (CAFI): deep learning-based temporal super-resolution for fast bioimaging
Content-aware frame interpolation (CAFI) improves the temporal resolution in time-lapse imaging by accurately predicting images in between image pairs. By allowing fewer frames to be imaged, CAFI also enables gentler live-cell imaging.
- Martin Priessner
- , David C. A. Gaboriau
- & Romain F. Laine
-
Article |
 Massively parallel single-cell sequencing of diverse microbial populations
Massively parallel single-cell sequencing of diverse microbial populations
DoTA-seq leverages a microfluidic droplet system to isolate and lyse diverse microbes and amplify target genetic loci, enabling high-throughput single-cell sequencing of microbial populations.
- Freeman Lan
- , Jason Saba
- & Ophelia S. Venturelli
-
-
-
Editorial |
Year in review 2023
As we begin a new year, we reflect on some of our favorites among the papers published in 2023 in Nature Methods.
-
Research Highlight |
 Predicting neural activity from facial expressions
Predicting neural activity from facial expressions
Facemap tracks keypoints on the mouse face and feeds the information into a deep neural network to predict neural activity.
- Nina Vogt
-
Research Briefing |
 ARTR-seq for determining RNA-binding protein target sites through in situ reverse transcription
ARTR-seq for determining RNA-binding protein target sites through in situ reverse transcription
ARTR-seq uses antibody-guided in situ reverse transcription to efficiently and accurately identify RNA-binding protein target sites in as few as 20 cells, or in a formaldehyde-fixed tissue section. The high temporal resolution of ARTR-seq opens opportunities for the investigation of dynamic RNA-binding protein–RNA interactions.
-
Correspondence |
 RepoRT: a comprehensive repository for small molecule retention times
RepoRT: a comprehensive repository for small molecule retention times
- Fleming Kretschmer
- , Eva-Maria Harrieder
- & Michael Witting
-
Article |
 Real-time visualization of structural dynamics of synapses in live cells in vivo
Real-time visualization of structural dynamics of synapses in live cells in vivo
SynapShot combines ddFPs with engineered synaptic adhesion molecules for real-time observation of the structural plasticity of synapses in cultured cells and animals.
- Seungkyu Son
- , Kenichiro Nagahama
- & Won Do Heo
-
Article |
 Mapping enzyme activity in living systems by real-time mid-infrared photothermal imaging of nitrile chameleons
Mapping enzyme activity in living systems by real-time mid-infrared photothermal imaging of nitrile chameleons
Real-time mid-infrared photothermal imaging of nitrile chameleons enables simultaneous, multiplexed measurement of enzymatic activity in living systems and is poised to reveal the spatiotemporal regulation of enzymes in health and disease.
- Hongjian He
- , Jiaze Yin
- & Ji-Xin Cheng
-
Article
| Open Access Image restoration of degraded time-lapse microscopy data mediated by near-infrared imaging
Image restoration of degraded time-lapse microscopy data mediated by near-infrared imaging
InfraRed-mediated Image Restoration (IR2) uses deep learning to combine the benefits of deep-tissue imaging with NIR probes and the convenience of imaging with GFP for improved time-lapse imaging of embryogenesis.
- Nicola Gritti
- , Rory M. Power
- & Jan Huisken
-
Article |
 mEnrich-seq: methylation-guided enrichment sequencing of bacterial taxa of interest from microbiome
mEnrich-seq: methylation-guided enrichment sequencing of bacterial taxa of interest from microbiome
mEnrich-seq leverages DNA methylation differences between bacteria to enrich taxa of interest from metagenomic samples for selective sequencing.
- Lei Cao
- , Yimeng Kong
- & Gang Fang
-
Technology Feature |
 Aging research comes of age
Aging research comes of age
As money pours into aging research, the field can combine its many methods to home in on what underpins aging. Approaches differ, but researchers share the desire to not overpromise quick-fix anti-aging methods.
- Vivien Marx
-
This Month |
If at first you don’t succeed
Scientists have successes to celebrate but must also cope with the sting of failures. In the way she handles both, Nobel laureate Katalin Karikó inspires others.
- Vivien Marx
-
Article |
 Smart lattice light-sheet microscopy for imaging rare and complex cellular events
Smart lattice light-sheet microscopy for imaging rare and complex cellular events
smartLLSM uses artificial intelligence-based instrument control to switch between epiflouorescence and lattice light-sheet microscopy to monitor cells at the population level while also capturing multicolor three-dimensional datasets of rare events of interest.
- Yu Shi
- , Jimmy S. Tabet
- & Wesley R. Legant
-
Article
| Open Access Thermal-plex: fluidic-free, rapid sequential multiplexed imaging with DNA-encoded thermal channels
Thermal-plex: fluidic-free, rapid sequential multiplexed imaging with DNA-encoded thermal channels
The thermal-plex method for highly multiplexed imaging uses DNA probes activated when briefly elevated to designated temperatures for rapid, fluidics-free sequential imaging in cells and tissues.
- Fan Hong
- , Jocelyn Y. Kishi
- & Peng Yin
-
Research Briefing |
 Ultra-long-working-distance multiphoton objective unlocks new possibilities for imaging
Ultra-long-working-distance multiphoton objective unlocks new possibilities for imaging
In 1858, the first standard for microscope objectives was established to encourage interchangeable components. Over the following 150 years, standards have evolved to constrain the size of objectives, which limits the parameters of working distance, field of view and resolution. A new design breaks out of this conventional envelope, offering an ultra-long working distance in air and enabling new neuroscience experiments.
-
Article
| Open Access The Cousa objective: a long-working distance air objective for multiphoton imaging in vivo
The Cousa objective: a long-working distance air objective for multiphoton imaging in vivo
The Cousa objective is an ultra-long working distance air objective optimized for two- and three-photon imaging. Bypassing challenges caused by water immersion and short working distances, the Cousa enables and improves imaging of diverse specimens.
- Che-Hang Yu
- , Yiyi Yu
- & Spencer LaVere Smith
-
Article
| Open Access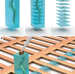 Serial Lift-Out: sampling the molecular anatomy of whole organisms
Serial Lift-Out: sampling the molecular anatomy of whole organisms
Serial Lift-Out creates a series of lamellae from one lift-out volume for cryo-ET, increasing the ease and throughput of cryo-lift-out and enabling the study of molecular anatomy in multicellular systems including C. elegans larvae.
- Oda Helene Schiøtz
- , Christoph J. O. Kaiser
- & Jürgen M. Plitzko
-
Article |
 A data analysis framework for combining multiple batches increases the power of isobaric proteomics experiments
A data analysis framework for combining multiple batches increases the power of isobaric proteomics experiments
msTrawler improves quantitative analysis of multiplexed mass spectrometry proteomics data when combining isobaric data across multiple batches.
- Jonathon J. O’Brien
- , Anil Raj
- & Fiona E. McAllister
-
Article |
 DeepMainmast: integrated protocol of protein structure modeling for cryo-EM with deep learning and structure prediction
DeepMainmast: integrated protocol of protein structure modeling for cryo-EM with deep learning and structure prediction
DeepMainmast is a protein structure modeling protocol for cryo-EM that combines the strengths of a deep-learning-based de novo protein main-chain-tracing approach with AlphaFold2-based structure predictions for improved performance.
- Genki Terashi
- , Xiao Wang
- & Daisuke Kihara
-
Method to Watch |
 Imaging across scales
Imaging across scales
New twists on established methods and multimodal imaging are poised to bridge gaps between cellular and organismal imaging.
- Rita Strack
-
Method to Watch |
 Nanopores for sequencing proteins
Nanopores for sequencing proteins
Developments in nanopore-based peptide detection and sequencing show promise of a breakthrough.
- Arunima Singh
-
Method to Watch |
 Spatially resolved multiomics
Spatially resolved multiomics
Spatially resolved multimodal omics offers a collective way to capture molecular information in complex tissues.
- Lei Tang
-
Method to Watch |
 Visual proteomics
Visual proteomics
Advances will enable proteome-scale structure determination in cells.
- Rita Strack
-
Method to Watch |
 Synthetic tissue environments
Synthetic tissue environments
Artificial ECMs enable recapitulation of tissue microenvironments.
- Madhura Mukhopadhyay
-
-
-
Method to Watch |
 Human brain mapping
Human brain mapping
High-resolution connectomics of the human brain is the next frontier in neuroscience.
- Nina Vogt
-
Research Highlight |
 Greater diversity for lineage tracing
Greater diversity for lineage tracing
DARLIN enables the generation of a massive diversity of barcodes for in vivo lineage tracing and the combination with single-cell multi-omics measurements.
- Lei Tang
-
Editorial |
 Method of the Year 2023: methods for modeling development
Method of the Year 2023: methods for modeling development
In vitro embryo models supported by methods development in adjacent fields have revolutionized our understanding of embryogenesis.
Browse broader subjects
Browse narrower subjects
- Analytical biochemistry
- Behavioural methods
- Bioinformatics
- Biological models
- Biophysical methods
- Cytological techniques
- Electrophysiology
- Epigenetics analysis
- Experimental organisms
- Gene delivery
- Gene expression analysis
- Genetic engineering
- Genetic techniques
- Genomic analysis
- High-throughput screening
- Imaging
- Immunological techniques
- Isolation, separation and purification
- Lab-on-a-chip
- Mass spectrometry
- Metabolomics
- Microbiology techniques
- Microscopy
- Molecular engineering
- Nanobiotechnology
- Optogenetics
- Proteomic analysis
- Sensors and probes
- Sequencing
- Software
- Optical spectroscopy
- Structure determination

 TREX identifies region-specific protein interactors of RNA molecules
TREX identifies region-specific protein interactors of RNA molecules Chroma is a generative model for protein design
Chroma is a generative model for protein design Better detection for localization microscopy
Better detection for localization microscopy