Article
|
Open Access
Featured
-
-
Brief Communication |
 Square condenser apertures for square cameras in low-dose transmission electron microscopy
Square condenser apertures for square cameras in low-dose transmission electron microscopy
Square and rectangular beams made with new condenser apertures enable more efficient image capture in low-dose transmission electron microscopy with no compromise to image quality in single-particle cryoelectron microscopy.
- Hamish G. Brown
- , Dan Smith
- & Eric Hanssen
-
Article |
 Learning structural heterogeneity from cryo-electron sub-tomograms with tomoDRGN
Learning structural heterogeneity from cryo-electron sub-tomograms with tomoDRGN
TomoDRGN is a deep learning framework designed to model conformational and compositional heterogeneity from cryo-ET datasets on a per-particle basis.
- Barrett M. Powell
- & Joseph H. Davis
-
Brief Communication
| Open Access Square beams for optimal tiling in transmission electron microscopy
Square beams for optimal tiling in transmission electron microscopy
A square electron beam improves imaging of large fields of view in transmission electron microscopes by facilitating montage tomography of vitrified specimens with no loss in data quality relative to conventional round beams.
- Eugene Y. D. Chua
- , Lambertus M. Alink
- & Alex de Marco
-
Article
| Open Access Serial Lift-Out: sampling the molecular anatomy of whole organisms
Serial Lift-Out: sampling the molecular anatomy of whole organisms
Serial Lift-Out creates a series of lamellae from one lift-out volume for cryo-ET, increasing the ease and throughput of cryo-lift-out and enabling the study of molecular anatomy in multicellular systems including C. elegans larvae.
- Oda Helene Schiøtz
- , Christoph J. O. Kaiser
- & Jürgen M. Plitzko
-
Method to Watch |
 Visual proteomics
Visual proteomics
Advances will enable proteome-scale structure determination in cells.
- Rita Strack
-
Brief Communication |
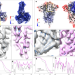 Improving resolution and resolvability of single-particle cryoEM structures using Gaussian mixture models
Improving resolution and resolvability of single-particle cryoEM structures using Gaussian mixture models
This manuscript describes a refinement protocol that extends the e2gmm method to optimize both the orientation and conformation estimation of particles to improve the alignment for flexible domains of proteins.
- Muyuan Chen
- , Michael F. Schmid
- & Wah Chiu
-
Article
| Open Access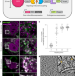 Genetically encoded multimeric tags for subcellular protein localization in cryo-EM
Genetically encoded multimeric tags for subcellular protein localization in cryo-EM
Genetically encoded multimeric particles (GEMs) are 25-nm tags with recognizable structural signatures, which can be used to label specific proteins in mammalian cells to facilitate their subcellular localization in cryo-ET.
- Herman K. H. Fung
- , Yuki Hayashi
- & Julia Mahamid
-
Article
| Open Access OPUS-DSD: deep structural disentanglement for cryo-EM single-particle analysis
OPUS-DSD: deep structural disentanglement for cryo-EM single-particle analysis
OPUS-DSD is a neural network-based algorithm that reconstructs distinct conformations or continuous dynamics of the macromolecular structural landscape, starting from single-particle cryo-EM data.
- Zhenwei Luo
- , Fengyun Ni
- & Jianpeng Ma
-
Article |
 CryoREAD: de novo structure modeling for nucleic acids in cryo-EM maps using deep learning
CryoREAD: de novo structure modeling for nucleic acids in cryo-EM maps using deep learning
Few methods for three-dimensional structure modeling of nucleic acids from cryo-EM data exist. CryoREAD, a fully automated DNA/RNA atomic structure modeling method based on deep learning, was developed to fill this gap.
- Xiao Wang
- , Genki Terashi
- & Daisuke Kihara
-
Research Briefing |
 Capturing detailed cellular landscapes by montage cryo-electron tomography
Capturing detailed cellular landscapes by montage cryo-electron tomography
To capture expansive, seamless fields of view from frozen hydrated specimens by cryo-electron tomography, we developed methods for the collection and processing of montage data. This approach enables rapid acquisition of contiguous regions of specimens using a montaged tilt series collection scheme.
-
Article |
 Time-resolved cryo-EM using a combination of droplet microfluidics with on-demand jetting
Time-resolved cryo-EM using a combination of droplet microfluidics with on-demand jetting
This article describes a time-resolved cryo-plunger that combines a droplet-based microfluidic mixer with a laser-induced generator of microjets that allows rapid reaction initiation and plunge-freezing of cryo-EM grids. A time resolution of 5 ms was achieved using this approach.
- Stefania Torino
- , Mugdha Dhurandhar
- & Rouslan G. Efremov
-
Brief Communication |
 New measures of anisotropy of cryo-EM maps
New measures of anisotropy of cryo-EM maps
This paper proposes two new anisotropy metrics—the Fourier shell occupancy and the Bingham test—that can be used to understand the quality of cryogenic electron microscopy maps.
- Jose-Luis Vilas
- & Hemant D. Tagare
-
Article
| Open Access TomoTwin: generalized 3D localization of macromolecules in cryo-electron tomograms with structural data mining
TomoTwin: generalized 3D localization of macromolecules in cryo-electron tomograms with structural data mining
TomoTwin is a deep metric learning-based particle picking method for cryo-electron tomograms. TomoTwin obviates the need for annotating training data and retraining a picking model for each protein.
- Gavin Rice
- , Thorsten Wagner
- & Stefan Raunser
-
Research Briefing |
 Mapping the motion and structure of flexible proteins from cryo-EM data
Mapping the motion and structure of flexible proteins from cryo-EM data
A deep learning algorithm maps out the continuous conformational changes of flexible protein molecules from single-particle cryo-electron microscopy images, allowing the visualization of the conformational landscape of a protein with improved resolution of its moving parts.
-
Article
| Open Access 3DFlex: determining structure and motion of flexible proteins from cryo-EM
3DFlex: determining structure and motion of flexible proteins from cryo-EM
3D Flexible Refinement (3DFlex) is a generative neural network model for continuous molecular heterogeneity for cryo-EM data that can be used to determine the structure and motion of flexible biomolecules. It enables visualization of nonrigid motion and improves 3D structure resolution by aggregating information from particle images spanning the conformational landscape of the target molecule.
- Ali Punjani
- & David J. Fleet
-
Correspondence |
 DAQ-Score Database: assessment of map–model compatibility for protein structure models from cryo-EM maps
DAQ-Score Database: assessment of map–model compatibility for protein structure models from cryo-EM maps
- Tsukasa Nakamura
- , Xiao Wang
- & Daisuke Kihara
-
Technology Feature |
 Tuning in to epigenetic cross-talk
Tuning in to epigenetic cross-talk
Chemical modifications to DNA, histones and RNA make changes happen. Scientists are exploring ways to track these modifications and how they interact.
- Vivien Marx
-
Review Article |
 Cryo-electron tomography on focused ion beam lamellae transforms structural cell biology
Cryo-electron tomography on focused ion beam lamellae transforms structural cell biology
This Review describes advances in cryogenic electron tomography on focused ion beam lamellae, highlighting the key benefits of this technology for in situ structural biology and discussing important future directions.
- Casper Berger
- , Navya Premaraj
- & Peter J. Peters
-
-
News & Views |
 Seeing the wood for the trees
Seeing the wood for the trees
A deep learning approach called DeepPiCt facilitates segmentation and macromolecular identification in the cellular jungle of electron cryotomography data.
- Olivia E. R. Smith
- & Tanmay A. M. Bharat
-
Article
| Open Access Integrated multimodality microscope for accurate and efficient target-guided cryo-lamellae preparation
Integrated multimodality microscope for accurate and efficient target-guided cryo-lamellae preparation
Cryogenic correlated light, ion and electron microscopy (cryo-CLIEM) integrates three-dimensional confocal microscopy with focused ion beam–scanning electron microscopy for efficient preparation of lamellae containing target structures for in situ structural biology with cryo-electron tomography.
- Weixing Li
- , Jing Lu
- & Wei Ji
-
Article
| Open Access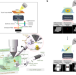 ELI trifocal microscope: a precise system to prepare target cryo-lamellae for in situ cryo-ET study
ELI trifocal microscope: a precise system to prepare target cryo-lamellae for in situ cryo-ET study
The ELI-TriScope advances cryo-CLEM by focusing light, electron and ion beams on cryopreserved samples for markedly improved preparation of cryo-lamellae containing target structures.
- Shuoguo Li
- , Ziyan Wang
- & Fei Sun
-
Method to Watch |
 Fluorescence in structural biology
Fluorescence in structural biology
Advances in fluorescence microscopy and spectroscopy show their promise for applications that complement in situ structural biology methods like cryoelectron tomography.
- Rita Strack
-
Article
| Open Access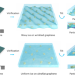 Uniform thin ice on ultraflat graphene for high-resolution cryo-EM
Uniform thin ice on ultraflat graphene for high-resolution cryo-EM
This paper shows that the uniformity of vitreous ice thickness relies on the surface flatness of the supporting film, and presents a method to use ultraflat graphene as the support for cryo-EM specimen preparation.
- Liming Zheng
- , Nan Liu
- & Hailin Peng
-
Research Briefing |
 Fast cryo-electron tomography data acquisition to determine molecular structures in situ
Fast cryo-electron tomography data acquisition to determine molecular structures in situ
To accelerate data acquisition for in situ cryo-electron tomography, we created a method that takes into consideration sample geometry for the robust prediction of sample movement while the microscope stage is tilted. This approach enabled the parallel collection of tens to hundreds of tilt series.
-
Article |
 Parallel cryo electron tomography on in situ lamellae
Parallel cryo electron tomography on in situ lamellae
The parallel cryo electron tomography (PACE-tomo) method increases the throughput on in situ samples by parallelizing acquisition. It maximizes the usable sample area on individual lamellae without compromising data quality.
- Fabian Eisenstein
- , Haruaki Yanagisawa
- & Radostin Danev
-
Research Briefing |
 Obtaining cryo-EM structures by scanning transmission electron microscopy
Obtaining cryo-EM structures by scanning transmission electron microscopy
Scanning transmission electron microscopy (STEM) techniques reveal atomic-resolution details of organic and inorganic materials. The application of STEM to biological vitrified specimens under low-dose cryogenic imaging conditions demonstrates that STEM also achieves near-atomic-resolution 3D structures of biological macromolecules.
-
Article
| Open Access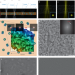 Single-particle cryo-EM structures from iDPC–STEM at near-atomic resolution
Single-particle cryo-EM structures from iDPC–STEM at near-atomic resolution
This paper explores the use of scanning transmission electron microscopy (STEM) to vitrified biological samples for biomolecular structure elucidation. Integrated differential phase contrast (iDPC)–STEM imaging of keyhole limpet hemocyanin and tobacco mosaic virus enabled cryo-EM structure determination at 6.5 and 3.5 Å resolution, respectively.
- Ivan Lazić
- , Maarten Wirix
- & Carsten Sachse
-
This Month |
 Tamir Gonen
Tamir Gonen
MicroED for structure determination without a model and with heavy metal guitar riffs.
- Vivien Marx
-
Article
| Open Access Ab initio phasing macromolecular structures using electron-counted MicroED data
Ab initio phasing macromolecular structures using electron-counted MicroED data
This article reports sub- and near-atomic structures of triclinic lysozyme and serine protease proteinase K, respectively, providing first demonstrations of ab initio phasing using electron counted MicroED data to solve macromolecular structures.
- Michael W. Martynowycz
- , Max T. B. Clabbers
- & Tamir Gonen
-
Article |
 Sub-3-Å cryo-EM structure of RNA enabled by engineered homomeric self-assembly
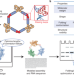
Sub-3-Å cryo-EM structure of RNA enabled by engineered homomeric self-assembly
ROCK (RNA oligomerization-enabled cryo-EM via installing kissing loops) enables improved single-particle cryo-EM of RNAs. ROCK was used to generate high-quality structures of three diverse RNAs, including the Tetrahymena group I intron.
- Di Liu
- , François A. Thélot
- & Peng Yin
-
Editorial |
 Methods lead the way
Methods lead the way
The COVID-19 pandemic has highlighted the importance of methodological advancements in basic biological research. We believe that method development will continue to propel both fundamental and applied studies on SARS-CoV-2 and other pathogens.
-
Comment |
 Fighting SARS-CoV-2 with structural biology methods
Fighting SARS-CoV-2 with structural biology methods
High-resolution structural information is critical for rapid development of vaccines and therapeutics against emerging human pathogens. Structural biology methods have been at the forefront of research on SARS-CoV-2 since the beginning of the COVID-19 pandemic. These technologies will continue to be powerful tools to fend off future public health threats.
- Jun Zhang
- & Bing Chen
-
Article |
 CR-I-TASSER: assemble protein structures from cryo-EM density maps using deep convolutional neural networks
CR-I-TASSER: assemble protein structures from cryo-EM density maps using deep convolutional neural networks
CR-I-TASSER integrates deep neural-network learning with I-TASSER assembly simulations for automated cryo-EM structure determination.
- Xi Zhang
- , Biao Zhang
- & Yang Zhang
-
Method to Watch |
 A dynamic direction for cryo-EM
A dynamic direction for cryo-EM
Emerging algorithms are extracting information about macromolecular motions from cryo-EM data.
- Allison Doerr
-
Comment |
 A paradigm shift in structural biology
A paradigm shift in structural biology
The release of protein structure predictions from AlphaFold will increase the number of protein structural models by almost three orders of magnitude. Structural biology and bioinformatics will never be the same, and the need for incisive experimental approaches will be greater than ever. Combining these advances in structure prediction with recent advances in cryo-electron microscopy suggests a new paradigm for structural biology.
- Sriram Subramaniam
- & Gerard J. Kleywegt
-
News & Views |
 Neural networks learn the motions of molecular machines
Neural networks learn the motions of molecular machines
New computational approaches capture molecular motion from cryo-EM images and provide a more complete understanding of protein dynamics.
- Timothy Grant
-
Article |
 Deep learning-based mixed-dimensional Gaussian mixture model for characterizing variability in cryo-EM
Deep learning-based mixed-dimensional Gaussian mixture model for characterizing variability in cryo-EM
e2gmm uses a deep neural network with a Gaussian representation to resolve the compositional and conformational variability within biomolecules using cryo-EM data.
- Muyuan Chen
- & Steven J. Ludtke
-
Comment |
 Cryo-electron tomography: observing the cell at the atomic level
Cryo-electron tomography: observing the cell at the atomic level
Interest in cryo-ET is rapidly growing, but many technical challenges still require solutions.
- Xueming Li
-
Comment |
 Molecular visualization of cellular complexity
Molecular visualization of cellular complexity
Cryo-ET, which yields both structural and spatial information, will help unlock the molecular architecture of cells.
- Michael R. Wozny
- & Wanda Kukulski
-
Comment |
 Expanding capabilities and infrastructure for cryo-EM
Expanding capabilities and infrastructure for cryo-EM
Many challenges and considerations must be evaluated when expanding and supporting new cryo-electron microscopy facilities.
- Kutti R. Vinothkumar
-
Comment |
 RNA structure: a renaissance begins?
RNA structure: a renaissance begins?
Advances in cryo-EM technology will open a new era of RNA-only 3D structure determination.
- Rhiju Das
-
News & Views |
 Improving cryo-EM structure validation
Improving cryo-EM structure validation
A community-wide challenge yields recommendations for improving cryo-EM structure validation.
- Alexis Rohou
-
This Month |
 Julia Mahamid
Julia Mahamid
Jumping down rabbit holes for a 3.5-Ångstrom peek at molecules.
- Vivien Marx
-
Article |
 Multi-particle cryo-EM refinement with M visualizes ribosome-antibiotic complex at 3.5 Å in cells
Multi-particle cryo-EM refinement with M visualizes ribosome-antibiotic complex at 3.5 Å in cells
The software M establishes a reference-based multi-particle refinement framework for cryo-EM data. Combined with CTF correction and map denoising, M enables residue-level structure determination inside cells.
- Dimitry Tegunov
- , Liang Xue
- & Julia Mahamid
-
Article |
 CryoDRGN: reconstruction of heterogeneous cryo-EM structures using neural networks
CryoDRGN: reconstruction of heterogeneous cryo-EM structures using neural networks
CryoDRGN is an unsupervised machine learning algorithm that reconstructs continuous distributions of three-dimensional density maps from heterogeneous single-particle cryo-EM data.
- Ellen D. Zhong
- , Tristan Bepler
- & Joseph H. Davis
-
Analysis
| Open Access Cryo-EM model validation recommendations based on outcomes of the 2019 EMDataResource challenge
Cryo-EM model validation recommendations based on outcomes of the 2019 EMDataResource challenge
A multi-laboratory study in the form of a community challenge assesses the quality of models that can be produced from cryo-EM maps using different software tools, the reproducibility of models generated by different users and the performance of metrics used for model validation.
- Catherine L. Lawson
- , Andriy Kryshtafovych
- & Wah Chiu
-
Article |
 A ‘Build and Retrieve’ methodology to simultaneously solve cryo-EM structures of membrane proteins
A ‘Build and Retrieve’ methodology to simultaneously solve cryo-EM structures of membrane proteins
The iterative Build and Retrieve (BaR) methodology facilitates the solving of cryo-EM structures of multiple membrane (and soluble) proteins simultaneously, including small and low-abundance membrane proteins.
- Chih-Chia Su
- , Meinan Lyu
- & Edward W. Yu
-
Research Highlight |
 Cryo-EM goes atomic
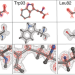
Cryo-EM goes atomic
Technological advances push the limits of single-particle cryo-EM to achieve atomic resolution.
- Rita Strack

 Electrospray-assisted cryo-EM sample preparation to mitigate interfacial effects
Electrospray-assisted cryo-EM sample preparation to mitigate interfacial effects Contrast enhancement for cryo-ET
Contrast enhancement for cryo-ET