Technology Feature |
Featured
-
-
Editorial |
 Methods lead the way
Methods lead the way
The COVID-19 pandemic has highlighted the importance of methodological advancements in basic biological research. We believe that method development will continue to propel both fundamental and applied studies on SARS-CoV-2 and other pathogens.
-
Method to Watch |
 A dynamic direction for cryo-EM
A dynamic direction for cryo-EM
Emerging algorithms are extracting information about macromolecular motions from cryo-EM data.
- Allison Doerr
-
-
-
Research Highlight |
 High-speed protein crystallography
High-speed protein crystallography
The European XFEL produces its first protein structures.
- Allison Doerr
-
Research Highlights |
 Seeing DNA
Seeing DNA
DNA and chromatin structures can be visualized in situ with electron tomography.
- Zachary J Lapin
-
Technology Feature |
 Structural biology: doors open at the European XFEL
Structural biology: doors open at the European XFEL
X-ray beams at 27,000 pulses per second promise high-resolution views of macromolecules.
- Vivien Marx
-
Research Highlights |
Zooming into the larval zebrafish brain
A serial-section electron microscopy data set of larval zebrafish brain—imaged at several scales—provides a resource for structure–function analyses of the animals' neural circuitry.
- Nina Vogt
-
Tools in Brief |
GPCR function insights by cryo-EM
-
Research Highlights |
Protein holography
Low-energy holography enables imaging single proteins and protein complexes.
- Zachary J Lapin
-
News & Views |
 Cool and dynamic: single-molecule fluorescence-based structural biology
Cool and dynamic: single-molecule fluorescence-based structural biology
New methods exploit single-molecule measurements to study protein structure and dynamics.
- Timothy D Craggs
-
Tools in Brief |
Faster to a structure with RELION-2
-
Research Highlights |
Blend-and-shoot crystallography
A rapid mix-and-inject serial femtosecond crystallography approach enables structure determination of ligand-binding intermediates.
- Allison Doerr
-
Editorial |
Method of the Year 2015
The end of 'blob-ology': single-particle cryo-electron microscopy (cryo-EM) is now being used to solve macromolecular structures at high resolution.
-
Research Highlights |
Metalloenzyme structures in a shot
Researchers study the structure of the metalloenzyme photosystem II by applying femtosecond X-ray pulses to simultaneously record X-ray diffraction and X-ray emission spectroscopy data.
- Allison Doerr
-
Resource |
 Interactome3D: adding structural details to protein networks
Interactome3D: adding structural details to protein networks
Interactome3D is an interactive resource allowing biologists to use available structural information to model protein-protein interactions and structurally annotate protein interaction networks.
- Roberto Mosca
- , Arnaud Céol
- & Patrick Aloy
-
News & Views |
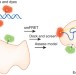 Six steps closer to FRET-driven structural biology
Six steps closer to FRET-driven structural biology
A new toolbox for structural biology that combines single-molecule fluorescence and molecular modeling is used to generate high-precision structures of protein complexes.
- Timothy D Craggs
- & Achillefs N Kapanidis
-
Article |
 A toolkit and benchmark study for FRET-restrained high-precision structural modeling
A toolkit and benchmark study for FRET-restrained high-precision structural modeling
A collection of simulation tools and workflow for single-molecule Förster resonance energy transfer (smFRET) allows highly quantitative structural modeling. This hybrid approach yields a model of reverse-transcriptase binding to DNA at sub-angstrom accuracy when benchmarked against a crystal structure and can resolve a flexible single-stranded template overhang.
- Stanislav Kalinin
- , Thomas Peulen
- & Claus A M Seidel
-
News & Views |
 Spinning up mass spectrometry for whole protein complexes
Spinning up mass spectrometry for whole protein complexes
Researchers adapt a popular Orbitrap-based mass spectrometer to detect and analyze large, intact protein complexes, accelerating a new frontier in structural biology.
- Philip D Compton
- & Neil L Kelleher
-
Brief Communication |
 High-sensitivity Orbitrap mass analysis of intact macromolecular assemblies
High-sensitivity Orbitrap mass analysis of intact macromolecular assemblies
Modifications to an Orbitrap-based mass spectrometer enable analysis of large protein complexes in native-like states by mass spectrometry with very high sensitivity and mass resolution.
- Rebecca J Rose
- , Eugen Damoc
- & Albert J R Heck
-
Brief Communication |
 Reversible labeling of native and fusion-protein motifs
Reversible labeling of native and fusion-protein motifs
Removing phosphopantetheine-tagged labels from acyl carrier proteins (ACPs) and ACP fusion proteins contributes to a versatile labeling method in which tags can be iteratively swapped.
- Nicolas M Kosa
- , Robert W Haushalter
- & Michael D Burkart
-
Article |
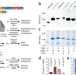 Facile backbone structure determination of human membrane proteins by NMR spectroscopy
Facile backbone structure determination of human membrane proteins by NMR spectroscopy
The combination of cell-free protein expression and combinatorial dual labeling–aided NMR analysis allows for the rapid backbone structure assessment of human membrane proteins.
- Christian Klammt
- , Innokentiy Maslennikov
- & Senyon Choe
-
Tools in Brief |
Tags to disentangle Dicer
-
Article |
 Three-dimensional RNA structure refinement by hydroxyl radical probing
Three-dimensional RNA structure refinement by hydroxyl radical probing
Hydroxyl radical probing measurements of large RNA molecules drive molecular dynamics simulations to generate three-dimensional structural models.
- Feng Ding
- , Christopher A Lavender
- & Nikolay V Dokholyan
-
Brief Communication |
 Lipidic phase membrane protein serial femtosecond crystallography
Lipidic phase membrane protein serial femtosecond crystallography
Lipidic sponge phase crystallization yields membrane protein microcrystals that can be injected into an X-ray free electron laser beam, yielding diffraction patterns that can be processed to recover the crystal structure.
- Linda C Johansson
- , David Arnlund
- & Richard Neutze
-
Brief Communication |
 In vivo protein crystallization opens new routes in structural biology
In vivo protein crystallization opens new routes in structural biology
Expression of a protein in Sf9 insect cells at high concentration triggers formation of in vivo crystals that can be analyzed by serial femtosecond X-ray crystallography.
- Rudolf Koopmann
- , Karolina Cupelli
- & Michael Duszenko
-
Brief Communication |
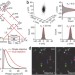 Dual-objective STORM reveals three-dimensional filament organization in the actin cytoskeleton
Dual-objective STORM reveals three-dimensional filament organization in the actin cytoskeleton
The use of dual-objective detection with astigmatism-based three-dimensional stochastic optical reconstruction microscopy (STORM) imaging improves resolution more than twofold and removes noise in resulting super-resolution images. This allowed detailed fluorescence imaging of distinctive features of the three-dimensional actin cytoskeleton ultrastructure with single-filament resolution in cells.
- Ke Xu
- , Hazen P Babcock
- & Xiaowei Zhuang
-
Review Article |
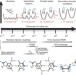 Characterizing RNA dynamics at atomic resolution using solution-state NMR spectroscopy
Characterizing RNA dynamics at atomic resolution using solution-state NMR spectroscopy
In this Review, the authors thoroughly discuss solution-state NMR spectroscopy methods used for characterizing the dynamic structure landscape of RNA molecules, from picosecond to second timescale motions.
- Jameson R Bothe
- , Evgenia N Nikolova
- & Hashim M Al-Hashimi
-
News in Brief |
High-resolution structures from low-resolution data
-
Research Highlights |
News in brief
-
Research Highlights |
Taming crystals' whimsy
Molecularly imprinted polymers act as 'smart' nucleants for protein crystallization.
- Petya V Krasteva
-
Research Highlights |
News in brief
-
Brief Communication |
 Near-infrared branding efficiently correlates light and electron microscopy
Near-infrared branding efficiently correlates light and electron microscopy
Fiducial marks that can be visualized by both light and electron microscopy are generated by 'branding' fixed tissue with a near-infrared laser and will facilitate correlative light and electron microscopy.
- Derron Bishop
- , Ivana Nikić
- & Thomas Misgeld
-
Article |
 Sequence-based identification of 3D structural modules in RNA with RMDetect
Sequence-based identification of 3D structural modules in RNA with RMDetect
Three-dimensional structural RNA modules, defined as ensembles of stacked arrays of ordered non-Watson-Crick base pairs, are found in many RNAs and play important functional roles. The presented computational tool, RMDetect, allows the identification of common RNA modules from sequence alone.
- José Almeida Cruz
- & Eric Westhof
-
Research Highlights |
News in brief
-
Brief Communication |
 High-throughput single-molecule optofluidic analysis
High-throughput single-molecule optofluidic analysis
A microfluidic mixing device for multiple, rapid and automated single-molecule measurements permits the study of macromolecule properties under varying environmental conditions. Also in this issue, Gambin et al. present another microfluidic mixing device for rapid single-molecule measurements.
- Soohong Kim
- , Aaron M Streets
- & Devdoot S Majumdar
-
This Month |
 Erik Jorgensen
Erik Jorgensen
Fluorescent proteins can be located in electron micrographs.
- Monya Baker
- & Erik Jorgensen
-
Article |
 Protein localization in electron micrographs using fluorescence nanoscopy
Protein localization in electron micrographs using fluorescence nanoscopy
Methods are reported for the combination of fluorescence nanoscopy using either stimulated emission depletion microscopy (STED) or photoactivated localization microscopy (PALM) with electron microscopy, to achieve correlative imaging in which the super-resolved fluorescence signal is placed in the context of cellular ultrastructure.
- Shigeki Watanabe
- , Annedore Punge
- & Erik M Jorgensen
-
Article |
 Maltose–neopentyl glycol (MNG) amphiphiles for solubilization, stabilization and crystallization of membrane proteins
Maltose–neopentyl glycol (MNG) amphiphiles for solubilization, stabilization and crystallization of membrane proteins
Readily synthesized maltose–neopentyl glycol (MNG) amphiphiles are useful reagents for stabilizing, extracting and crystallizing a variety of integral membrane proteins and have favorable properties relative to conventional detergents.
- Pil Seok Chae
- , Søren G F Rasmussen
- & Samuel H Gellman
-
Research Highlights |
 Protein structure gets exciting
Protein structure gets exciting
Researchers determined the excited-state structure of a small protein using nuclear magnetic resonance spectroscopy.
- Allison Doerr
-
Article |
 Monitoring multiple distances within a single molecule using switchable FRET
Monitoring multiple distances within a single molecule using switchable FRET
Single-molecule fluorescence resonance energy transfer (FRET) is a useful technique for monitoring biomolecular dynamics. A new method, termed switchable FRET, facilitates monitoring of multiple distances in single molecules, using a single donor and multiple spectrally identical acceptors that are switched on and off between a fluorescent state and a dark state.
- Stephan Uphoff
- , Seamus J Holden
- & Achillefs N Kapanidis
-
Research Highlights |
NMR and the elusive GPCR
Researchers solve a high-resolution structure of a seven-helix transmembrane protein using nuclear magnetic resonance (NMR) spectroscopy.
- Allison Doerr
-
Technology Feature |
 Making membrane proteins for structures: a trillion tiny tweaks
Making membrane proteins for structures: a trillion tiny tweaks
Researchers try multiple means to get high-quality membrane proteins for X-ray crystallography and nuclear magnetic resonance (NMR) spectroscopy studies.
- Monya Baker
-
Brief Communication |
 Detecting the conformation of individual proteins in live cells
Detecting the conformation of individual proteins in live cells
Single-molecule fluorescence resonance energy transfer (smFRET) is applied in live cells and reveals the conformational changes of individual SNARE proteins upon entering a SNARE complex.
- John J Sakon
- & Keith R Weninger

 Tuning in to epigenetic cross-talk
Tuning in to epigenetic cross-talk