Featured
-
-
-
Article
| Open Access Quantification of absolute labeling efficiency at the single-protein level
Quantification of absolute labeling efficiency at the single-protein level
Super-resolution imaging of reference and target structures enables precise determination of the labeling efficiency of high-affinity binding proteins in cells for improved quantitative assessment of protein organization at the single-molecule level.
- Joschka Hellmeier
- , Sebastian Strauss
- & Ralf Jungmann
-
Article
| Open Access Virtual reality-empowered deep-learning analysis of brain cells
Virtual reality-empowered deep-learning analysis of brain cells
Generating training data for training deep-learning-based tools is time consuming. The DELiVR pipeline facilitates this process as demonstrated in this study on detecting c-Fos+ cells or microglia in the brain, following tissue clearing and imaging with light-sheet microscopy.
- Doris Kaltenecker
- , Rami Al-Maskari
- & Ali Ertürk
-
Article |
 Selective-plane-activation structured illumination microscopy
Selective-plane-activation structured illumination microscopy
The combination of light sheet illumination and reversibly switchable fluorophores enables improved structured illumination microscopy for fast, low-background super-resolution imaging in cells and spheroids.
- Kenta Temma
- , Ryosuke Oketani
- & Katsumasa Fujita
-
Article
| Open Access A multicolor suite for deciphering population coding of calcium and cAMP in vivo
A multicolor suite for deciphering population coding of calcium and cAMP in vivo
Improved green cAMP and red calcium sensors were developed to facilitate dual-color imaging in vivo. These sensors will allow studying the relationship between calcium and cAMP signaling.
- Tatsushi Yokoyama
- , Satoshi Manita
- & Masayuki Sakamoto
-
Article |
 Bright and stable monomeric green fluorescent protein derived from StayGold
Bright and stable monomeric green fluorescent protein derived from StayGold
mBaoJin is a monomeric derivative of the bright and photostable green fluorescent protein StayGold. mBaoJin offers favorable photophysical properties for use in diverse protein tagging and subcellular labeling applications.
- Hanbin Zhang
- , Gleb D. Lesnov
- & Fedor V. Subach
-
Article
| Open Access Image restoration of degraded time-lapse microscopy data mediated by near-infrared imaging
Image restoration of degraded time-lapse microscopy data mediated by near-infrared imaging
InfraRed-mediated Image Restoration (IR2) uses deep learning to combine the benefits of deep-tissue imaging with NIR probes and the convenience of imaging with GFP for improved time-lapse imaging of embryogenesis.
- Nicola Gritti
- , Rory M. Power
- & Jan Huisken
-
Article |
 Smart lattice light-sheet microscopy for imaging rare and complex cellular events
Smart lattice light-sheet microscopy for imaging rare and complex cellular events
smartLLSM uses artificial intelligence-based instrument control to switch between epiflouorescence and lattice light-sheet microscopy to monitor cells at the population level while also capturing multicolor three-dimensional datasets of rare events of interest.
- Yu Shi
- , Jimmy S. Tabet
- & Wesley R. Legant
-
Article
| Open Access Thermal-plex: fluidic-free, rapid sequential multiplexed imaging with DNA-encoded thermal channels
Thermal-plex: fluidic-free, rapid sequential multiplexed imaging with DNA-encoded thermal channels
The thermal-plex method for highly multiplexed imaging uses DNA probes activated when briefly elevated to designated temperatures for rapid, fluidics-free sequential imaging in cells and tissues.
- Fan Hong
- , Jocelyn Y. Kishi
- & Peng Yin
-
Research Briefing |
 Ultra-long-working-distance multiphoton objective unlocks new possibilities for imaging
Ultra-long-working-distance multiphoton objective unlocks new possibilities for imaging
In 1858, the first standard for microscope objectives was established to encourage interchangeable components. Over the following 150 years, standards have evolved to constrain the size of objectives, which limits the parameters of working distance, field of view and resolution. A new design breaks out of this conventional envelope, offering an ultra-long working distance in air and enabling new neuroscience experiments.
-
-
Article |
 Automated neuron tracking inside moving and deforming C. elegans using deep learning and targeted augmentation
Automated neuron tracking inside moving and deforming C. elegans using deep learning and targeted augmentation
Targettrack is a deep-learning-based pipeline for automatic tracking of neurons within freely moving C. elegans. Using targeted augmentation, the pipeline has a reduced need for manually annotated training data.
- Core Francisco Park
- , Mahsa Barzegar-Keshteli
- & Sahand Jamal Rahi
-
Article
| Open Access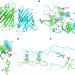 StayGold variants for molecular fusion and membrane-targeting applications
StayGold variants for molecular fusion and membrane-targeting applications
Monomeric and tandem dimer derivatives of the bright and photostable green fluorescent protein StayGold offer versatile tools for tagging target proteins and membranes in extended live-cell imaging.
- Ryoko Ando
- , Satoshi Shimozono
- & Atsushi Miyawaki
-
Research Briefing |
 Fluorescent actinometers for fast and simple quantitative measurement of light intensity
Fluorescent actinometers for fast and simple quantitative measurement of light intensity
Fluorescent actinometers enable the measurement of light intensity even in the depths of samples and over wide ranges of wavelengths and intensities. We introduce two protocols to quantitatively characterize the spatial distribution of light of various fluorescence imaging systems and to calibrate the illumination of commercially available instruments and light sources.
-
Article
| Open Access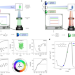 Fluorescence to measure light intensity
Fluorescence to measure light intensity
Two methods for fluorescence-based actinometry using organic dyes and photoconvertible fluorescent proteins enable rapid and precise measurement of light intensity at the sample in fluorescence microscopes.
- Aliénor Lahlou
- , Hessam Sepasi Tehrani
- & Ludovic Jullien
-
Article
| Open Access Bio-friendly long-term subcellular dynamic recording by self-supervised image enhancement microscopy
Bio-friendly long-term subcellular dynamic recording by self-supervised image enhancement microscopy
DeepSeMi is a self-supervised denoising framework that can enhance SNR over 12 dB across diverse samples and imaging modalities. DeepSeMi enables extended longitudinal imaging of subcellular dynamics with high spatiotemporal resolution.
- Guoxun Zhang
- , Xiaopeng Li
- & Qionghai Dai
-
Article
| Open Access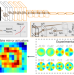 Deep learning-driven adaptive optics for single-molecule localization microscopy
Deep learning-driven adaptive optics for single-molecule localization microscopy
A deep learning approach bypasses iterative trials associated with sensorless adaptive optics to compensate for wavefront deformations when imaging biological specimens, enabling improved deep tissue localization microscopy.
- Peiyi Zhang
- , Donghan Ma
- & Fang Huang
-
Research Briefing |
 Mapping deformations and increasing quantitative accuracy in expansion microscopy
Mapping deformations and increasing quantitative accuracy in expansion microscopy
We introduce GelMap, a flexible workflow for reporting deformations and anisotropy in expansion microscopy. By intrinsically calibrating the expansion hydrogel using a fluorescent grid that scales with expansion and deforms with anisotropy, GelMap enables the reliable quantification of expansion factors and correction of deformations.
-
Article
| Open Access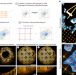 GelMap: intrinsic calibration and deformation mapping for expansion microscopy
GelMap: intrinsic calibration and deformation mapping for expansion microscopy
The GelMap workflow adds a fluorescent grid into samples before expansion, allowing for precise determination of expansion factor and subsequent deformation correction in ExM. GelMap works with diverse samples and expansion methods.
- Hugo G. J. Damstra
- , Josiah B. Passmore
- & Lukas C. Kapitein
-
Article |
 Large Stokes shift fluorescent RNAs for dual-emission fluorescence and bioluminescence imaging in live cells
Large Stokes shift fluorescent RNAs for dual-emission fluorescence and bioluminescence imaging in live cells
The Clivias are a series of small, monomeric fluorescent RNAs that emit with a large Stokes shift in the orange–red. They enable multiplexed RNA imaging in live cells and BRET-based detection of protein–RNA interactions in mice.
- Li Jiang
- , Xin Xie
- & Yi Yang
-
Article
| Open Access Statistically unbiased prediction enables accurate denoising of voltage imaging data
Statistically unbiased prediction enables accurate denoising of voltage imaging data
Statistically unbiased prediction utilizing spatiotemporal information in imaging data (SUPPORT) is a self-supervised deep learning approach to accurately denoise voltage and calcium imaging data while preserving true dynamic signals.
- Minho Eom
- , Seungjae Han
- & Young-Gyu Yoon
-
Research Highlight |
 A closer look at chromatin
A closer look at chromatin
An expansion microscopy technique called ChromExM offers detailed views into the organization chromatin and associated gene expression machinery in embryos.
- Rita Strack
-
Registered Report |
 Quantitative assessment of near-infrared fluorescent proteins
Quantitative assessment of near-infrared fluorescent proteins
This Registered Report describes an extensive comparison of 22 near-infrared fluorescent proteins in vitro, in cultured mammalian cells, and in model animals, clarifying top performers in diverse biological settings.
- Hanbin Zhang
- , Stavrini Papadaki
- & Kiryl D. Piatkevich
-
Research Briefing |
 Ultrafast nLight indicators for sensitive and specific in vivo imaging of norepinephrine
Ultrafast nLight indicators for sensitive and specific in vivo imaging of norepinephrine
We developed, characterized and validated nLight sensors, a new family of genetically encoded green and red fluorescent norepinephrine indicators based on an alpha-1 adrenergic receptor. nLight probes can detect norepinephrine in living animals with superior sensitivity, ligand specificity and temporal resolution as compared with previous tools.
-
Brief Communication
| Open Access Open-3DSIM: an open-source three-dimensional structured illumination microscopy reconstruction platform
Open-3DSIM: an open-source three-dimensional structured illumination microscopy reconstruction platform
Open-3DSIM is a versatile open-source software for high-fidelity reconstruction of three-dimensional structured illumination microscopy data (with polarization). It is available in three convenient forms for user-friendly and customizable applications.
- Ruijie Cao
- , Yaning Li
- & Peng Xi
-
Article |
 Sensitive multicolor indicators for monitoring norepinephrine in vivo
Sensitive multicolor indicators for monitoring norepinephrine in vivo
Red and green genetically encoded indicators for norepinephrine have been developed and employed to monitor norepinephrine during locomotion and reward behavior in mice. The strategy used for generating these indicators also produced indicators for other neuromodulators.
- Zacharoula Kagiampaki
- , Valentin Rohner
- & Tommaso Patriarchi
-
Article
| Open Access Voltage-Seq: all-optical postsynaptic connectome-guided single-cell transcriptomics
Voltage-Seq: all-optical postsynaptic connectome-guided single-cell transcriptomics
Voltage-Seq combines voltage imaging, optogenetics and single-cell RNA-seq for high-throughput analysis of functional and transcriptomic properties of neurons in situ.
- Veronika Csillag
- , Marianne Hiriart Bizzozzero
- & János Fuzik
-
Editorial |
 What’s next for bioimage analysis?
What’s next for bioimage analysis?
Advanced bioimage analysis tools are poised to disrupt the way in which microscopy images are acquired and analyzed. This Focus issue shares the hopes and opinions of experts on the near and distant future of image analysis.
-
News & Views |
 Lighting up action potentials with fast and bright voltage sensors
Lighting up action potentials with fast and bright voltage sensors
Three groundbreaking studies have created a new generation of genetically encoded voltage indicators, empowering us to tackle a host of questions on our path toward understanding the brain.
- Alessio Andreoni
- & Lin Tian
-
Article |
 A positively tuned voltage indicator for extended electrical recordings in the brain
A positively tuned voltage indicator for extended electrical recordings in the brain
The ASAP4 family of genetically encoded voltage indicators allows recording of action potentials and subthreshold activity with either one- or two-photon microscopy over extended periods of time.
- S. Wenceslao Evans
- , Dong-Qing Shi
- & Michael Z. Lin
-
Research Highlight |
 Capturing hyperspectral images
Capturing hyperspectral images
A single-shot hyperspectral phasor camera (SHy-Cam) enables fast, multiplexed volumetric imaging.
- Rita Strack
-
Research Highlight |
 Rethinking microscope objectives
Rethinking microscope objectives
A microscope objective inspired by the Schmidt telescope offers a large field of view, high numerical aperture, long working distance and compatibility with all homogeneous immersion media for versatile bioimaging.
- Rita Strack
-
Article
| Open Access Virtual-scanning light-field microscopy for robust snapshot high-resolution volumetric imaging
Virtual-scanning light-field microscopy for robust snapshot high-resolution volumetric imaging
Virtual-scanning light-field microscopy (VsLFM) uses a physics-based deep learning model to improve the quality and speed of LFM, reducing motion artifacts and enabling challenging demonstrations such as fast 3D voltage imaging in Drosophila.
- Zhi Lu
- , Yu Liu
- & Qionghai Dai
-
Article
| Open Access Rapid detection of neurons in widefield calcium imaging datasets after training with synthetic data
Rapid detection of neurons in widefield calcium imaging datasets after training with synthetic data
DeepWonder removes background signals from widefield calcium recordings and enables accurate and efficient neuronal segmentation with high throughput.
- Yuanlong Zhang
- , Guoxun Zhang
- & Qionghai Dai
-
Article |
 ERnet: a tool for the semantic segmentation and quantitative analysis of endoplasmic reticulum topology
ERnet: a tool for the semantic segmentation and quantitative analysis of endoplasmic reticulum topology
ERnet is a deep learning-based software tool for automatic segmentation and classification of structures in the endoplasmic reticulum. ERnet is compatible with many fluorescence imaging modalities and can uncover subtle phenotypic changes.
- Meng Lu
- , Charles N. Christensen
- & Clemens F. Kaminski
-
Brief Communication |
 An optical design enabling lightweight and large field-of-view head-mounted microscopes
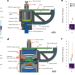
An optical design enabling lightweight and large field-of-view head-mounted microscopes
Two miniature microscopes with innovative light paths are described and applied to imaging of juvenile zebra finches and mice.
- Joseph R. Scherrer
- , Galen F. Lynch
- & Michale S. Fee
-
News & Views |
 Deep brain imaging on the move
Deep brain imaging on the move
New three-photon miniature microscopes open the study of neuronal networks to those deep in the brains of behaving animals.
- Jérôme A. Lecoq
- , Roman Boehringer
- & Benjamin F. Grewe
-
Article |
 Miniature three-photon microscopy maximized for scattered fluorescence collection
Miniature three-photon microscopy maximized for scattered fluorescence collection
A three-photon miniature microscope with optimized light-collection efficiency facilitates imaging of neuronal activity throughout the cortex, as well as in the hippocampus, in freely moving mice.
- Chunzhu Zhao
- , Shiyuan Chen
- & Heping Cheng
-
Research Highlight |
 Next-generation expansion microscopy
Next-generation expansion microscopy
A new twist on expansion microscopy called Magnify uses a mechanically sturdy gel to simultaneously anchor and expand diverse biological samples for super-resolution imaging.
- Rita Strack
-
Brief Communication |
 LILAC: enhanced actin imaging with an optogenetic Lifeact
LILAC: enhanced actin imaging with an optogenetic Lifeact
LILAC is a photoactivatable version of Lifeact, a tool for labeling F-actin. LILAC can help avoid cytotoxicity, which is sometimes associated with the use of Lifeact.
- Kourtney L. Kroll
- , Alexander R. French
- & Ronald S. Rock
-
Article
| Open Access HyU: Hybrid Unmixing for longitudinal in vivo imaging of low signal-to-noise fluorescence
HyU: Hybrid Unmixing for longitudinal in vivo imaging of low signal-to-noise fluorescence
Hybrid Unmixing offers enhanced imaging of multiplexed fluorescence labels, enabling longitudinal imaging of multiple fluorescent signals with reduced illumination intensities.
- Hsiao Ju Chiang
- , Daniel E. S. Koo
- & Francesco Cutrale
-
Article
| Open Access Integrated multimodality microscope for accurate and efficient target-guided cryo-lamellae preparation
Integrated multimodality microscope for accurate and efficient target-guided cryo-lamellae preparation
Cryogenic correlated light, ion and electron microscopy (cryo-CLIEM) integrates three-dimensional confocal microscopy with focused ion beam–scanning electron microscopy for efficient preparation of lamellae containing target structures for in situ structural biology with cryo-electron tomography.
- Weixing Li
- , Jing Lu
- & Wei Ji
-
Method to Watch |
 A light switch for targeted genomics
A light switch for targeted genomics
The combination of microscopy, targeted illumination and single-cell sequencing is driving applications from direct evolution to spatial omics.
- Rita Strack
-
Matters Arising |
 Assessment of 3D MINFLUX data for quantitative structural biology in cells
Assessment of 3D MINFLUX data for quantitative structural biology in cells
- Kirti Prakash
- & Alistair P. Curd
-
Matters Arising |
 Reply to: Assessment of 3D MINFLUX data for quantitative structural biology in cells
Reply to: Assessment of 3D MINFLUX data for quantitative structural biology in cells
- Klaus C. Gwosch
- , Francisco Balzarotti
- & Stefan W. Hell
-
Brief Communication |
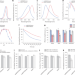 Deep-tissue SWIR imaging using rationally designed small red-shifted near-infrared fluorescent protein
Deep-tissue SWIR imaging using rationally designed small red-shifted near-infrared fluorescent protein
miRFP718nano is a rationally designed small near-infrared fluorescent protein with an emission tail that extends into the short-wave infrared range for improved multiplexed and deep-tissue imaging applications.
- Olena S. Oliinyk
- , Chenshuo Ma
- & Vladislav V. Verkhusha
-
Brief Communication |
 An improved imaging system that corrects MS2-induced RNA destabilization
An improved imaging system that corrects MS2-induced RNA destabilization
An improved version of the MS2-MCP system for imaging RNA dynamics involves tethering translation termination factors to tagged mRNAs to bypass destabilization caused by NMD machinery.
- Weihan Li
- , Anna Maekiniemi
- & Robert H. Singer
-
Research Highlight |
 Imaging-based spatial transcriptomics goes electric
Imaging-based spatial transcriptomics goes electric
Researchers use electric fields to transfer RNA from a tissue sample onto a surface for subsequent fluorescence in situ hybridization-based profiling of transcriptomes at the single-cell level.
- Rita Strack
-
Article
| Open Access Incorporating the image formation process into deep learning improves network performance
Incorporating the image formation process into deep learning improves network performance
Richardson–Lucy Network (RLN) combines the traditional Richardson–Lucy iteration with deep learning for improved deconvolution. RLN is more generalizable, offers fewer artifacts and requires less computing time than alternative approaches.
- Yue Li
- , Yijun Su
- & Hari Shroff

 Dissecting cell membrane tension dynamics and its effect on Piezo1-mediated cellular mechanosensitivity using force-controlled nanopipettes
Dissecting cell membrane tension dynamics and its effect on Piezo1-mediated cellular mechanosensitivity using force-controlled nanopipettes Fast and multiplexed DNA-PAINT
Fast and multiplexed DNA-PAINT Better detection for localization microscopy
Better detection for localization microscopy