Abstract
E26 avian leukemia oncogene 2, 3′ domain (Ets2) has been implicated in various biological processes. An Ets2 mutant model (Ets2db1/db1), which lacks the DNA-binding domain, was previously reported to exhibit embryonic lethality caused by a trophoblast abnormality. This phenotype could be rescued by tetraploid complementation, resulting in pups with wavy hair and curly whiskers. Here, we generated new Ets2 mutant models with a frame-shift mutation in exon 8 using the CRISPR/Cas9 method. Homozygous mutants could not be obtained by natural mating as embryonic development stopped before E8.5, as previously reported. When we rescued them by tetraploid complementation, these mice did not exhibit wavy hair or curly whisker phenotypes. Our newly generated mice exhibited exon 8 skipping, which led to in-frame mutant mRNA expression in the skin and thymus but not in E7.5 Ets2em1/em1 embryos. This exon 8-skipped Ets2 mRNA was translated into protein, suggesting that this Ets2 mutant protein complemented the Ets2 function in the skin. Our data implies that novel splicing variants incidentally generated after genome editing may complicate the phenotypic analysis but may also give insight into the new mechanisms related to biological gene functions.
Similar content being viewed by others
Introduction
E26 avian leukemia oncogene 2, 3′ domain (Ets2), a member of the ETS family, is a transcription factor that contains an ETS winged helix-loop-helix DNA-binding domain (ETS domain) that binds to GGA(A/T) DNA sequences. It is conserved in various species, including mice and humans1,2,3. Ets2 has been implicated in various biological contexts, including placentation, hair formation, mammary tumors, inflammatory responses, angiogenesis, and the pulmonary fibrosis4,5,6,7,8.
In a previous study, Ets2-deficient mice (Ets2db1/db1 mice), which lack the ETS domain through deletion of exons 9 and 10, were found to exhibit early embryonic lethality due to a trophectoderm abnormality. The tetraploid complementation technique could rescue this placental abnormality, allowing for survival of the offspring4, indicating that Ets2 is essential for placental development. Ets2db1/db1 mice created using the tetraploid complementation technique exhibit a variety of phenotypes, such as wavy hair, curly whisker, and a rounded forehead, allowing them to be identified. However, their fertility is normal, and they exhibit no lethal phenotype after birth. Therefore, the Ets2db1/db1 mouse is a useful model for studying treatment methods for placental abnormalities9.
The generation of gene-deficient animal models is now commonly performed using CRISPR/Cas9-based genome engineering10,11. Model organisms made using this technique can completely mimic the genome mutations found in human diseases, such as indel mutations and substitutions, which were previously difficult to generate using the conventional knockout method. Further, homozygous mutant mice can be obtained efficiently in the founder generation by directly delivering the crRNA/tracrRNA/Cas9 ribonucleoprotein complex into a mouse zygote via electroporation12. Unfortunately, if the homozygous mutant exhibits embryonic lethality, it cannot be obtained in this way. However, it is possible to obtain placental-deficient mutant mice, such as Ets2, in the founder generation using the tetraploid complementation method13 in combination with genome-edited zygotes or their embryonic stem cells.
Using the above strategy, we established three new Ets2 mutant mouse lines. Two of those contained a frame-shift deletion in exon 8, which was located before the ETS domain encoded by exons 9 and 10. These genomic mutations were predicted to produce a transcriptional product that would undergo nonsense-mediated mRNA decay (NMD) or, if translated, a protein lacking the ETS domain. The third one had the null mutant allele which lacked all open reading frame regions. We found that some of the phenotypes exhibited by the frame-shift mutant mice differed from the previous study, whose origin was investigated in this work.
Results
Generation of new Ets2 mutant mice by CRISPR/Cas9 system using zygotes
On the basis of a previous study4, we designed three gRNA targeted to sites in exon 8 that would induce a frame-shift mutation, leading to a deficiency in the ETS domain, encoded by exons 9 and 10 (Fig. 1A). The riboprotein complex, which consisted of three designed crRNAs, tracrRNAs, and Cas9 protein, was electroporated into one-cell stage zygotes, which developed until the eight-cell stage. These genome-edited embryos were used for the tetraploid complementation method in order to obtain homozygous mutant mice in the founder generation (Fig. 1B). However, no homozygous mutant mice were born from the 29 transferred embryos.
Generation of new Ets2 mutant models using zygotes. (A) Design of crRNA targeting sites in exon 8 of the Ets2 gene and checking primer positions. (B) Strategy for obtaining Ets2 homozygous mutant l in F0 generation using the electroporation technique and the tetraploid method. (C) Genotyping of the two newly generated Ets2 mutants in F0 generation. Both had a wildtype (WT: 564 bp) allele and a deletion allele (em1 or em2), which were detected as shorter bands than WT. (D) DNA sequence of mutant alleles and crRNA targeted sequences. The 205 bp deleted (Δ205 bp) allele was named em1, and the 82 bp deleted (Δ82 bp) allele was named em2.
Two out of three delivered pups had a heterozygous deletion mutation, which was determined by PCR analysis. One mutation was a 205-bp deletion (hereafter referred to as em1), and the other was an 82 bp deletion (hereafter referred to as em2) (Fig. 1C,D and full-length gel of Fig. 1C is presented in Supplemental Fig. S1). Expectedly, both had frame-shift mutations.
Assessment of the development of the newly generated Ets2 mutant mice
Previous reports indicated that Ets2db1/db1 mice exhibit an embryonic lethal phenotype due to a placental deficiency4. Sixteen pups were obtained from three derivations, and as expected, none of the pups included the double mutant alleles (em1/em2) (Table 1). Further, we analyzed the developmental ability of Ets2 mutant mice by performing a test cross using Ets2+/em1 mice and assessed the genotypes of the offspring. No homozygous mutant pups (Ets2em1/em1) were born (wild:hetero:homo = 45:91:0, Table 1).
A previous study reported that Ets2db1/db1 embryos were degenerated by the placental deficiency around E7.5 and disappeared after E8.5. To investigate whether the Ets2em1/em1 mutant phenocopies the Ets2db1/db1 mutant, we crossed Ets2+/em1 animals and observed embryos at several stages. Ets2em1/em1 embryos at E7.5 were slightly delayed in their developmental stage but clearly progressed in a comparable manner to embryos from Ets2db1/db1 animals. The Ets2em1/em1 embryos had survived at E8.5, but all of them were retarded. By E9.5 and E10.5, some malformed Ets2em1/em1 embryos were present and developed before the turning of the embryo, which usually occurred at approximately E8.5 (Fig. 2A; Table 2).
Development of Ets2em1/em1 embryos. (A) From E7.5 to E10.5, embryos were observed after crossing Ets2+/em1 females and males. Ets2em1/em1 embryos are indicated as circles in each picture. The regions outside of the circles correspond to Ets2+/+ or Ets2+/em1 embryos. The genotype of the embryo indicated by an asterisk at E9.5 could not be determined. All scale bars indicate 1 mm. (B) Gene expressions of E7.5 WT and Ets2em1/em1 (em1) embryos. Embryos were separated into trophectodermal tissue (TE), including ectoplacental corn, and epiblast (Epi), from which RNA and cDNA were prepared. Both em1 bands were shifted to be lower than the WT bands. The DNA sequences of the em1 bands were the same as em1L in Fig. 4B.
As the frame-shift mutation in Ets2em1/em1 is located in exon 8, the stop codon occurs before exon 9, and the original stop codon is located in exon 10. For this reason, NMD might occur, such that the em1 mutant mRNA may be degraded in Ets2em1/em1 embryos. To confirm this, we performed RT-PCR using E7.5 embryos. Embryos were separated into the posterior trophectoderm (TE) and anterior epiblast (Epi). Both regions expressed em1 mutant mRNA (Fig. 2B, and full-length gels are presented in Supplemental Fig S2), and their sequences included 205 nt deletions that were predicted from the genomic sequence. This result showed that em1 mutant RNA had escaped NMD. Furthermore, it is likely the case that the Ets2em1/em1 exhibited a distinct phenotype compared with Ets2db1/db1 even if the em1 mRNA was translated into a protein product (Supplemental Fig. S7).
Establishment of Ets2 homozygous mutant ESC lines and phenotypic analysis after birth
By rescuing placental function using the tetraploid complementation method, Ets2db1/db1 offspring were successfully developed to term. Therefore, we attempted the same experiment to define whether the embryonic lethal phenotype of Ets2em1/em1 was dependent on the placental deficiency or not.
Before conducting tetraploid complementation, we established Ets2em1/em1 and Ets2em2/em2 ESC lines. In this way, we improved the efficiency of obtaining homozygous mutant mice because the ratio of homozygous mutant embryos was only one out of four when we used embryos from a heterozygous crossing. After crossing heterozygotes, two-cell embryos were collected and developed until the blastocyst stage. ESC lines were established from the collected blastocysts and analyzed by genotyping. The rate of homozygous mutant ESC line establishment for both the Ets2em1/em1 and Ets2em2/em2 mutants followed Mendel’s law (Table 3).
Using the tetraploid complementation method, offspring were obtained from Ets2em1/em1 and Ets2em2/em2 ESC lines (Fig. 3A–C; Supplemental Fig. S3, and Table 4). This result indicated that the embryonic lethality observed for the Ets2em1/em1 and Ets2em2/em2 genotypes was due to a dysfunction of placental differentiation, the same as that seen for the Ets2db1/db1 mutant. Unexpectedly, wavy hair and curly whisker phenotypes were not observed in Ets2em1/em1 or Ets2em2/em2 mice (Fig. 3B,C; Supplemental Fig. S3).
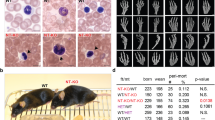
Generation of homozygous Ets2 mutant models using ESCs and assessment of hair and whisker phenotypes after birth. (A) Strategy for the generation of Ets2 homozygous mutant mouse after establishing the ESC from homozygous mutant blastocysts with tetraploid complementation. (B) 2-week-old Ets2em1/em1 mice. (C) Faces of 4-week-old in Ets2+/em1 (left picture) and Ets2em1/em1 (right picture). Curly whiskers were not observed in Ets2em1/em1. (D) Strategy for the generation of Ets2 homozygous null mutant mouse after genome-editing to the ESC with tetraploid complementation. (E) The hair of the Ets2null(em3/em3) mutant mouse was wavy (indicated as #). (F) Some whiskers of the null mutant mouse were not straight (arrow in the lower picture).
To corroborate the relationship between Ets2 and the wavy hair phenotype, we established a null mutant (hereafter referred to as Ets2null(em3/em3)) ES cell line, in which a region upstream of exon 2 through the 3′-UTR of exon 10 was deleted, including all open reading frame (ORF) regions (Supplemental Fig. S4). Pups were then generated using the tetraploid complementation method (Fig. 3D–F; Table 4). Both the wavy hair and curly whisker phenotypes were observed in Ets2null(em3/em3) mice from around 2-weeks of age, as was observed for Ets2db1/db1 mice.
Gene expression in newly established Ets2 mutant skins
In this study, newly established Ets2em1/em1 and Ets2em2/em2 mice exhibited an embryonic lethal phenotype due to placental dysfunction but did not exhibit the wavy hair phenotype of Ets2db1/db1 and Ets2null(em3/em3) mice, despite having a frame-shift mutation. Therefore, we next investigated the Ets2 gene expression from the Ets2 locus in their skins.
The expression of the mRNA was detected in the skin of wildtype, Ets2em1/em1 and Ets2em2/em2 mice, but not in the skin of Ets2null(em3/em3) mice. Notably, two sizes of fragments were detected in Ets2em1/em1 and Ets2em2/em2 skin samples (Fig. 4A, Supplemental Fig. S5A and full-length gels are presented in Supplemental Fig. S6).
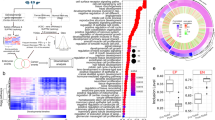
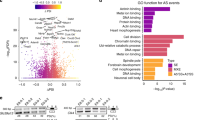
Gene expression in the Ets2em1/em1 skin. (A) Gene expression from the Ets2 locus in skins of WT, Ets2null(em3/em3) mutant (null) and Ets2em1/em1 (em1) mice. From em1 skin, two types of mRNA were expressed, although both were shorter than WT. The larger mRNA was called em1L, and the smaller mRNA was called em1S. (B) Sequences of em1L and em1S. Em1L was the expected sequence, but em1S contained a deleted locus that matched exon 8, shown in Supplemental Fig. S5. (C) Gene expression of Mmp3 and Mmp9 in the Ets2em1/em1 skin. There were significant differences in the expression level of Mmp3 between WT and null, em1 and null, and WT and em1. The expression level of Mmp9 in the null skin was slightly decreased compared with WT, but not significantly, and em1 showed a significant difference compared with null. *p < 0.05, **p < 0.01.
We hypothesized that the unexpected fragment size might be attributed to a splice variant and that this could explain the differences in phenotype between new Ets2 mutants and Ets2db1/db1 animals. Therefore, we next analyzed the sequences of the potential splice variants. Although the large bands observed for skins and thymuses of the Ets2em1/em1 and Ets2em2/em2 represented the expected frame-shifted sequences, the sequences of the smaller bands showed skipping of exon 8, which was in-frame and consisted of 264 bp (Fig. 4B; Supplemental Fig. S5B).
Further, we examined the gene expression of MMP-3 and MMP-9 in Ets2em1/em1 skins, since a previous report showed that expression of these genes was decreased in Ets2db1/db1 mice4. However, the expression levels of MMP-3 and MMP-9 in 4-week-old Ets2em1/em1 skins were not reduced compared with the wildtype, even though they were reduced in the Ets2null(em3/em3) (Fig. 4C).
Presence of Ets2 mutant protein in the skin and thymus
This Ets2 mutant protein, which skipped exon 8 (Ets2 ΔEx8), was predicted to contain the ETS domain based on the SMART online database14 (Supplemental Fig. S7). To confirm the presence of Ets2 mutant protein in the skin and thymus, we performed western blotting. Amino acid sequence sizes of Ets2, Ets2 em1, Ets2 em2, Ets2 ΔEx8 protein were of 468AA, 342AA, 298AA, and 380AA, respectively. Ets2 protein and Ets2 mutant proteins were distinguished by these molecular weights and the lack of band in the null mutant tissues.
The Ets2 em1 protein was detected at its expected size in Ets2em1/em1 skin and thymus. The Ets2 em2 protein was not found in both Ets2em2/em2 skin, even though Ets2 em2 mRNA (em2-L) was detected in the Ets2em2/em2 skin (Supplemental Fig. S5A and Supplemental Fig. S6B). Ets2 ΔEx8 protein, which has ETS domain, was present in the Ets2em1/em1 thymus, Ets2em2/em2 skin, but not clearly in the Ets2em1/em1 skin (Fig. 5, and full-length blots of Fig. 5 are presented in Supplemental Fig. S8).
Presence of proteins in the Ets2 mutant skin and thymus. Western blotting with anti-Ets2 antibody and anti-Gapdh antibody in the skin (A) and thymus (B). Native Ets2 protein (*) was detected only in WT samples. Ets2 ΔEx8 (**) and Ets2 em1 (***) mutant proteins were not detected in WT and null mutant samples. Ets2 em2 mutant protein (the predicted size indicates as ##) were not detected in Ets2em2/em2 (em2) skin.
Discussion
In this work, we attempted to generate Ets2 homozygous mutant models at F0 generation by combination with CRISPR/Cas9 system and tetraploid complementation. We could not obtain the Ets2 homozygous mutant model at F0 generation using zygotes but generated those using ES cells. In the previous study, the theoretical birth rate of Ets2db1/db1, which crossed with heterozygous parent and performed tetraploid complementation, was guessed 25% along with Mendelian law. However, it has been reported that the actual birth rate was 10%4. In this study, the birth rate of offspring from Ets2null(em3/em3) ES cells by tetraploid complementation was comparable to that from wild type ES cells as shown in Table 4. These results suggest that the mutant ES cells can be used to analyze gene functions after birth if the mutations were deleterious to the trophectoderm lineages.
Here, we newly generated two Ets2 frame-shift mutant models, namely, em1 and em2. Since both have the stop codon located before the ETS domain, we predicted that both mutant proteins would lack the ETS domain if they were translated. Therefore, we expected that the phenotypes in these homozygous mutants would mimic those described in previous reports for Ets2db1/db1 mice. Indeed, Ets2em1/em1 mice exhibited the same embryonic lethal phenotype, but the wavy hair and curly whisker phenotypes observed in Ets2db1/db1 and Ets2null(em3/em3) mice did not occur in Ets2em1/em1 and Ets2em2/em2 mice. Hair and whisker phenotypes of Ets2db1/db1 mice have been caused due to the lack of ETS domain4. We found that the skin and thymus of Ets2em1/em1 and Ets2em2/em2mice expressed a mutant mRNA lacking exon 8 that could potentially be translated into a protein, including the ETS domain. We detected Ets2 ΔEx8 protein in Ets2em2/em2 skin and Ets2em1/em1 and Ets2em2/em2 thymuses. These data demonstrated that Ets2 ΔEx8 mRNA was translated into the protein which contains the ETS domain. Therefore, we hypothesize that this Ets2 ΔEx8 protein could rescue the Ets2 function in hair and whisker. This was strongly suggested by the finding that these mice exhibited comparable levels of MMP-3 and MMP-9 mRNAs to wildtype, and Ets2db1/db1 and Ets2null(em3/em3) mice exhibited decreased expression.
DNA mutations in a genetic locus frequently lead to exon skipping, and several human diseases are linked to these types of exon skipping events. In addition, exon skipping can occur because of lack of exonic splicing enhancer sequences or an exonic splicing silencer sequence inside of an exon15,16. Some reports have also suggested that unexpected exon skips can occur when using the CRISPR/Cas9 system, indicating that a frame-shifted mutant exon induced by CRISPR/Cas9 is skipped but can be induced through alternative splicing or in-frame exon skipping17,18,19,20. These studies indicated that mutations located on the three multiple exons could induce its exon skipping. Due to our targeted exon 8 in Ets2 gene consists of 264 nts (88 multiples of three nts), exon skipping may have occurred in the tissue specifically.
Since there was no report about the mutation within exon 8 of Ets2 in human by ClinVar database21, the new Ets2 frame-shift mutant models could not applied to the human disease model. However, these new Ets2 frame-shift mutant models had the following issue.
The exon skip observed was only identified in the skin and thymus but not detected during embryonic stages. Thus, the induction of the exon skip was likely due to the changes in mRNA splicing that were dependent on the cell type or developmental stage, despite having the same genomic mutation. This phenomenon has a possible effect on the appearance of phenotypes making it a novel finding of this study.
Overall, our study suggested that all ORF-deletion models are adequate for analysis if we have to generate the gene knockouts. Furthermore, research related to indel mutation or point mutation in human disease models need to pay attention to the effects of alternative splicing.
Methods
Animals
All animal experiments were conducted in accordance with the guidelines of “Regulations and By-Laws of Animal Experimentation at the Nara Institute for Science and Technology”, and were approved by the Animal experimental Committee at the Nara Institute of Science and Technology (the approval no. 1639). Study of the animal experiments were carried out in compliance with the ARRIVE guidelines22. B6D2F1 female mice and ICR mice were purchased from SLC (Japan). C57BL/6J male mice were purchased from CLEA (Japan).
Collection of zygotes
Female mice were treated by PMSG and hCG for superovulation, then mated with male mice. Pronuclear stage zygotes were collected from female oviducts after 20 h of hCG injection. After removing cumulus cells using hyaluronidase, zygotes were incubated in KSOM at 37 °C under 5% CO2 in the air until use. 2-cell stage zygotes were collected from female oviducts after 42–46 h of hCG injection by the flush-out method. Collected 2-cell stage embryos were incubated until use the same as above.
Generation of Ets2 mutant zygote by CRISPR/Cas9 system using electroporation
Target sites of guide RNA (gRNA) were designed using the web tool CRISPR direct23. Genome editing by the electroporation was referred to as the previous study12.
CFB16-HB and LF501PT1-10 electrode (BEXCo.Ltd., Tokyo, Japan) were used for electroporation. 30–40 pronuclear stage zygotes were subjected to electroporation at one time. Zygotes were washed with Opti-MEM I (Thermofisher) three times, subsequently placed in a line in the electrode gap filled with five μl the mixture of 120 ng/μl Cas9 protein (TaKaRa, Japan), 300 ng/μl tracrRNA, and 200 ng/μl crRNA (HPLC grade, Fasmac) in Opti-MEM I. The electroporation condition was performed were 30 V (3 ms ON ± 97 ms OFF) four times. After electroporation, zygotes were washed with KSOM three times then cultured until developing the eight-cell stage. Eight-cell stage embryos were provided to the tetraploid complementation.
Establishment of Ets2 mutated ESC lines
To establish the Ets2 homozygous mutant model, collected 2-cell stage embryos from Ets2 heterozygous mutant parents were incubated until the blastocyst stage, removing the Zona pellucida (ZP) using Acidic Tyrode solution (Sigma T1788). Blastocyst embryos without the ZP were seed on gelatin-coated 60-mm dishes and cultured on mouse embryonic fibroblast (MEF) with N2B27 medium supplemented with 3 μM CHIR99021(Axon1386), 1.5 μM CGP77675 (Sigma SML0314), and mouse LIF (N2B27-a2i/L medium)24. After seven days, the outgrowth of blastocysts was disaggregated by 0.25% trypsin in 1 mM EDTA in PBS (−). Half of the cells were seeded on MEF with the gelatin-coated dishes for expanding. The others were seeded on the gelatin-coated dishes without MEF for genotyping by PCR. Ets2 homozygous mutant ESC lines were provided for tetraploid complementation.
The Ets2 null mutant model was established using ESCs, referred to as the previous study25. mF1-05 ESC line, which was newly established from 129X1and C57BL6/J F1 embryo, was seeded on MEF then transfected with two designed pSpCas9(BB)-2A-Puro (pX459) V2.0 (Addgene #62988) plasmids using Lipofectamine 3000 (Thermofisher). Transfected cells were selected by transient treatment with 1 μg/ml puromycin, then ESC colonies were subject to genotyping with PCR and sequencing. The Ets2 null mutant ESC line was provided for tetraploid complementation.
Tetraploid complementation
Tetraploid embryos were prepared as described previously9,26. In brief, ICR two-cell stage embryos were placed in the fusion buffer, and electrofusion was performed by applying 140 V for 50 ms after aligning embryos between the electrodes. CFB16-HB and LF501PT1-10 electrode (BEXCo.Ltd., Tokyo, Japan) were used for cell fusion.
A wild-type tetraploid four-cell embryo and a genome-edited diploid eight-cell embryo were aggregated after removing the Zonae pellucid for the aggregation method. For the injection method, Ets2 mutant ESCs were injected into a wild-type tetraploid four-cell embryo or blastocyst. These embryos were cultured until the blastocysts stage and transferred into the uterus of E2.5 pseudopregnant ICR mice. Offspring were recovered by natural delivery or Caesarean section on E19.5. The mutation of offspring was detected by genotyping with PCR and sequencing.
Genotyping
Genotyping primers for detecting Ets2-wild, em 1, and em 2 alleles were 5′-ctgagtttaagagtgctcggagg-3′ (Ets2_Fw) and 5′- gccctataggacttgtgtacagg-3′ (Ets2_Rev). Primers for Ets2 null (em3) mutant allele(s) were 5′-tgtggagtctcacatcgaag-3′ (Ets2_Ex2_F) and 5′- gggcctgctcggtgccacgg-3′ (Ets2_EX10_R). DNA fragments were amplified using GoTaq (Promega) for 40 cycles under the following conditions: 94 °C for 30 s, 60 °C for 30 s and 68 °C for 40 s for detecting wild, em1 or em2 allele, and 94 °C for 30 s, 60 °C for 30 s and 68 °C for 20 s for detecting the null allele, respectively.
RNA expression analysis
Mouse cDNAs were prepared from 4-week old skin, adult skin, and adult thymus using SuperScript III Reverse Transcriptase (Thermo Fisher Scientific) after purified RNA by Trizol reagent (Thermo Fisher Scientific). RT-PCR was performed using 20 ng of cDNA with the following primers: 5′-CGTGAATTTGCTCAACAACAATTCTG-3′ and 5′-GAGAGGCTATGCCGGT-3′ for Ets2, 5′-CCAGTATGACTCCACTCACG-3′ and 5′-GACTCCACGACATACTCAGC-3 for Gapdh 27. cDNA fragments were amplified using KOD Fx Neo (TOYOBO) or GoTaq (Promega) for 35 cycles under the following conditions: 94 °C for 30 s, 60 °C for 30 s and 72 °C for 40 s for Ets2, and 94 °C for 30 s, 53 °C for 30 s and 72 °C for 30 s for Gapdh.
Quantitative Real-time PCR was performed using 20 ng of cDNA with following primers: 5′-TTAAAGACAGGCACTTTTGG-3′ and 5′-CAGGGTGTGAATGCTTTTAG-3′for Mmp3, 5′-CGTCTGAGAATTGAATCAGC-3′ and 5′-AGTAGGGGCAACTGAATACC-3′ for Mmp9 expression5. Gene expression level was normalized by Gapdh, the same cDNA. The primer set for Gapdh was the same as above. Real-time PCR was performed by LightCycler96 (Roche) using the Luna Universal qPCR Master Mix (NEB), and the data were analyzed by the LightCycler96 software (Roche, version 1.1.0.1320).
Western blotting
Whole cell extracts of 2 weeks old skin or thymus were prepared by homogenization in RIPA buffer (Nacalai). Equivalent amount of protein were separated by 4–12% Bolt Bis–Tris gel (Invitrogen) and run under reducing conditions. After gel electrophoresis, they transferred to immobilon-P membrane (Merck). Membranes were blocked in 5% skim milk powder in phosphate-buffered saline with Tween20. The membrane was incubated with anti-Gapdh (WAKO, 010-25521) or anti-Ets2 (Genetex, GTX104527) followed by a secondary antibody conjugated with horseradish peroxidase (Cytiva, NA931, NA934). Peroxidase activity was detected with Chemi-Lumi One (Nacalai) using the FUSION -Chemiluminescence Imaging System (Vilber-Lourmat).
Statistics analysis
The statistical difference was determined using the Student t-test. Differences were considered statistically significant if the P-value was less than 0.05.
References
Karim, F. D. et al. The ETS-domain: A new DNA-binding motif that recognizes a purine-rich core DNA sequence. Genes Dev. 4, 1451–1453. https://doi.org/10.1101/gad.4.9.1451 (1990).
Sharrocks, A. D. The ETS-domain transcription factor family. Nat. Rev. Mol. Cell Biol. 2, 827–837. https://doi.org/10.1038/35099076 (2001).
Seidel, J. J. & Graves, B. J. An ERK2 docking site in the Pointed domain distinguishes a subset of ETS transcription factors. Genes Dev. 16, 127–137. https://doi.org/10.1101/gad.950902 (2002).
Yamamoto, H. et al. Defective trophoblast function in mice with a targeted mutation of Ets2. Genes Dev. 12, 1315–1326. https://doi.org/10.1101/gad.12.9.1315 (1998).
Man, A. K. et al. Ets2-dependent stromal regulation of mouse mammary tumors. Mol. Cell Biol. 23, 8614–8625. https://doi.org/10.1128/mcb.23.23.8614-8625.2003 (2003).
Wei, G. et al. Ets1 and Ets2 are required for endothelial cell survival during embryonic angiogenesis. Blood 114, 1123–1130. https://doi.org/10.1182/blood-2009-03-211391 (2009).
Baran, C. P. et al. Transcription factor ets-2 plays an important role in the pathogenesis of pulmonary fibrosis. Am. J. Respir. Cell Mol. Biol. 45, 999–1006. https://doi.org/10.1165/rcmb.2010-0490OC (2011).
Wei, G. et al. Activated Ets2 is required for persistent inflammatory responses in the motheaten viable model. J. Immunol. 173, 1374–1379. https://doi.org/10.4049/jimmunol.173.2.1374 (2004).
Okada, Y. et al. Complementation of placental defects and embryonic lethality by trophoblast-specific lentiviral gene transfer. Nat. Biotechnol. 25, 233–237. https://doi.org/10.1038/nbt1280 (2007).
Cong, L. et al. Multiplex genome engineering using CRISPR/Cas systems. Science 339, 819–823. https://doi.org/10.1126/science.1231143 (2013).
Mali, P. et al. RNA-guided human genome engineering via Cas9. Science 339, 823–826. https://doi.org/10.1126/science.1232033 (2013).
Hashimoto, M., Yamashita, Y. & Takemoto, T. Electroporation of Cas9 protein/sgRNA into early pronuclear zygotes generates non-mosaic mutants in the mouse. Dev. Biol. 418, 1–9. https://doi.org/10.1016/j.ydbio.2016.07.017 (2016).
Nagy, A., Rossant, J., Nagy, R., Abramow-Newerly, W. & Roder, J. C. Derivation of completely cell culture-derived mice from early-passage embryonic stem cells. Proc. Natl. Acad. Sci. U S A 90, 8424–8428, https://doi.org/10.1073/pnas.90.18.8424 (1993).
SMART. http://smart.embl-heidelberg.de/.
Cartegni, L., Chew, S. L. & Krainer, A. R. Listening to silence and understanding nonsense: Exonic mutations that affect splicing. Nat. Rev. Genet. 3, 285–298. https://doi.org/10.1038/nrg775 (2002).
Baralle, F. E. & Giudice, J. Alternative splicing as a regulator of development and tissue identity. Nat. Rev. Mol. Cell Biol. 18, 437–451. https://doi.org/10.1038/nrm.2017.27 (2017).
Mou, H. et al. CRISPR/Cas9-mediated genome editing induces exon skipping by alternative splicing or exon deletion. Genome Biol. 18, 108. https://doi.org/10.1186/s13059-017-1237-8 (2017).
Chen, D. et al. CRISPR/Cas9-mediated genome editing induces exon skipping by complete or stochastic altering splicing in the migratory locust. BMC Biotechnol. 18, 60. https://doi.org/10.1186/s12896-018-0465-7 (2018).
Sui, T. et al. CRISPR-induced exon skipping is dependent on premature termination codon mutations. Genome Biol. 19, 164. https://doi.org/10.1186/s13059-018-1532-z (2018).
Tuladhar, R. et al. CRISPR-Cas9-based mutagenesis frequently provokes on-target mRNA misregulation. Nat. Commun. 10, 4056. https://doi.org/10.1038/s41467-019-12028-5 (2019).
ClinVar. https://www.ncbi.nlm.nih.gov/clinvar/.
Kilkenny, C. et al. Improving bioscience research reporting: The ARRIVE guidelines for reporting animal research. PLoS Biol. 8, e1000412. https://doi.org/10.1371/journal.pbio.1000412 (2010).
CRISPRdirect. http://crispr.dbcls.jp/.
Choi, J. et al. Prolonged Mek1/2 suppression impairs the developmental potential of embryonic stem cells. Nature 548, 219–223. https://doi.org/10.1038/nature23274 (2017).
Oji, A. et al. CRISPR/Cas9 mediated genome editing in ES cells and its application for chimeric analysis in mice. Sci. Rep. 6, 31666. https://doi.org/10.1038/srep31666 (2016).
Kokubu, C. et al. A transposon-based chromosomal engineering method to survey a large cis-regulatory landscape in mice. Nat. Genet. 41, 946–952. https://doi.org/10.1038/ng.397 (2009).
Wen, F. et al. Ets2 is required for trophoblast stem cell self-renewal. Dev. Biol. 312, 284–299. https://doi.org/10.1016/j.ydbio.2007.09.024 (2007).
Acknowledgements
This work was supported by JSPS KAKENHI Grant Number 16K07091, 18H04885, Start Up Fund for female researchers in NAIST, and KAC 40th Anniversary Research Grant. The authors would like to thank Enago (https://www.enago.jp) for the English language review.
Author information
Authors and Affiliations
Contributions
A.I., Y.K., I.N., and W.H. performed most experiments, assisted by N.Y. who performed tetraploid complementation, and S.Y., performed qPCR. A.I., S.Y., and M.I. analyzed the data. A.I. wrote the manuscript and all authors discussed the results and commented on the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Kishimoto, Y., Nishiura, I., Hirata, W. et al. A novel tissue specific alternative splicing variant mitigates phenotypes in Ets2 frame-shift mutant models. Sci Rep 11, 8297 (2021). https://doi.org/10.1038/s41598-021-87751-5
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-021-87751-5
This article is cited by
-
Escaping from CRISPR–Cas-mediated knockout: the facts, mechanisms, and applications
Cellular & Molecular Biology Letters (2024)
-
Generating an organ-deficient animal model using a multi-targeted CRISPR-Cas9 system
Scientific Reports (2024)
-
Identification and characterization of novel ETV4 splice variants in prostate cancer
Scientific Reports (2023)
-
Scheduled simple production method of pseudopregnant female mice for embryo transfer using the luteinizing hormone-releasing hormone agonist
Scientific Reports (2022)
-
Genome editing
Scientific Reports (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.