Article
|
Open Access
Featured
-
-
Article
| Open Access Longshot enables accurate variant calling in diploid genomes from single-molecule long read sequencing
Longshot enables accurate variant calling in diploid genomes from single-molecule long read sequencing
Single-molecule sequencing (SMS) such as Pacific Biosciences and Oxford Nanopore generate long reads with high error rate. Here, the authors develop Longshot, a computational method that detects and phases single nucleotide variants (SNV) in diploid genomes using SMS data.
- Peter Edge
- & Vikas Bansal
-
Article
| Open Access A systematic evaluation of single cell RNA-seq analysis pipelines
A systematic evaluation of single cell RNA-seq analysis pipelines
There has been a rapid rise in single cell RNA-seq methods and associated pipelines. Here the authors use simulated data to systematically evaluate the performance of 3000 possible pipelines to derive recommendations for data processing and analysis of different types of scRNA-seq experiments.
- Beate Vieth
- , Swati Parekh
- & Ines Hellmann
-
Article
| Open Access DC3 is a method for deconvolution and coupled clustering from bulk and single-cell genomics data
DC3 is a method for deconvolution and coupled clustering from bulk and single-cell genomics data
Single-cell omics analysis can reveal heterogeneity among individual cells at different levels. Here, the authors develop DC3, a computational method for joint analysis of various bulk and single-cell data from the same heterogeneous cell population.
- Wanwen Zeng
- , Xi Chen
- & Wing Hung Wong
-
Article
| Open Access Thermodynamic control of −1 programmed ribosomal frameshifting
Thermodynamic control of −1 programmed ribosomal frameshifting
Programmed ribosomal frameshifting (PRF) is an alternative translation strategy that causes controlled slippage of the ribosome along the mRNA, changing the sequence of the synthesized protein. Here the authors provide a thermodynamic framework that explains how mRNA sequence determines the efficiency of frameshifting.
- Lars V. Bock
- , Neva Caliskan
- & Helmut Grubmüller
-
Article
| Open Access SCALE method for single-cell ATAC-seq analysis via latent feature extraction
SCALE method for single-cell ATAC-seq analysis via latent feature extraction
Single-cell ATAC-seq data is challenging to analyse for reasons such as high dimensionality and sparsity. Here, the authors develop SCALE, a deep learning method that leverages latent feature extraction for various tasks of scATACseq data analysis.
- Lei Xiong
- , Kui Xu
- & Qiangfeng Cliff Zhang
-
Article
| Open Access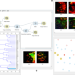 Democratized image analytics by visual programming through integration of deep models and small-scale machine learning
Democratized image analytics by visual programming through integration of deep models and small-scale machine learning
Deep learning approaches for image preprocessing and analysis offer important advantages, but these are rarely incorporated into user-friendly software. Here the authors present an easy-to-use visual programming toolbox integrating deep-learning and interactive data visualization for image analysis.
- Primož Godec
- , Matjaž Pančur
- & Blaž Zupan
-
Article
| Open Access Evolution and regulation of nitrogen flux through compartmentalized metabolic networks in a marine diatom
Evolution and regulation of nitrogen flux through compartmentalized metabolic networks in a marine diatom
Here, using the diatom Phaeodactylum tricornutum as a model organism, the authors combine functional genomics, phylogenetics, and metabolic modeling to describe how diatoms might have functionally integrated nitrogen metabolism during evolution and how metabolic flux is regulated across cellular compartments
- Sarah R. Smith
- , Chris L. Dupont
- & Andrew E. Allen
-
Article
| Open Access De novo identification of essential protein domains from CRISPR-Cas9 tiling-sgRNA knockout screens
De novo identification of essential protein domains from CRISPR-Cas9 tiling-sgRNA knockout screens
Tiling-sgRNA designs allow the in situ evaluation of protein domain functions. Here the authors present ProTiler - a computational method to predict CRISPR knockout hyper-sensitive regions, revealing previously unannotated domains.
- Wei He
- , Liang Zhang
- & Han Xu
-
Article
| Open Access The genome-wide multi-layered architecture of chromosome pairing in early Drosophila embryos
The genome-wide multi-layered architecture of chromosome pairing in early Drosophila embryos
Homologs are paired in Drosophila somatic cells from embryogenesis to adulthood. Using a computational approach for haplotype-resolved Hi-C, the authors reveal highly structured homolog pairing in Drosophila embryos during zygotic genome activation and demonstrate its application to mammalian embryos.
- Jelena Erceg
- , Jumana AlHaj Abed
- & C.-ting Wu
-
Article
| Open Access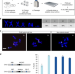 Highly structured homolog pairing reflects functional organization of the Drosophila genome
Highly structured homolog pairing reflects functional organization of the Drosophila genome
Trans-homolog interactions, such as homolog pairing, are highly structured and associated with gene function in Drosophila cells. Here, the authors use haplotype-resolved Hi-C to identify genome-wide trans-homolog interactions in a Drosophila hybrid cell line and investigate their patterns and functional roles.
- Jumana AlHaj Abed
- , Jelena Erceg
- & C.-ting Wu
-
Article
| Open Access A tunable dual-input system for on-demand dynamic gene expression regulation
A tunable dual-input system for on-demand dynamic gene expression regulation
Cellular systems have numerous mechanisms to control gene expression. Here the authors build a Tet-On system with conditional destablising elements to regulate gene expression and protein stability, allowing fine modulation of mESC signalling pathways.
- Elisa Pedone
- , Lorena Postiglione
- & Lucia Marucci
-
Article
| Open Access Functional interpretation of single cell similarity maps
Functional interpretation of single cell similarity maps
The increasing accessibility of single cell RNA sequencing demands tools that enable data visualization and interpretation. Here, the authors introduce Vision, a flexible annotation tool that operates directly on the manifold of cell-cell similarity and aids interpretation of cellular heterogeneity.
- David DeTomaso
- , Matthew G. Jones
- & Nir Yosef
-
Article
| Open Access Massive computational acceleration by using neural networks to emulate mechanism-based biological models
Massive computational acceleration by using neural networks to emulate mechanism-based biological models
Mechanistic models provide valuable insights, but large-scale simulations are computationally expensive. Here, the authors show that it is possible to explore the dynamics of a mechanistic model over a large set of parameters by training an artificial neural network on a smaller set of simulations.
- Shangying Wang
- , Kai Fan
- & Lingchong You
-
Article
| Open Access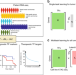 Chromatin-informed inference of transcriptional programs in gynecologic and basal breast cancers
Chromatin-informed inference of transcriptional programs in gynecologic and basal breast cancers
Epigenomic data on chromatin accessibility and transcription factor occupancy can reveal enhancer landscapes in cancer. Here, the authors develop a computational strategy called PSIONIC (patient-specific inference of networks informed by chromatin) to model the impact of enhancers on transcriptional programs in gynecologic and basal breast cancers.
- Hatice U. Osmanbeyoglu
- , Fumiko Shimizu
- & Christina S. Leslie
-
Article
| Open Access Ageing affects DNA methylation drift and transcriptional cell-to-cell variability in mouse muscle stem cells
Ageing affects DNA methylation drift and transcriptional cell-to-cell variability in mouse muscle stem cells
Age-related tissue alterations have been associated with a decline in stem cell number and function. Here the authors report a single cell multi-omics study of mouse muscle stem cells, combining single cell transcriptome and DNA methylome profiling and find that aged cells have a global increase of uncoordinated transcriptional heterogeneity biased towards genes regulating cell-niche interactions.
- Irene Hernando-Herraez
- , Brendan Evano
- & Wolf Reik
-
Article
| Open Access Simultaneous measurement of excitation-contraction coupling parameters identifies mechanisms underlying contractile responses of hiPSC-derived cardiomyocytes
Simultaneous measurement of excitation-contraction coupling parameters identifies mechanisms underlying contractile responses of hiPSC-derived cardiomyocytes
Cardiomyocytes obtained from human induced pluripotent stem cells are increasingly used for drug testing, but they are not always predictive of the heart contractile responses. Here the authors develop a method to measure cytosolic calcium, action potentials and contraction simultaneously, to achieve higher sensitivity for drug screenings.
- Berend J. van Meer
- , Ana Krotenberg
- & Christine L. Mummery
-
Article
| Open Access Genome-wide recombination map construction from single individuals using linked-read sequencing
Genome-wide recombination map construction from single individuals using linked-read sequencing
Variation of recombination rates within genomes has important implications in genetics and evolution. Here, the authors develop a method for building genome-wide recombination maps from single individuals using linked-read sequencing data, and report its application in mouse and stickleback fish.
- Andreea Dréau
- , Vrinda Venu
- & Felicity C. Jones
-
Article
| Open Access Linked-read sequencing of gametes allows efficient genome-wide analysis of meiotic recombination
Linked-read sequencing of gametes allows efficient genome-wide analysis of meiotic recombination
Meiotic crossovers (COs) generate genetic variation and ensure proper chromosome segregation. Here, the authors develop a method for identifying COs at kilobase resolution in pooled recombinants using linked-read sequencing data, and apply it to investigate genome-wide CO landscapes of Arabidopsis thaliana.
- Hequan Sun
- , Beth A. Rowan
- & Korbinian Schneeberger
-
Article
| Open Access ImmGen report: sexual dimorphism in the immune system transcriptome
ImmGen report: sexual dimorphism in the immune system transcriptome
Sexual dimorphism is observed frequently in immune disorders, but the underlying insights are still unclear. Here the authors analyze transcriptome and epigenome changes induced by interferon in various mouse immune cell types, and find only a restricted set of sexual dimorphism genes in innate immunity and macrophages.
- Shani Talia Gal-Oz
- , Barbara Maier
- & Tal Shay
-
Article
| Open Access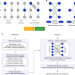 Discovering genetic interactions bridging pathways in genome-wide association studies
Discovering genetic interactions bridging pathways in genome-wide association studies
Genetic interactions may contribute to phenotypic traits but are challenging to decipher. Here, the authors develop BridGE, a computational approach for identifying pathways connected by genetic interactions from GWAS data.
- Gang Fang
- , Wen Wang
- & Chad L. Myers
-
Article
| Open Access Enhancing and shaping the immunogenicity of native-like HIV-1 envelope trimers with a two-component protein nanoparticle
Enhancing and shaping the immunogenicity of native-like HIV-1 envelope trimers with a two-component protein nanoparticle
Nanoparticles are a promising approach to increase immunogenicity of protein antigens for vaccines. Here, Brouwer et al. design self-assembling, two-component protein NPs that present native-like SOSIP trimers of HIV envelope protein and determine immunogenicity in a small animal model.
- Philip J. M. Brouwer
- , Aleksandar Antanasijevic
- & Rogier W. Sanders
-
Article
| Open Access An African Salmonella Typhimurium ST313 sublineage with extensive drug-resistance and signatures of host adaptation
An African Salmonella Typhimurium ST313 sublineage with extensive drug-resistance and signatures of host adaptation
Invasive non-typhoidal Salmonella (iNTS) infections are dominated by antibiotic resistant isolates of the sequence type (ST) 313. Here, the authors identify the ST313 sublineage II.1 in the Democratic Republic of the Congo exhibiting extensive drug resistance and genetic signatures potentially associated with host adaptation.
- Sandra Van Puyvelde
- , Derek Pickard
- & Stijn Deborggraeve
-
Article
| Open Access Optimized CRISPR guide RNA design for two high-fidelity Cas9 variants by deep learning
Optimized CRISPR guide RNA design for two high-fidelity Cas9 variants by deep learning
Application of highly specific Cas9 variants can be restricted by the design of the guide RNA. Here the authors present DeepHF, a gRNA activity prediction tool built from genome-scale screens of 50,000 guides covering 20,000 genes.
- Daqi Wang
- , Chengdong Zhang
- & Yongming Wang
-
Article
| Open Access Identification of significant chromatin contacts from HiChIP data by FitHiChIP
Identification of significant chromatin contacts from HiChIP data by FitHiChIP
HiChIP/PLAC-seq assay is popular for profiling 3D genome interactions among regulatory elements at kilobase resolution. Here the authors describe FitHiChIP an empirical null-based, flexible computational method for statistical significance estimation and loop calling from HiChIP data.
- Sourya Bhattacharyya
- , Vivek Chandra
- & Ferhat Ay
-
Article
| Open Access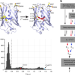 Learning the pattern of epistasis linking genotype and phenotype in a protein
Learning the pattern of epistasis linking genotype and phenotype in a protein
Epistasis underlies the complexity of genotype-phenotype maps. Here, the authors analyze 8,192 mutants that link two phenotypically distinct variants of the Entacmaea quadricolor fluorescent protein, and show the existence, but also the sparsity, of high-order epistatic interactions.
- Frank J. Poelwijk
- , Michael Socolich
- & Rama Ranganathan
-
Article
| Open Access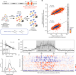 The mutational landscape of a prion-like domain
The mutational landscape of a prion-like domain
TDP43 aggregates are a hallmark of amyotrophic lateral sclerosis. By using deep mutagenesis to measure the toxicity of more than 50,000 mutations in the prion domain of TDP43, the authors conclude that mutations that increase toxicity promote formation of liquid-like condensates, while aggregation of TDP43 is protective for the cell.
- Benedetta Bolognesi
- , Andre J. Faure
- & Ben Lehner
-
Article
| Open Access UPF1/SMG7-dependent microRNA-mediated gene regulation
UPF1/SMG7-dependent microRNA-mediated gene regulation
UPF1 mediates the decay of target mRNA in a 3′ untranslated region (UTR)-length-dependent manner. Here the authors reveal that the 3′UTR-length-dependent regulation of UPF1-dependent mRNA decay occurs through EJC-independent but miRNA-dependent regulation.
- Jungyun Park
- , Jwa-Won Seo
- & Jin-Wu Nam
-
Article
| Open Access The distinction of CPR bacteria from other bacteria based on protein family content
The distinction of CPR bacteria from other bacteria based on protein family content
Recent studies have identified a large, phylogenetically distinct clade of bacteria, the candidate phyla radiation (CPR). Here, Méheust and colleagues analyze almost 3600 genomes to characterize the protein family content of CPR versus other bacteria and archaea.
- Raphaël Méheust
- , David Burstein
- & Jillian F. Banfield
-
Article
| Open Access RuvC uses dynamic probing of the Holliday junction to achieve sequence specificity and efficient resolution
RuvC uses dynamic probing of the Holliday junction to achieve sequence specificity and efficient resolution
Holliday junctions (HJs) are four-way DNA structures that occur in DNA repair by homologous recombination which are removed by nucleases such as RuvC. Here the authors provide structural and biochemical details on the steps required for the completion of the process.
- Karolina Maria Górecka
- , Miroslav Krepl
- & Marcin Nowotny
-
Article
| Open Access Exploring use of unsupervised clustering to associate signaling profiles of GPCR ligands to clinical response
Exploring use of unsupervised clustering to associate signaling profiles of GPCR ligands to clinical response
Identifying ligands which activate the specific effectors driving particular in vivo drug effects remains challenging. Here, the authors apply unsupervised clustering of pharmacodynamic parameters to classify GPCR ligands into different categories with similar signaling profiles and shared frequency of report of side effects.
- Besma Benredjem
- , Jonathan Gallion
- & Graciela Pineyro
-
Article
| Open Access Accurate detection of m6A RNA modifications in native RNA sequences
Accurate detection of m6A RNA modifications in native RNA sequences
We currently lack generic methods to map RNA modifications across the entire transcriptome. Here, the authors demonstrate that m6A RNA modifications can be detected with high accuracy using nanopore direct RNA sequencing.
- Huanle Liu
- , Oguzhan Begik
- & Eva Maria Novoa
-
Article
| Open Access Components of genetic associations across 2,138 phenotypes in the UK Biobank highlight adipocyte biology
Components of genetic associations across 2,138 phenotypes in the UK Biobank highlight adipocyte biology
While many pleiotropic genetic loci have been identified, how they contribute to phenotypes across traits and diseases is unclear. Here, the authors propose decomposition of genetic associations (DeGAs), which uses singular value decomposition, to characterize the underlying latent structure of genetic associations of 2,138 phenotypes.
- Yosuke Tanigawa
- , Jiehan Li
- & Manuel A. Rivas
-
Article
| Open Access Spatially clustered loci with multiple enhancers are frequent targets of HIV-1 integration
Spatially clustered loci with multiple enhancers are frequent targets of HIV-1 integration
HIV-1 usually targets active genes and integrates near the nuclear pore compartment. Here the authors show that recurrently targeted genes are proximal to super-enhancer genomic elements, which cluster in specific spatial compartments of the T cell nucleus, suggesting a role for nuclear organisation in viral infection.
- Bojana Lucic
- , Heng-Chang Chen
- & Marina Lusic
-
Article
| Open Access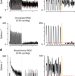 Neural sensitization improves encoding fidelity in the primate retina
Neural sensitization improves encoding fidelity in the primate retina
Light intensity on the retina can fluctuate rapidly during natural vision, posing a challenge for encoding visual information. Here, the authors report that mechanisms of sensitization/facilitation maintain the sensitivity of the numerically dominant neural pathway in the primate retina during dynamic vision.
- Todd R. Appleby
- & Michael B. Manookin
-
Article
| Open Access Translational coupling via termination-reinitiation in archaea and bacteria
Translational coupling via termination-reinitiation in archaea and bacteria
Archaea and bacteria often have gene pairs with overlapping stop and start codons, suggesting translational coupling. Here, Huber et al. analyse overlapping gene pairs from 720 genomes, and validate translational coupling via termination-reinitiation for 14 gene pairs in Haloferax volcanii and Escherichia coli.
- Madeleine Huber
- , Guilhem Faure
- & Jörg Soppa
-
Article
| Open Access Deep learning extends de novo protein modelling coverage of genomes using iteratively predicted structural constraints
Deep learning extends de novo protein modelling coverage of genomes using iteratively predicted structural constraints
Prediction of protein structures on the scale of genomes remains a challenge. Here the authors introduce a protein structure prediction method that uses deep learning to predict inter-atomic distances, torsion angles and hydrogen bonds, and apply it to predict the structures of 1475 Pfam domains.
- Joe G. Greener
- , Shaun M. Kandathil
- & David T. Jones
-
Comment
| Open Access Technology to advance infectious disease forecasting for outbreak management
Technology to advance infectious disease forecasting for outbreak management
Forecasting is beginning to be integrated into decision-making processes for infectious disease outbreak response. We discuss how technologies could accelerate the adoption of forecasting among public health practitioners, improve epidemic management, save lives, and reduce the economic impact of outbreaks.
- Dylan B. George
- , Wendy Taylor
- & Nicholas G. Reich
-
Article
| Open Access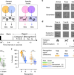 A dual role of prestimulus spontaneous neural activity in visual object recognition
A dual role of prestimulus spontaneous neural activity in visual object recognition
The effect of spontaneous variations in prestimulus neural activity on subsequent perception is incompletely understood. Here, using MEG, the authors identify two distinct neural processes that can influence object recognition in different ways.
- Ella Podvalny
- , Matthew W. Flounders
- & Biyu J. He
-
Article
| Open Access Single-cell transcriptomics reveals multi-step adaptations to endocrine therapy
Single-cell transcriptomics reveals multi-step adaptations to endocrine therapy
The development of resistance to endocrine therapy is a significant, clinical problem in breast cancer. Here, the authors identify a rare subpopulation of cells that drive resistance following transcriptional reprogramming.
- Sung Pil Hong
- , Thalia E. Chan
- & Luca Magnani
-
Article
| Open Access Changes in gene expression predictably shift and switch genetic interactions
Changes in gene expression predictably shift and switch genetic interactions
Non-additive genetic interactions are plastic and can complicate genetic prediction. Here, using deep mutagenesis of the lambda repressor, Li et al. reveal that changes in gene expression can alter the strength and direction of genetic interactions between mutations in many genes and develop mathematical models for predicting them.
- Xianghua Li
- , Jasna Lalić
- & Ben Lehner
-
Article
| Open Access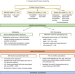 A phenotypic and genomics approach in a multi-ethnic cohort to subtype systemic lupus erythematosus
A phenotypic and genomics approach in a multi-ethnic cohort to subtype systemic lupus erythematosus
Systemic lupus erythematosus (SLE) is an autoimmune disease of substantial phenotypic heterogeneity in different ethnic groups. Here, using data from a multi-ethnic cohort, the authors describe an approach based on clinical and molecular data to subtype SLE patients into three clusters of severity.
- Cristina M. Lanata
- , Ishan Paranjpe
- & Lindsey A. Criswell
-
Article
| Open Access Identification of somatic mutations in single cell DNA-seq using a spatial model of allelic imbalance
Identification of somatic mutations in single cell DNA-seq using a spatial model of allelic imbalance
Single cell whole-genome sequencing data harbors information about somatic genetic variation but is challenging to analyze. Here, the authors develop a spatial model to correct for allelic amplification imbalance and a somatic SNV genotyper SCAN-SNV for analyzing single cell DNA sequencing data.
- Lovelace J. Luquette
- , Craig L. Bohrson
- & Peter J. Park
-
Article
| Open Access Integrative transcriptome imputation reveals tissue-specific and shared biological mechanisms mediating susceptibility to complex traits
Integrative transcriptome imputation reveals tissue-specific and shared biological mechanisms mediating susceptibility to complex traits
PrediXcan is a widely used gene expression imputation method that links genetic variants to gene expression. Here, the authors develop EpiXcan which leverages epigenetic annotations to inform transcriptomic imputation and further use the obtained gene-trait associations for computational drug repurposing.
- Wen Zhang
- , Georgios Voloudakis
- & Panos Roussos
-
Article
| Open Access Deep multiomics profiling of brain tumors identifies signaling networks downstream of cancer driver genes
Deep multiomics profiling of brain tumors identifies signaling networks downstream of cancer driver genes
Multi-omic profiling is a powerful approach to dissecting molecular mechanisms in disease. Here the authors generate whole proteome, phosphoproteome and transcriptome profiles from two mouse models of high-grade glioma driven by different oncogenes, and validate identified master regulators with a CRISPR screen.
- Hong Wang
- , Alexander K. Diaz
- & Junmin Peng
-
Article
| Open Access Cancer-associated mutations in DICER1 RNase IIIa and IIIb domains exert similar effects on miRNA biogenesis
Cancer-associated mutations in DICER1 RNase IIIa and IIIb domains exert similar effects on miRNA biogenesis
DICER is involved in the processing of miRNAs, where the RNase IIIa and IIIb domains are thought to cut the 3p and 5p hairpin arms, respectively. Here, in endometrial cancer, the authors identify an RNase IIIa mutation, which phenocopies mutations in the RNase IIIb domain.
- Jeffrey Vedanayagam
- , Walid K. Chatila
- & Eric C. Lai
-
Article
| Open Access A general approach for detecting expressed mutations in AML cells using single cell RNA-sequencing
A general approach for detecting expressed mutations in AML cells using single cell RNA-sequencing
The advent of single-cell RNA sequencing has revealed significant transcriptional heterogeneity in cancer, but its relationship to genomic heterogeneity remains unclear. Focusing on acute myeloid leukemia samples, the authors describe a general approach for linking mutation-containing cells to their transcriptional phenotypes using single-cell RNA sequencing data.
- Allegra A. Petti
- , Stephen R. Williams
- & Timothy J. Ley
-
Article
| Open Access Smu1 and RED are required for activation of spliceosomal B complexes assembled on short introns
Smu1 and RED are required for activation of spliceosomal B complexes assembled on short introns
Human spliceosome components Smu1 and RED regulate alternative splicing. Here the authors show that Smu1 and RED are also required for constitutive splicing of short introns.
- Sandra Keiper
- , Panagiotis Papasaikas
- & Reinhard Lührmann
-
Article
| Open Access The pause-initiation limit restricts transcription activation in human cells
The pause-initiation limit restricts transcription activation in human cells
Gene activation requires an increase of successful initiation events. Here, by employing a genome-wide kinetic analysis of transcription, the authors showed that gene activation generally requires a decrease in RNA Polymerase II (Pol II) promoter-proximal pausing while transcription of enhancer elements is not limited by Pol II pausing.
- Saskia Gressel
- , Björn Schwalb
- & Patrick Cramer
-
Article
| Open Access Past–future information bottleneck for sampling molecular reaction coordinate simultaneously with thermodynamics and kinetics
Past–future information bottleneck for sampling molecular reaction coordinate simultaneously with thermodynamics and kinetics
Efficient sampling of rare events in all-atom molecular dynamics simulations remains a challenge. Here, the authors adapt the Predictive Information Bottleneck framework to sample biomolecular structure and dynamics through iterative rounds of biased simulations and deep learning.
- Yihang Wang
- , João Marcelo Lamim Ribeiro
- & Pratyush Tiwary
Browse broader subjects
Browse narrower subjects
- Biochemical reaction networks
- Cellular signalling networks
- Classification and taxonomy
- Communication and replication
- Computational models
- Computational neuroscience
- Computational platforms and environments
- Data acquisition
- Data integration
- Data mining
- Data processing
- Data publication and archiving
- Databases
- Functional clustering
- Gene ontology
- Gene regulatory networks
- Genome informatics
- Hardware and infrastructure
- High-throughput screening
- Image processing
- Literature mining
- Machine learning
- Microarrays
- Network topology
- Phylogeny
- Power law
- Predictive medicine
- Probabilistic data networks
- Programming language
- Protein analysis
- Protein design
- Protein folding
- Protein function predictions
- Protein structure predictions
- Proteome informatics
- Quality control
- Scale invariance
- Sequence annotation
- Software
- Standards
- Statistical methods
- Virtual drug screening

 Species abundance information improves sequence taxonomy classification accuracy
Species abundance information improves sequence taxonomy classification accuracy