Featured
-
-
Article
| Open Access DynamicBind: predicting ligand-specific protein-ligand complex structure with a deep equivariant generative model
DynamicBind: predicting ligand-specific protein-ligand complex structure with a deep equivariant generative model
Proteins often function by changing conformations upon ligand binding. Efficient structural modelling of these interactions, crucial for drug discovery, is limited: here the authors address this with DynamicBind, a diffusion-based deep generative model.
- Wei Lu
- , Jixian Zhang
- & Shuangjia Zheng
-
Article
| Open Access ZeroBind: a protein-specific zero-shot predictor with subgraph matching for drug-target interactions
ZeroBind: a protein-specific zero-shot predictor with subgraph matching for drug-target interactions
Existing drug-target interaction (DTI) prediction methods generally fail to generalize well unseen proteins and drugs. Here the authors report a protein-specific meta-learning framework, ZeroBind, with subgraph matching for predicting protein-drug interactions from their structures.
- Yuxuan Wang
- , Ying Xia
- & Xiaoyong Pan
-
Article
| Open Access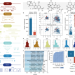 First fully-automated AI/ML virtual screening cascade implemented at a drug discovery centre in Africa
First fully-automated AI/ML virtual screening cascade implemented at a drug discovery centre in Africa
Streamlined data-driven drug discovery remains challenging, especially in resource-limited settings. Here, the authors present ZairaChem, an AI/ML tool that streamlines QSAR/QSPR modelling, implemented for the first time at the H3D Centre in South Africa.
- Gemma Turon
- , Jason Hlozek
- & Miquel Duran-Frigola
-
Article
| Open Access Transfer Learning with Kernel Methods
Transfer Learning with Kernel Methods
Transfer learning can be applied in computer vision and natural language processing to utilize knowledge from a source task to improve performance on a target task. The authors propose a framework for transfer learning with kernel methods for improved image classification and virtual drug screening.
- Adityanarayanan Radhakrishnan
- , Max Ruiz Luyten
- & Caroline Uhler
-
Article
| Open Access Sequence-based drug design as a concept in computational drug design
Sequence-based drug design as a concept in computational drug design
Conventional structure-based drug design pipeline is a complex, human-engineered process with multiple independently optimized steps. Here, the authors report a sequence-to-drug concept that discovers drug-like small molecule modulators directly from protein sequences.
- Lifan Chen
- , Zisheng Fan
- & Mingyue Zheng
-
Article
| Open Access Multi-omic underpinnings of epigenetic aging and human longevity
Multi-omic underpinnings of epigenetic aging and human longevity
Here, the authors integrate genomic, bulk and single-cell transcriptomic, and metabolomic data sets to compare the biological underpinning of four epigenetic clocks and human longevity, offering novel insights into aging biology.
- Lucas A. Mavromatis
- , Daniel B. Rosoff
- & Falk W. Lohoff
-
Article
| Open Access Computational pharmacogenomic screen identifies drugs that potentiate the anti-breast cancer activity of statins
Computational pharmacogenomic screen identifies drugs that potentiate the anti-breast cancer activity of statins
Statins are promising for breast cancer therapy; dipyridamole can potentiate their effects, but is contraindicated in some cases. Here, the authors develop a pharmacogenomics pipeline to predict other compounds that potentiate statins, and validate the top candidates in cell line screens and 3D cultures.
- Jenna E. van Leeuwen
- , Wail Ba-Alawi
- & Deena M. A. Gendoo
-
Article
| Open Access Integrating gene expression and clinical data to identify drug repurposing candidates for hyperlipidemia and hypertension
Integrating gene expression and clinical data to identify drug repurposing candidates for hyperlipidemia and hypertension
Prioritizing drug repurposing candidates for downstream studies remains challenging. Here, the authors present a high-throughput approach to identify and validate drug repurposing candidates, integrating human gene expression, drug perturbation, and clinical data from publicly available resources.
- Patrick Wu
- , QiPing Feng
- & Wei-Qi Wei
-
Article
| Open Access Sliding of HIV-1 reverse transcriptase over DNA creates a transient P pocket – targeting P-pocket by fragment screening
Sliding of HIV-1 reverse transcriptase over DNA creates a transient P pocket – targeting P-pocket by fragment screening
Here the authors observe a transient P-pocket created when HIV reverse transcriptase slides over DNA substrate, identify fragments targeting this pocket, and develop a cryo-EM platform for lead optimization.
- Abhimanyu K. Singh
- , Sergio E. Martinez
- & Kalyan Das
-
Article
| Open Access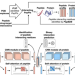 A deep-learning framework for multi-level peptide–protein interaction prediction
A deep-learning framework for multi-level peptide–protein interaction prediction
Peptide-protein interactions play fundamental roles in cellular processes and are crucial for designing peptide therapeutics. Here, the authors present a deep learning framework for simultaneously predicting peptide-protein interactions and identifying peptide binding residues involved in the interactions.
- Yipin Lei
- , Shuya Li
- & Jianyang Zeng
-
Article
| Open Access Causal network models of SARS-CoV-2 expression and aging to identify candidates for drug repurposing
Causal network models of SARS-CoV-2 expression and aging to identify candidates for drug repurposing
Given the severity of the SARS-CoV-2 pandemic, a major challenge is to rapidly repurpose existing approved drugs for clinical interventions. Here, the authors identify robust druggable protein targets within a principled causal framework that makes use of multiple data modalities and integrates aging signatures.
- Anastasiya Belyaeva
- , Louis Cammarata
- & Caroline Uhler
-
Article
| Open Access Computationally predicting clinical drug combination efficacy with cancer cell line screens and independent drug action
Computationally predicting clinical drug combination efficacy with cancer cell line screens and independent drug action
Computational models that can predict drug combination efficacy are often based on drug synergy. Here, the authors develop a different approach to computationally predict the efficacy of drug combinations using monotherapy data from high-throughput cancer cell line screens.
- Alexander Ling
- & R. Stephanie Huang
-
Article
| Open Access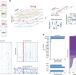 Genomic data integration by WON-PARAFAC identifies interpretable factors for predicting drug-sensitivity in vivo
Genomic data integration by WON-PARAFAC identifies interpretable factors for predicting drug-sensitivity in vivo
Integrative analyses that link molecular data to treatment sensitivity are essential for precision medicine. Here the authors introduce WON-PARAFAC to integrate multiple genomics data to identify sparse and interpretable factors.
- Yongsoo Kim
- , Tycho Bismeijer
- & Daniel J. Vis
-
Article
| Open Access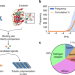 Pocket similarity identifies selective estrogen receptor modulators as microtubule modulators at the taxane site
Pocket similarity identifies selective estrogen receptor modulators as microtubule modulators at the taxane site
Taxanes are natural products which bind beta-tubulin, stabilize microtubules and have a broad spectrum of anticancer activity. Here authors employ a computational binding site similarity screen and cell-based assays to reveal a SERM cross-reactivity between the estrogen receptor and the beta-tubulin taxane binding pocket.
- Yu-Chen Lo
- , Olga Cormier
- & Russ B. Altman
-
Article
| Open Access A network integration approach for drug-target interaction prediction and computational drug repositioning from heterogeneous information
A network integration approach for drug-target interaction prediction and computational drug repositioning from heterogeneous information
Network-based data integration for drug–target prediction is a promising avenue for drug repositioning, but performance is wanting. Here, the authors introduce DTINet, whose performance is enhanced in the face of noisy, incomplete and high-dimensional biological data by learning low-dimensional vector representations.
- Yunan Luo
- , Xinbin Zhao
- & Jianyang Zeng
-
Article
| Open Access Reversal of cancer gene expression correlates with drug efficacy and reveals therapeutic targets
Reversal of cancer gene expression correlates with drug efficacy and reveals therapeutic targets
Increased availability of large-scale molecular profiling has enabled system-level monitoring of molecular effects of candidate therapeutics. Here, the authors take advantage of such data to show that the ability of a drug to reverse cancer-associated gene expression changes is indicative of itsin vitroanti-proliferative efficacy, allowing them to identify novel compounds against liver cancer.
- Bin Chen
- , Li Ma
- & Atul J. Butte
-
Article
| Open Access A new inhibitor of the β-arrestin/AP2 endocytic complex reveals interplay between GPCR internalization and signalling
A new inhibitor of the β-arrestin/AP2 endocytic complex reveals interplay between GPCR internalization and signalling
Beta-arrestins play central roles in the mechanisms regulating GPCR signalling and trafficking. Here the authors identify a selective inhibitor of the interaction between β-arrestin and the β2-adaptin subunit of the clathrin adaptor protein AP-2, which they use to dissect the role of the β-arrestin/β2-adaptin interaction in GPCR signalling.
- Alexandre Beautrait
- , Justine S. Paradis
- & Michel Bouvier

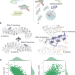 Design of target specific peptide inhibitors using generative deep learning and molecular dynamics simulations
Design of target specific peptide inhibitors using generative deep learning and molecular dynamics simulations