Article
|
Open Access
Featured
-
-
Article
| Open Access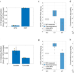 Inferring mitochondrial and cytosolic metabolism by coupling isotope tracing and deconvolution
Inferring mitochondrial and cytosolic metabolism by coupling isotope tracing and deconvolution
Studying metabolism in distinct subcellular compartments typically involves isolating organelles. Here, the authors demonstrate a quantitative approach to infer cytosolic and mitochondrial metabolic activities based on experiments with intact cells, maintaining physiological conditions.
- Alon Stern
- , Mariam Fokra
- & Tomer Shlomi
-
Article
| Open Access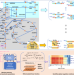 Local flux coordination and global gene expression regulation in metabolic modeling
Local flux coordination and global gene expression regulation in metabolic modeling
Genome-scale metabolic networks (GSMs) are a representation of a cell’s stoichiometrically balanced reactions. Here the authors report Decrem, a GSM reconstruction method, by integrating locally coupled reactions and global transcriptional regulation of metabolism by cell state.
- Gaoyang Li
- , Li Liu
- & Huansheng Cao
-
Article
| Open Access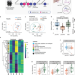 Portosystemic shunt placement reveals blood signatures for the development of hepatic encephalopathy through mass spectrometry
Portosystemic shunt placement reveals blood signatures for the development of hepatic encephalopathy through mass spectrometry
Patients with liver disease undergoing transjugular intrahepatic portosystemic shunt (TIPS) are at a higher risk of hepatic encephalopathy (HE). Here, the authors show intrahepatic shunting and specific metabolites, especially bile acids, as potential biomarkers and treatment targets for HE.
- Ana Carolina Dantas Machado
- , Stephany Flores Ramos
- & Amir Zarrinpar
-
Article
| Open Access Identification of gene function based on models capturing natural variability of Arabidopsis thaliana lipid metabolism
Identification of gene function based on models capturing natural variability of Arabidopsis thaliana lipid metabolism
The use of automated tools to reconstruct lipid metabolic pathways is not warranted in plants. Here, the authors construct Plant Lipid Module for Arabidopsis rosette using constraint-based modeling, demonstrate its integration in other plant metabolic models, and use it to dissect the genetic architecture of lipid metabolism.
- Sandra Correa Córdoba
- , Hao Tong
- & Zoran Nikoloski
-
Article
| Open Access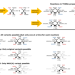 Network-wide thermodynamic constraints shape NAD(P)H cofactor specificity of biochemical reactions
Network-wide thermodynamic constraints shape NAD(P)H cofactor specificity of biochemical reactions
NADH and NADPH are redox cofactors coexisting in all living cells. Here, the authors present a computational study suggesting that evolved NAD(P)H reaction specificities in E. coli are largely shaped by metabolic network structure enabling maximal thermodynamic driving forces close to the theoretical optimum.
- Pavlos Stephanos Bekiaris
- & Steffen Klamt
-
Article
| Open Access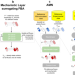 A neural-mechanistic hybrid approach improving the predictive power of genome-scale metabolic models
A neural-mechanistic hybrid approach improving the predictive power of genome-scale metabolic models
Mechanistic models estimate the phenotype of microorganisms in different environments but may have limited predictive capabilities. Here, authors develop trainable hybrid models with improved predictability using mechanistic insights and smaller training sets than conventional machine learning techniques.
- Léon Faure
- , Bastien Mollet
- & Jean-Loup Faulon
-
Article
| Open Access Genome-scale metabolic modeling of Aspergillus fumigatus strains reveals growth dependencies on the lung microbiome
Genome-scale metabolic modeling of Aspergillus fumigatus strains reveals growth dependencies on the lung microbiome
Here, the authors generate strain-specific genome-scale metabolic models of Aspergillus fumigatus and analyze fungal metabolism of infection of the lung of cystic fibrosis patients, finding that the fungus shapes the lung microbiome to promote its own growth.
- Mohammad H. Mirhakkak
- , Xiuqiang Chen
- & Gianni Panagiotou
-
Article
| Open Access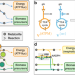 Functional decomposition of metabolism allows a system-level quantification of fluxes and protein allocation towards specific metabolic functions
Functional decomposition of metabolism allows a system-level quantification of fluxes and protein allocation towards specific metabolic functions
Quantifying the contribution of individual molecular components to complex cellular processes is a grand challenge in systems biology. Here, the authors present a general theoretical framework (Functional Decomposition of Metabolism, FDM) to quantify the contribution of every metabolic reaction to metabolic functions, e.g. the synthesis of biomass building blocks.
- Matteo Mori
- , Chuankai Cheng
- & Terence Hwa
-
Article
| Open Access Optimal enzyme utilization suggests that concentrations and thermodynamics determine binding mechanisms and enzyme saturations
Optimal enzyme utilization suggests that concentrations and thermodynamics determine binding mechanisms and enzyme saturations
One of the main challenges hampering the development of kinetic models is the lack of kinetic parameters for many enzymatic reactions. Here, the authors introduce a framework to explore the catalytically optimal operating conditions of any complex enzyme mechanism from an evolutionary perspective.
- Asli Sahin
- , Daniel R. Weilandt
- & Vassily Hatzimanikatis
-
Article
| Open Access Teasing out missing reactions in genome-scale metabolic networks through hypergraph learning
Teasing out missing reactions in genome-scale metabolic networks through hypergraph learning
A computational method for rapid and accurate gap-filling of metabolic networks without using phenotypic data is unavailable. Here, the authors address this problem by developing a deep learning based method that can predict missing reactions using topological features of the metabolic networks.
- Can Chen
- , Chen Liao
- & Yang-Yu Liu
-
Article
| Open Access Universal structures for adaptation in biochemical reaction networks
Universal structures for adaptation in biochemical reaction networks
At the molecular level, the evolution of life is driven by the generation and diversification of adaptation mechanisms. Here Araujo and Liotta identify definitive and universal structural requirements for adaptation via intermolecular interactions.
- Robyn P. Araujo
- & Lance A. Liotta
-
Article
| Open Access Data integration across conditions improves turnover number estimates and metabolic predictions
Data integration across conditions improves turnover number estimates and metabolic predictions
The construction of protein-constrained genome-scale metabolic models depends on the integration of organism-specific enzyme turnover numbers. Here, the authors show that correction of turnover numbers by simultaneous consideration of proteomics and physiological data leads to improved predictions of condition-specific growth rates.
- Philipp Wendering
- , Marius Arend
- & Zoran Nikoloski
-
Article
| Open Access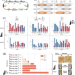 The automated Galaxy-SynBioCAD pipeline for synthetic biology design and engineering
The automated Galaxy-SynBioCAD pipeline for synthetic biology design and engineering
Automated design and build processes can rapidly accelerate work in synthetic biology and metabolic engineering. Here the authors present Galaxy-SynBioCAD, a toolshed for synthetic biology, metabolic engineering, and industrial biotechnology that they use to build and execute Galaxy scientific workflows from pathway design to strain engineering through the automated generation of scripts driving robotic workstations.
- Joan Hérisson
- , Thomas Duigou
- & Jean-Loup Faulon
-
Article
| Open Access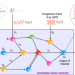 Frequency spectra and the color of cellular noise
Frequency spectra and the color of cellular noise
The invention of the Fourier integral in the 19th century laid the foundation for modern spectral analysis methods. Here the authors develop frequency-based methods for analyzing the reaction mechanisms within living cells from distinctively noisy single-cell output trajectories and present forward engineering of synthetic oscillators and controllers.
- Ankit Gupta
- & Mustafa Khammash
-
Article
| Open Access A versatile active learning workflow for optimization of genetic and metabolic networks
A versatile active learning workflow for optimization of genetic and metabolic networks
Optimization of biological networks is often limited by wet lab labor and cost, and the lack of convenient computational tools. Here, aimed at democratization and standardization, the authors describe METIS, a modular and versatile active machine learning workflow with a simple online interface for the optimization of biological target functions with minimal experimental datasets.
- Amir Pandi
- , Christoph Diehl
- & Tobias J. Erb
-
Article
| Open Access Reconstruction of a catalogue of genome-scale metabolic models with enzymatic constraints using GECKO 2.0
Reconstruction of a catalogue of genome-scale metabolic models with enzymatic constraints using GECKO 2.0
Genome-scale metabolic models have been widely used for quantitative exploration of the relation between genotype and phenotype. Here the authors present GECKO 2, an automated framework for continuous and version controlled update of enzyme-constrained models of metabolism, producing an interesting catalogue of high-quality models for diverse yeasts, bacteria and human metabolism, aiming to facilitate their use in basic science, metabolic engineering and synthetic biology purposes.
- Iván Domenzain
- , Benjamín Sánchez
- & Jens Nielsen
-
Article
| Open Access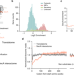 Ribosome profiling reveals multiple roles of SecA in cotranslational protein export
Ribosome profiling reveals multiple roles of SecA in cotranslational protein export
Using a combination of ribosome profiling methods, Zhu et al. investigate the principles governing the cotranslational interaction of SecA with nascent proteins and reveal a hierarchical organization of protein export pathways in bacteria.
- Zikun Zhu
- , Shuai Wang
- & Shu-ou Shan
-
Article
| Open Access Deep learning driven biosynthetic pathways navigation for natural products with BioNavi-NP
Deep learning driven biosynthetic pathways navigation for natural products with BioNavi-NP
The complete biosynthetic pathway from most natural products (NPs) are unknown. Here, the authors report BioNavi-NP, a computational toolkit for bio-retrosynthetic pathway elucidation or reconstruction for both NPs and NP-like compounds.
- Shuangjia Zheng
- , Tao Zeng
- & Ruibo Wu
-
Article
| Open Access A hierarchy of biomolecular proportional-integral-derivative feedback controllers for robust perfect adaptation and dynamic performance
A hierarchy of biomolecular proportional-integral-derivative feedback controllers for robust perfect adaptation and dynamic performance
The design of feedback biomolecular controllers is essential to synthetically regulate biological processes in a robust and timely fashion. Here the authors introduce a wide array of biomolecular Proportional-Integral-Derivative (PID) controllers that are capable of enhancing stability and dynamic performance, and also reducing stochastic noise.
- Maurice Filo
- , Sant Kumar
- & Mustafa Khammash
-
Article
| Open Access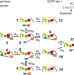 A general theoretical framework to design base editors with reduced bystander effects
A general theoretical framework to design base editors with reduced bystander effects
Base editors can edit target nucleotides, and identical ones that are within the editing window. Here the authors build an analytical model to propose general principles of editor design to reduce bystander effects.
- Qian Wang
- , Jie Yang
- & Anatoly B. Kolomeisky
-
Article
| Open Access Correcting for sparsity and interdependence in glycomics by accounting for glycan biosynthesis
Correcting for sparsity and interdependence in glycomics by accounting for glycan biosynthesis
Glycomics can uncover important molecular changes but measured glycans are highly interconnected and incompatible with common statistical methods, introducing pitfalls during analysis. Here, the authors develop an approach to identify glycan dependencies across samples to facilitate comparative glycomics.
- Bokan Bao
- , Benjamin P. Kellman
- & Nathan E. Lewis
-
Article
| Open Access An extended reconstruction of human gut microbiota metabolism of dietary compounds
An extended reconstruction of human gut microbiota metabolism of dietary compounds
The interplay between human diet and the gut microbiome is complex. Here, the authors present a model of human-microbiome interaction that can predict how phenolic compounds are metabolized by the human gut microbiome, identifying diet-specific metabolites in children of varied clinical conditions.
- Telmo Blasco
- , Sergio Pérez-Burillo
- & Francisco J. Planes
-
Article
| Open Access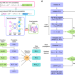 Neural network aided approximation and parameter inference of non-Markovian models of gene expression
Neural network aided approximation and parameter inference of non-Markovian models of gene expression
Cells are complex systems that make decisions biologists struggle to understand. Here, the authors use neural networks to approximate the solution of mathematical models that capture the history and randomness of biochemical processes in order to understand the principles of transcription control.
- Qingchao Jiang
- , Xiaoming Fu
- & Ramon Grima
-
Article
| Open Access Integration of machine learning and genome-scale metabolic modeling identifies multi-omics biomarkers for radiation resistance
Integration of machine learning and genome-scale metabolic modeling identifies multi-omics biomarkers for radiation resistance
Personalized prediction of tumor radiosensitivity would facilitate development of precision medicine workflows for cancer treatment. Here, the authors integrate machine learning and genome-scale metabolic modeling approaches to identify multi-omics biomarkers predictive of radiation response.
- Joshua E. Lewis
- & Melissa L. Kemp
-
Article
| Open Access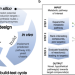 A computational workflow for the expansion of heterologous biosynthetic pathways to natural product derivatives
A computational workflow for the expansion of heterologous biosynthetic pathways to natural product derivatives
The top down cheminformatics method is usually used for the reconstitution of heterologous pathway to produce plant natural products. Here, the authors report a bottom up computational workflow for the identification of potential products and the enzymes required to make them in a noscapine pathway in yeast.
- Jasmin Hafner
- , James Payne
- & Christina Smolke
-
Article
| Open Access A ribosome-associated chaperone enables substrate triage in a cotranslational protein targeting complex
A ribosome-associated chaperone enables substrate triage in a cotranslational protein targeting complex
Biochemistry combined with biophysical measurements and mathematical modeling offer insight into the mechanism by which the cotranslational chaperone, nascent polypeptide-associated complex (NAC), modulates substrate selection by signal recognition particle (SRP) and reduces aberrant, nonspecific targeting of ribosomes to the ER.
- Hao-Hsuan Hsieh
- , Jae Ho Lee
- & Shu-ou Shan
-
Article
| Open Access Methanol-dependent Escherichia coli strains with a complete ribulose monophosphate cycle
Methanol-dependent Escherichia coli strains with a complete ribulose monophosphate cycle
The engineering of methanol-dependent growth in Escherichia coli is challenging. Here, the authors predict and experimentally validate methanol-dependent strains with a complete RuMP cycle and high potential for the development of a methylotrophic platform organism.
- Philipp Keller
- , Elad Noor
- & Julia A. Vorholt
-
Article
| Open Access A strategy to incorporate prior knowledge into correlation network cutoff selection
A strategy to incorporate prior knowledge into correlation network cutoff selection
Correlation network inference is typically based on the significance of the correlation coefficients, but this procedure is not guaranteed to capture biological mechanisms. Here, the authors develop a cutoff selection algorithm that maximizes the overlap between inferred networks and prior knowledge.
- Elisa Benedetti
- , Maja Pučić-Baković
- & Jan Krumsiek
-
Article
| Open Access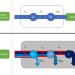 An analytical theory of balanced cellular growth
An analytical theory of balanced cellular growth
Genome-scale models of microbial metabolism largely ignore reaction kinetics. Here, the authors develop a general mathematical framework for modeling cellular growth with explicit non-linear reaction kinetics and use it to glean insights into the principles of cellular resource allocation and growth.
- Hugo Dourado
- & Martin J. Lercher
-
Article
| Open Access Minimal exposure of lipid II cycle intermediates triggers cell wall antibiotic resistance
Minimal exposure of lipid II cycle intermediates triggers cell wall antibiotic resistance
Antibiotics targeting cell wall synthesis display an unexplained gap between in vivo efficacy and in vitro binding affinity for their target. Here, Piepenbreier et al. develop a model for bacterial cell wall biosynthesis, show how it is affected by antibiotics, and use it to predict in vivo efficacy of antibiotics.
- Hannah Piepenbreier
- , Angelika Diehl
- & Georg Fritz
-
Article
| Open Access Probabilistic controllability approach to metabolic fluxes in normal and cancer tissues
Probabilistic controllability approach to metabolic fluxes in normal and cancer tissues
Metabolic rewiring is a feature of many cancers. Here, the authors combine control theory and flux correlation analysis to study the transition of healthy metabolic networks to cancer states, and find that cancer metabolism is characterized by more streamlined flux distributions.
- Jean-Marc Schwartz
- , Hiroaki Otokuni
- & Jose C. Nacher
-
Article
| Open Access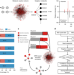 Metabolic reaction network-based recursive metabolite annotation for untargeted metabolomics
Metabolic reaction network-based recursive metabolite annotation for untargeted metabolomics
Untargeted metabolomics detects large numbers of metabolites but their annotation remains challenging. Here, the authors develop a metabolic reaction network-based recursive algorithm that expands metabolite annotation by taking advantage of the mass spectral similarity of reaction-paired neighbor metabolites.
- Xiaotao Shen
- , Ruohong Wang
- & Zheng-Jiang Zhu
-
Article
| Open Access Costless metabolic secretions as drivers of interspecies interactions in microbial ecosystems
Costless metabolic secretions as drivers of interspecies interactions in microbial ecosystems
In considering cross-feeding among microbes within communities, it is typically assumed that metabolic secretions are costly to produce. However, Pacheco et al. use metabolic models to show that ‘costless’ secretions could be common in some environments and important for structuring interactions among microbes.
- Alan R. Pacheco
- , Mauricio Moel
- & Daniel Segrè
-
Article
| Open Access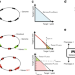 MoVE identifies metabolic valves to switch between phenotypic states
MoVE identifies metabolic valves to switch between phenotypic states
Interventions in metabolic networks can improve microbes for chemical production. Here, the authors develop a tool to identify metabolic valves that can decouple growth and production to systematically improve the rate and yield of biochemical production processes.
- Naveen Venayak
- , Axel von Kamp
- & Radhakrishnan Mahadevan
-
Article
| Open Access Machine learning applied to enzyme turnover numbers reveals protein structural correlates and improves metabolic models
Machine learning applied to enzyme turnover numbers reveals protein structural correlates and improves metabolic models
Experimental data on enzyme turnover numbers is sparse and noisy. Here, the authors use machine learning to successfully predict enzyme turnover numbers for E. coli, and show that using these to parameterize mechanistic genome-scale models enhances their predictive accuracy.
- David Heckmann
- , Colton J. Lloyd
- & Bernhard O. Palsson
-
Article
| Open Access Evolution of gene knockout strains of E. coli reveal regulatory architectures governed by metabolism
Evolution of gene knockout strains of E. coli reveal regulatory architectures governed by metabolism
The function of metabolic genes in the context of regulatory networks is not well understood. Here, the authors investigate the adaptive responses of E. coli after knockout of metabolic genes and highlight the influence of metabolite levels in the evolution of regulatory function.
- Douglas McCloskey
- , Sibei Xu
- & Bernhard O. Palsson
-
Article
| Open Access Exploring the role of stromal osmoregulation in cancer and disease using executable modelling
Exploring the role of stromal osmoregulation in cancer and disease using executable modelling
Aberrant ion transporter expression disrupts osmoregulation in many cancers. Here, the authors introduce an executable model of osmotic regulation and membrane transport, illuminating the mechanistic basis of multiple cellular cancer phenotypes and suggesting therapeutic avenues.
- David Shorthouse
- , Angela Riedel
- & Benjamin A. Hall
-
Article
| Open Access Systems analysis of intracellular pH vulnerabilities for cancer therapy
Systems analysis of intracellular pH vulnerabilities for cancer therapy
Tumors often exhibit a pH gradient, with an acidic extracellular space and alkaline cytoplasm. Here the authors develop a computational model to show how alkaline pHi supports changes inherent to cancer cell metabolism and acidification disables these adaptations, and demonstrate the effect of acidic pHi on breast cancer cell survival.
- Erez Persi
- , Miquel Duran-Frigola
- & Eytan Ruppin
-
Article
| Open Access HEPATOKIN1 is a biochemistry-based model of liver metabolism for applications in medicine and pharmacology
HEPATOKIN1 is a biochemistry-based model of liver metabolism for applications in medicine and pharmacology
In silico models of cells can provide insight into the causes and effects of disease states and reduce the need for in vivo studies. Here, the authors present a kinetic model of hepatocyte metabolism including energy, carbohydrate, lipid and nitrogen metabolism and hormonal and allosteric regulation of enzymatic activity.
- Nikolaus Berndt
- , Sascha Bulik
- & Hermann-Georg Holzhütter
-
Article
| Open Access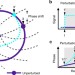 Design principles for enhancing phase sensitivity and suppressing phase fluctuations simultaneously in biochemical oscillatory systems
Design principles for enhancing phase sensitivity and suppressing phase fluctuations simultaneously in biochemical oscillatory systems
Biochemical processes require both high sensitivity and low fluctuation which is incompatible with the fluctuation dissipation theorem. Here Fei et al. model biochemical oscillators to show how free energy dissipation leads to both a suppression of phase fluctuation and an enhancement of phase sensitivity.
- Chenyi Fei
- , Yuansheng Cao
- & Yuhai Tu
-
Article
| Open Access Pathway design using de novo steps through uncharted biochemical spaces
Pathway design using de novo steps through uncharted biochemical spaces
Existing pathway design tools make use of existing reactions from databases or successively apply retrosynthetic rules. novoStoic provides an integrated optimization-based framework combining known reactions with novel steps in pathway design allowing for constraints on thermodynamic feasibility, product yield, pathway length and number of novel steps.
- Akhil Kumar
- , Lin Wang
- & Costas D. Maranas
-
Article
| Open Access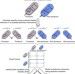 Genome-driven evolutionary game theory helps understand the rise of metabolic interdependencies in microbial communities
Genome-driven evolutionary game theory helps understand the rise of metabolic interdependencies in microbial communities
The rise of metabolic interdependencies among microbes is still poorly understood. Here, taking the underlying biochemical networks into consideration, Zomorrodi and Segrè integrate genome-scale metabolic models with evolutionary game theory to study the rise of cross-feeding in microbial communities.
- Ali R. Zomorrodi
- & Daniel Segrè
-
Article
| Open Access An in-silico approach to predict and exploit synthetic lethality in cancer metabolism
An in-silico approach to predict and exploit synthetic lethality in cancer metabolism
Exploiting synthetic lethality is a promising approach for cancer therapy. Here, the authors present an approach to identifying such interactions by finding genetic minimal cut sets (gMCSs) that block cancer proliferation, and apply it to study the lethality of RRM1 inhibition in multiple myeloma.
- Iñigo Apaolaza
- , Edurne San José-Eneriz
- & Francisco J. Planes
-
Article
| Open Access Growth-coupled overproduction is feasible for almost all metabolites in five major production organisms
Growth-coupled overproduction is feasible for almost all metabolites in five major production organisms
Coupling of growth and product synthesis is an important principle in metabolic engineering, but its range of applicability is unclear. Here, the authors use a dedicated computational framework to study the feasibility of coupling the production of metabolites to growth in the genome-scale metabolic models of five production organisms, and show that coupling can be achieved for most metabolites.
- Axel von Kamp
- & Steffen Klamt
-
Article
| Open Access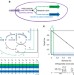 Redesigning metabolism based on orthogonality principles
Redesigning metabolism based on orthogonality principles
Growth-coupled designs for chemical production are limited by native metabolic networks’ optimality for growth. Here, the authors introduce pathway orthogonality as a measure of the independence of biomass and chemical production pathways, identify metabolic valves that allow substrate utilization to be switched between the two, and demonstrate advantages of orthogonal designs.
- Aditya Vikram Pandit
- , Shyam Srinivasan
- & Radhakrishnan Mahadevan
-
Article
| Open Access An analytic approximation of the feasible space of metabolic networks
An analytic approximation of the feasible space of metabolic networks
Large-scale metabolic models of organisms from microbes to mammals can provide great insight into cellular function, but their analysis remains challenging. Here, the authors provide an approximate analytic method to estimate the feasible solution space for the flux vectors of metabolic networks, enabling more accurate analysis under a wide range of conditions of interest.
- Alfredo Braunstein
- , Anna Paola Muntoni
- & Andrea Pagnani
-
Article
| Open Access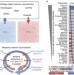 Reconciled rat and human metabolic networks for comparative toxicogenomics and biomarker predictions
Reconciled rat and human metabolic networks for comparative toxicogenomics and biomarker predictions
The rat is a widely-used model for human biology, but we must be aware of metabolic differences. Here, the authors reconstruct the genome-scale metabolic network of the rat, and after reconciling it with an improved human metabolic model, demonstrate the power of the models to integrate toxicogenomics data, providing species-specific biomarker predictions in response to a panel of drugs.
- Edik M. Blais
- , Kristopher D. Rawls
- & Jason A. Papin
-
Article
| Open Access A network property necessary for concentration robustness
A network property necessary for concentration robustness
Absolute concentration robustness (ACR), independence of the steady-state concentration of a molecule from the environment, is difficult to predict. Here, the authors derive a network structure-based necessary condition for ACR, and suggest that metabolites satisfying the condition are prevalent.
- Jeanne M. O. Eloundou-Mbebi
- , Anika Küken
- & Zoran Nikoloski
-
Article
| Open Access Forward design of a complex enzyme cascade reaction
Forward design of a complex enzyme cascade reaction
Building multi-component enzymatic processes in one pot is challenged by the inherent complexity of each biochemical system. Here, the authors use online mass spectroscopy and engineering systems theory to achieve forward design of a ten-membered reaction cascade.
- Christoph Hold
- , Sonja Billerbeck
- & Sven Panke

 Viscosity-dependent control of protein synthesis and degradation
Viscosity-dependent control of protein synthesis and degradation