Featured
-
-
Article
| Open Access Reconstructing the evolution history of networked complex systems
Reconstructing the evolution history of networked complex systems
Evolution processes of complex networked systems in biology and social sciences, and their underlying mechanisms, still need better understanding. The authors propose a machine learning approach to reconstruct the evolution history of complex networks.
- Junya Wang
- , Yi-Jiao Zhang
- & Yanqing Hu
-
Article
| Open Access Nutritional redundancy in the human diet and its application in phenotype association studies
Nutritional redundancy in the human diet and its application in phenotype association studies
Studying human diet may help us identify measures to treat or prevent chronic diseases. Here, the authors discover the nutritional redundancy phenomenon in human diet and demonstrate its association with cardiovascular disease and type 2 diabetes.
- Xu-Wen Wang
- , Yang Hu
- & Yang-Yu Liu
-
Article
| Open Access Revealing proteome-level functional redundancy in the human gut microbiome using ultra-deep metaproteomics
Revealing proteome-level functional redundancy in the human gut microbiome using ultra-deep metaproteomics
Here, Li et al. show that functional redundancy, which has not previously been quantified at the proteome level, can arise when different microbes play similar roles in the gut microbiome, revealing that proteomes are nested among gut microbes, favoring high functional redundancy.
- Leyuan Li
- , Tong Wang
- & Daniel Figeys
-
Article
| Open Access A positive statistical benchmark to assess network agreement
A positive statistical benchmark to assess network agreement
In the variety of biological and social networks, the validation of experimental data is done by comparing an overlap with reference networks. The authors introduce a positive statistical benchmark corresponding to the best possible overlap between two networks to threshold and validate new experimental datasets.
- Bingjie Hao
- & István A. Kovács
-
Article
| Open Access Universal structures for adaptation in biochemical reaction networks
Universal structures for adaptation in biochemical reaction networks
At the molecular level, the evolution of life is driven by the generation and diversification of adaptation mechanisms. Here Araujo and Liotta identify definitive and universal structural requirements for adaptation via intermolecular interactions.
- Robyn P. Araujo
- & Lance A. Liotta
-
Article
| Open Access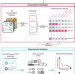 Assessment of community efforts to advance network-based prediction of protein–protein interactions
Assessment of community efforts to advance network-based prediction of protein–protein interactions
Comprehensive understanding of the human protein-protein interaction network, aka the human interactome, can provide important insights into the molecular mechanisms of complex biological processes and diseases. Here the authors summarize the community efforts initiated by the International Network Medicine Consortium to benchmark the ability of 26 representative network-based methods to predict protein-protein interactions.
- Xu-Wen Wang
- , Lorenzo Madeddu
- & Yang-Yu Liu
-
Article
| Open Access Precision dynamical mapping using topological data analysis reveals a hub-like transition state at rest
Precision dynamical mapping using topological data analysis reveals a hub-like transition state at rest
Although spontaneous brain activity is complex and clinically relevant, it is still unclear whether transitions in resting brain activity follow an underlying arrangement or whether they are unpredictable. In this work, the authors revealed a transition state of the brain that acts like a switch between states and forms the basis for the continuous evolution of brain activity patterns at rest.
- Manish Saggar
- , James M. Shine
- & Damien Fair
-
Article
| Open Access A randomized controlled trial for response of microbiome network to exercise and diet intervention in patients with nonalcoholic fatty liver disease
A randomized controlled trial for response of microbiome network to exercise and diet intervention in patients with nonalcoholic fatty liver disease
Exercise and diet interventions are treatments for nonalcoholic fatty liver disease (NAFLD). Here the authors report that in randomized, controlled trial in patients with NAFLD exercise and diet intervention were associated with diversified gut microbiome keystone taxa. Exploratory analysis suggests gut microbial network may be used to predict the individual liver fat response to exercise intervention, if validated in future studies.
- Runtan Cheng
- , Lu Wang
- & Sulin Cheng
-
Article
| Open Access Leveraging the Cell Ontology to classify unseen cell types
Leveraging the Cell Ontology to classify unseen cell types
Classifying cells into unseen cell types remains challenging in scRNA-seq analysis. Here we show that Cell Ontology enables an accurate classification of unseen cell types through considering the cell type relationships in the Cell Ontology graph.
- Sheng Wang
- , Angela Oliveira Pisco
- & Russ B. Altman
-
Article
| Open Access Divide-and-conquer: machine-learning integrates mammalian and viral traits with network features to predict virus-mammal associations
Divide-and-conquer: machine-learning integrates mammalian and viral traits with network features to predict virus-mammal associations
A more comprehensive map of viral host ranges can help identify and mitigate zoonotic and animal-disease risks. A divide-and-conquer approach which separates viral, mammalian and network features predicts over 20,000 unknown associations between known viruses and susceptible mammalian species.
- Maya Wardeh
- , Marcus S. C. Blagrove
- & Matthew Baylis
-
Article
| Open Access The VRNetzer platform enables interactive network analysis in Virtual Reality
The VRNetzer platform enables interactive network analysis in Virtual Reality
Data-rich networks can be difficult to interpret beyond a certain size. Here, the authors introduce a platform that uses virtual reality to allow the visual exploration of large networks, while interfacing with data repositories and other analytical methods to improve the interpretation of big data.
- Sebastian Pirch
- , Felix Müller
- & Jörg Menche
-
Article
| Open Access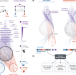 Identification of disease treatment mechanisms through the multiscale interactome
Identification of disease treatment mechanisms through the multiscale interactome
Most diseases disrupt multiple proteins, and drugs treat such diseases by restoring the functions of the disrupted proteins; how drugs restore these functions, however, is often unknown. Here, the authors develop the multiscale interactome, a powerful approach to explain disease treatment.
- Camilo Ruiz
- , Marinka Zitnik
- & Jure Leskovec
-
Article
| Open Access Forecasting influenza activity using machine-learned mobility map
Forecasting influenza activity using machine-learned mobility map
Human mobility plays a central role in the spread of infectious diseases and can help in forecasting incidence. Here the authors show a comparison of multiple mobility benchmarks in forecasting influenza, and demonstrate the value of a machine-learned mobility map with global coverage at multiple spatial scales.
- Srinivasan Venkatramanan
- , Adam Sadilek
- & Madhav Marathe
-
Article
| Open Access Individualized interactomes for network-based precision medicine in hypertrophic cardiomyopathy with implications for other clinical pathophenotypes
Individualized interactomes for network-based precision medicine in hypertrophic cardiomyopathy with implications for other clinical pathophenotypes
Understanding patient-specific pathobiological pathways is a critical step for advancing precision medicine. Here the authors show that individualized protein-protein interaction networks provide key insight on patient-level pathobiology and clinically relevant pathophenotypic characteristics in a complex disease.
- Bradley A. Maron
- , Rui-Sheng Wang
- & Joseph Loscalzo
-
Article
| Open Access Nonlinear mechanics of lamin filaments and the meshwork topology build an emergent nuclear lamina
Nonlinear mechanics of lamin filaments and the meshwork topology build an emergent nuclear lamina
Mechanical strength of in situ assembled nuclear lamin filaments arranged in a 3D meshwork is unclear. Here, using mechanical, structural and simulation tools, the authors report the hierarchical organization of the lamin meshwork that imparts strength and toughness to lamin filaments at par with silk and Kevlar®
- K. Tanuj Sapra
- , Zhao Qin
- & Ohad Medalia
-
Article
| Open Access Deciphering functional redundancy in the human microbiome
Deciphering functional redundancy in the human microbiome
Here, the authors develop a genome evolution model to investigate the origin of functional redundancy in the human microbiome by analyzing its genomic content network and illustrate potential ecological and evolutionary processes that may contribute to its resilience.
- Liang Tian
- , Xu-Wen Wang
- & Yang-Yu Liu
-
Article
| Open Access Mutualistic networks emerging from adaptive niche-based interactions
Mutualistic networks emerging from adaptive niche-based interactions
Nested and modular patterns are vastly observed in mutualistic networks across genres and geographic conditions. Here, the authors show a unified mechanism that underlies the assembly and evolution of such networks, based on adaptive niche interactions of the participants.
- Weiran Cai
- , Jordan Snyder
- & Raissa M. D’Souza
-
Article
| Open Access Mapping allosteric communications within individual proteins
Mapping allosteric communications within individual proteins
The computational prediction of protein allostery can guide experimental studies of protein function and cellular activity. Here, the authors develop a network-based method to detect allosteric coupling within proteins solely based on their structures, and set up a webserver for allostery prediction.
- Jian Wang
- , Abha Jain
- & Nikolay V. Dokholyan
-
Article
| Open Access Exploring the SARS-CoV-2 virus-host-drug interactome for drug repurposing
Exploring the SARS-CoV-2 virus-host-drug interactome for drug repurposing
Information developed to understand the molecular mechanisms of SARS-CoV-2 infection for predicting drug repurposing candidates is time-consuming to integrate and explore. Here, the authors develop an interactive online platform for virus-host interactome exploration and drug (target) identification.
- Sepideh Sadegh
- , Julian Matschinske
- & Jan Baumbach
-
Article
| Open Access Partial cross mapping eliminates indirect causal influences
Partial cross mapping eliminates indirect causal influences
It is crucial yet challenging to identify cause-consequence relation in complex dynamical systems where direct causal links can mix with indirect ones. Leng et al. propose a data-driven model-independent method to distinguish direct from indirect causality and test its applicability to real-world data.
- Siyang Leng
- , Huanfei Ma
- & Luonan Chen
-
Article
| Open Access Discovering the genes mediating the interactions between chronic respiratory diseases in the human interactome
Discovering the genes mediating the interactions between chronic respiratory diseases in the human interactome
Complex diseases often share genetic determinants and symptoms, but the mechanistic basis of disease interactions remains elusive. Here, the authors propose a network topological measure to identify proteins linking complex diseases in the interactome, and identify mediators between COPD and asthma.
- Enrico Maiorino
- , Seung Han Baek
- & Amitabh Sharma
-
Article
| Open Access Mapping the perturbome network of cellular perturbations
Mapping the perturbome network of cellular perturbations
Our understanding of the mechanisms of drug interactions remains limited. Here the authors introduce a framework to study how complex cellular perturbations induced by different drugs affect each other in morphological feature space.
- Michael Caldera
- , Felix Müller
- & Jörg Menche
-
Article
| Open Access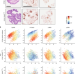 Cell type-dependent differential activation of ERK by oncogenic KRAS in colon cancer and intestinal epithelium
Cell type-dependent differential activation of ERK by oncogenic KRAS in colon cancer and intestinal epithelium
KRASG12V and BRAFV600E are oncogenic mutations that activate ERK signalling. Here, the authors use single cell analysis in intestinal organoids and show that BRAFV600E activates ERK in all intestinal cell types, while KRASG12V induces ERK activation in only a subset of cells, depending on cell differentiation state.
- Raphael Brandt
- , Thomas Sell
- & Markus Morkel
-
Article
| Open Access Feed-forward regulation adaptively evolves via dynamics rather than topology when there is intrinsic noise
Feed-forward regulation adaptively evolves via dynamics rather than topology when there is intrinsic noise
Feed‐forward loops (FFLs) can filter out noise, but whether their overrepresentation in GRNs reflects adaptive evolution for this function is debated. Here, the authors develop a null model of regulatory evolution and find that FFLs evolve readily under selection for the noise filtering function.
- Kun Xiong
- , Alex K. Lancaster
- & Joanna Masel
-
Article
| Open Access Network-based prediction of drug combinations
Network-based prediction of drug combinations
Combination therapy holds great promise, but discovery remains challenging. Here, the authors propose a method to identify efficacious drug combinations for specific diseases, and find that successful combinations tend to target separate neighbourhoods of the disease module in the human interactome.
- Feixiong Cheng
- , István A. Kovács
- & Albert-László Barabási
-
Article
| Open Access Topological scoring of protein interaction networks
Topological scoring of protein interaction networks
Inferring direct protein−protein interactions (PPIs) and modules in PPI networks remains a challenge. Here, the authors introduce an algorithm to infer potential direct PPIs from quantitative proteomic AP-MS data by identifying enriched interactions of each bait relative to the other baits.
- Mihaela E. Sardiu
- , Joshua M. Gilmore
- & Michael P. Washburn
-
Article
| Open Access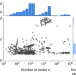 Scale-free networks are rare
Scale-free networks are rare
Real-world networks are often said to be ”scale free”, meaning their degree distribution follows a power law. Broido and Clauset perform statistical tests of this claim using a large and diverse corpus of real-world networks, showing that scale-free structure is far from universal.
- Anna D. Broido
- & Aaron Clauset
-
Article
| Open Access An interpretable approach for social network formation among heterogeneous agents
An interpretable approach for social network formation among heterogeneous agents
Complex networks can be a useful tool to investigate problems in social science. Here the authors use game theory to establish a network model and then use a machine learning approach to characterize the role of nodes within a social network.
- Yuan Yuan
- , Ahmad Alabdulkareem
- & Alex ‘Sandy’ Pentland
-
Article
| Open Access Network integration of multi-tumour omics data suggests novel targeting strategies
Network integration of multi-tumour omics data suggests novel targeting strategies
Tumours of different tissues can show similarities in genomic alterations. Here, the authors combine tumour transcriptome and protein interaction data in a network-based analysis of 11 tumours types, and identify clusters of tumours with specific signatures for multi-tumour drug targeting and survival prognosis.
- Ítalo Faria do Valle
- , Giulia Menichetti
- & Daniel Remondini
-
Article
| Open Access Reprogramming of regulatory network using expression uncovers sex-specific gene regulation in Drosophila
Reprogramming of regulatory network using expression uncovers sex-specific gene regulation in Drosophila
For many applications knowledge of context-specific gene regulatory networks (GRNs) is desirable, but their inference remains a challenge. Here, the authors introduce a method for construction of context-specific GRNs, and apply it to construct sex-specific Drosophila GRNs.
- Yijie Wang
- , Dong-Yeon Cho
- & Teresa M. Przytycka
-
Article
| Open Access Network enhancement as a general method to denoise weighted biological networks
Network enhancement as a general method to denoise weighted biological networks
Technical noise in experiments is unavoidable, but it introduces inaccuracies into the biological networks we infer from the data. Here, the authors introduce a diffusion-based method for denoising undirected, weighted networks, and show that it improves the performances of downstream analyses.
- Bo Wang
- , Armin Pourshafeie
- & Jure Leskovec
-
Article
| Open Access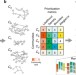 Prioritizing network communities
Prioritizing network communities
Community detection allows one to decompose a network into its building blocks. While communities can be identified with a variety of methods, their relative importance can’t be easily derived. Here the authors introduce an algorithm to identify modules which are most promising for further analysis.
- Marinka Zitnik
- , Rok Sosič
- & Jure Leskovec
-
Article
| Open Access Experimental noise cutoff boosts inferability of transcriptional networks in large-scale gene-deletion studies
Experimental noise cutoff boosts inferability of transcriptional networks in large-scale gene-deletion studies
Reliable inference of gene interactions from perturbation experiments remains a challenge. Here, the authors quantify the upper limits of transcriptional network inference from knockout screens, identify the key determinants of accuracy, and introduce an unbiased and scalable inference algorithm.
- C. F. Blum
- , N. Heramvand
- & M. Kollmann
-
Article
| Open Access Model-free inference of direct network interactions from nonlinear collective dynamics
Model-free inference of direct network interactions from nonlinear collective dynamics
Network dynamical systems can represent the interactions involved in the collective dynamics of gene regulatory networks or metabolic circuits. Here Casadiego et al. present a method for inferring these types of interactions directly from observed time series without relying on their model.
- Jose Casadiego
- , Mor Nitzan
- & Marc Timme
-
Article
| Open Access Percolation transition of cooperative mutational effects in colorectal tumorigenesis
Percolation transition of cooperative mutational effects in colorectal tumorigenesis
Cancer is caused by accumulating genetic mutations. Here, the authors investigate the cooperative effect of these mutations in colorectal cancer patients and identify a giant cluster of mutation-propagating modules that undergoes percolation transition during tumorigenesis.
- Dongkwan Shin
- , Jonghoon Lee
- & Kwang-Hyun Cho
-
Article
| Open Access A network integration approach for drug-target interaction prediction and computational drug repositioning from heterogeneous information
A network integration approach for drug-target interaction prediction and computational drug repositioning from heterogeneous information
Network-based data integration for drug–target prediction is a promising avenue for drug repositioning, but performance is wanting. Here, the authors introduce DTINet, whose performance is enhanced in the face of noisy, incomplete and high-dimensional biological data by learning low-dimensional vector representations.
- Yunan Luo
- , Xinbin Zhao
- & Jianyang Zeng
-
Article
| Open Access The interdependent network of gene regulation and metabolism is robust where it needs to be
The interdependent network of gene regulation and metabolism is robust where it needs to be
Although networks of interacting genes and metabolic reactions are interdependent, they have largely been treated as separate systems. Here the authors apply a statistical framework for interdependent networks to E. coli, and show that it is sensitive to gene and protein perturbations but robust against metabolic changes.
- David F. Klosik
- , Anne Grimbs
- & Marc-Thorsten Hütt
-
Article
| Open Access Single-cell entropy for accurate estimation of differentiation potency from a cell’s transcriptome
Single-cell entropy for accurate estimation of differentiation potency from a cell’s transcriptome
Robust quantification of the differentiation potential of single cells is a task of great importance. Here the authors integrate single-cell RNA-Seq profiles with a cellular interaction network to compute the signaling entropy, and show that it can identify normal and cancer stem-cell phenotypes.
- Andrew E. Teschendorff
- & Tariq Enver
-
Article
| Open Access Exercise contagion in a global social network
Exercise contagion in a global social network
Some argue that health-related behaviours, such as obesity, are contagious, but empirical evidence of health contagion remains inconclusive. Here, using a large scale quasi-experiment in a global network of runners, Aral and Nicolaides show that this type of contagion exists in fitness behaviours.
- Sinan Aral
- & Christos Nicolaides
-
Article
| Open Access Tor forms a dimer through an N-terminal helical solenoid with a complex topology
Tor forms a dimer through an N-terminal helical solenoid with a complex topology
The target of rapamycin (Tor) is a Ser/Thr protein kinase that regulates a wide range of anabolic and catabolic processes. Here the authors describe a sub-nanometer cryo-EM structure of a yeast Tor–Lst8 complex and propose an overall topology that differs from that previously suggested for mTORC1.
- Domagoj Baretić
- , Alex Berndt
- & Roger L. Williams
-
Article |
 Network physiology reveals relations between network topology and physiological function
Network physiology reveals relations between network topology and physiological function
Humans are a network of complex physiological systems, but quantifying these diverse systems is a challenge. This study presents a method to show that each physiological state is characterized by a specific network structure, demonstrating a connection between network topology and function.
- Amir Bashan
- , Ronny P. Bartsch
- & Plamen Ch. Ivanov

 Early detection of emerging viral variants through analysis of community structure of coordinated substitution networks
Early detection of emerging viral variants through analysis of community structure of coordinated substitution networks