Article
|
Open Access
Featured
-
-
Article
| Open Access High-throughput prediction of protein conformational distributions with subsampled AlphaFold2
High-throughput prediction of protein conformational distributions with subsampled AlphaFold2
Protein dynamics, crucial for life, are difficult and expensive to predict. This study shows that AI-based structure prediction methods can be modified for rapidly predicting the conformational landscapes of proteins, with strong correlations with experimentally-measured relative state populations.
- Gabriel Monteiro da Silva
- , Jennifer Y. Cui
- & Brenda M. Rubenstein
-
Article
| Open Access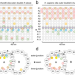 Uncovering structural themes across cilia microtubule inner proteins with implications for human cilia function
Uncovering structural themes across cilia microtubule inner proteins with implications for human cilia function
The inside surface of microtubules contains so-called microtubule inner proteins, but little is known about their identity. Here the authors use bioinformatics to identify structural motifs within this class of proteins and potential new members.
- Jens S. Andersen
- , Aaran Vijayakumaran
- & Kenneth Bødtker Schou
-
Article
| Open Access Accurate global and local 3D alignment of cryo-EM density maps using local spatial structural features
Accurate global and local 3D alignment of cryo-EM density maps using local spatial structural features
Density map alignment is a fundamental step in Cryo-EM data postprocessing. Here, authors propose an accurate global and local density map alignment method using local density features.
- Bintao He
- , Fa Zhang
- & Renmin Han
-
Article
| Open Access DynamicBind: predicting ligand-specific protein-ligand complex structure with a deep equivariant generative model
DynamicBind: predicting ligand-specific protein-ligand complex structure with a deep equivariant generative model
Proteins often function by changing conformations upon ligand binding. Efficient structural modelling of these interactions, crucial for drug discovery, is limited: here the authors address this with DynamicBind, a diffusion-based deep generative model.
- Wei Lu
- , Jixian Zhang
- & Shuangjia Zheng
-
Article
| Open Access From interaction networks to interfaces, scanning intrinsically disordered regions using AlphaFold2
From interaction networks to interfaces, scanning intrinsically disordered regions using AlphaFold2
Here, the authors show that AlphaFold2 accurately predicts protein interfaces involving disordered regions. Combining different delimitations and sequence alignments increases the success rate, while scanning short overlapping fragments identifies binding sites.
- Hélène Bret
- , Jinmei Gao
- & Raphaël Guerois
-
Article
| Open Access Enhancing geometric representations for molecules with equivariant vector-scalar interactive message passing
Enhancing geometric representations for molecules with equivariant vector-scalar interactive message passing
Utilising geometric information and reducing computational costs are key challenges in the molecular modelling field. Here, authors propose ViSNet, which efficiently extracts geometric features, accurately predicts molecular properties, and drives simulations with interpretability.
- Yusong Wang
- , Tong Wang
- & Tie-Yan Liu
-
Article
| Open Access Merizo: a rapid and accurate protein domain segmentation method using invariant point attention
Merizo: a rapid and accurate protein domain segmentation method using invariant point attention
Proteins contain modular structural and functional units called domains. Here, the authors have developed Merizo, a deep learning method for domain segmentation applicable to experimental structures as well as those generated by AlphaFold2.
- Andy M. Lau
- , Shaun M. Kandathil
- & David T. Jones
-
Article
| Open Access Accurate prediction of protein assembly structure by combining AlphaFold and symmetrical docking
Accurate prediction of protein assembly structure by combining AlphaFold and symmetrical docking
Current methods to predict structures of proteins cannot handle large assemblies with complex symmetries. Here, the authors demonstrate that structures of proteins with cubic symmetries can be accurately predicted with a method combining AlphaFold with symmetrical assembly simulations.
- Mads Jeppesen
- & Ingemar André
-
Article
| Open Access HLA3DB: comprehensive annotation of peptide/HLA complexes enables blind structure prediction of T cell epitopes
HLA3DB: comprehensive annotation of peptide/HLA complexes enables blind structure prediction of T cell epitopes
Structure prediction of peptide/HLA (pHLA) complexes has been mired by the inability to accurately model the middle of the peptide. Here, the authors present a curated database of pHLA structures (HLA3DB) and identify discrete peptide backbone conformations that are used for high fidelity modelling.
- Sagar Gupta
- , Santrupti Nerli
- & Nikolaos G. Sgourakis
-
Article
| Open Access The net electrostatic potential and hydration of ABCG2 affect substrate transport
The net electrostatic potential and hydration of ABCG2 affect substrate transport
ABCG2, an ATP-binding cassette transporter, extrudes hundreds of hydrophilic and hydrophobic compounds from cells, playing roles in xenobiotic clearance or multidrug resistance in cancer. Gose et al provide key insights into ABCG2 substrate selection.
- Tomoka Gose
- , Heather M. Aitken
- & John D. Schuetz
-
Article
| Open Access Deep transfer learning for inter-chain contact predictions of transmembrane protein complexes
Deep transfer learning for inter-chain contact predictions of transmembrane protein complexes
Membrane proteins are encoded by approximately a quarter of human genes. Here, the authors propose a deep transfer learning method for predicting inter-chain residue-residue contacts of transmembrane protein complexes.
- Peicong Lin
- , Yumeng Yan
- & Sheng-You Huang
-
Article
| Open Access Targeted cross-linker delivery for the in situ mapping of protein conformations and interactions in mitochondria
Targeted cross-linker delivery for the in situ mapping of protein conformations and interactions in mitochondria
Current methods for analysing protein structures and interactions generally require the separation of specific organelles or changes to the intracellular environment. Here, authors developed nanocarrier-based cross-linking mass spectrometry techniques to assess mitochondrial proteins within living cells.
- Yuwan Chen
- , Wen Zhou
- & Yukui Zhang
-
Article
| Open Access Designed active-site library reveals thousands of functional GFP variants
Designed active-site library reveals thousands of functional GFP variants
Mutations in a protein active site can alter function in useful ways, but the active site is sensitive to changes. Here the authors present a general strategy to design combinatorial mutation libraries. Applied to GFP, the authors isolate thousands of fluorescent designs that exhibit large and useful changes in spectral properties.
- Jonathan Yaacov Weinstein
- , Carlos Martí-Gómez
- & Sarel J. Fleishman
-
Article
| Open Access Improving de novo protein binder design with deep learning
Improving de novo protein binder design with deep learning
Recently, a pipeline for the design of protein-binding proteins using only the structure of the target protein was reported. Here, the authors report that the incorporation of deep learning methods into the original pipeline increases experimental success rate by ten-fold.
- Nathaniel R. Bennett
- , Brian Coventry
- & David Baker
-
Article
| Open Access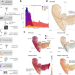 Sequence-structure-function relationships in the microbial protein universe
Sequence-structure-function relationships in the microbial protein universe
Advances in protein structure prediction have led to a significant influx of protein structure data. Here the authors exploit this data to offer an unbiased overview of complex sequence-structure-function relationships in the protein universe. This work opens up new uses for 3D structure data repositories in meta-omics and other fields of biology.
- Julia Koehler Leman
- , Pawel Szczerbiak
- & Tomasz Kosciolek
-
Article
| Open Access Fast, accurate antibody structure prediction from deep learning on massive set of natural antibodies
Fast, accurate antibody structure prediction from deep learning on massive set of natural antibodies
Prediction of antibody structures is critical for understanding and designing novel therapeutic and diagnostic molecules. Here, the authors present IgFold: a fast, accurate method for antibody structure prediction using an end-to-end deep learning model.
- Jeffrey A. Ruffolo
- , Lee-Shin Chu
- & Jeffrey J. Gray
-
Article
| Open Access PeSTo: parameter-free geometric deep learning for accurate prediction of protein binding interfaces
PeSTo: parameter-free geometric deep learning for accurate prediction of protein binding interfaces
Predicting protein interactions is crucial for understanding biological functions. Here, authors introduce a geometric transformer that accurately identifies protein binding interfaces, enabling new insights into unexplored biology.
- Lucien F. Krapp
- , Luciano A. Abriata
- & Matteo Dal Peraro
-
Article
| Open Access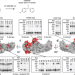 Structural details of a Class B GPCR-arrestin complex revealed by genetically encoded crosslinkers in living cells
Structural details of a Class B GPCR-arrestin complex revealed by genetically encoded crosslinkers in living cells
The conformation of GPCR-arrestin complexes at the cell membrane, despite available structures, remains uncertain. This work reveals structure and dynamics of the PTH1R-arrestin2 complex, including flexible regions, in live cells.
- Yasmin Aydin
- , Thore Böttke
- & Irene Coin
-
Article
| Open Access Evaluating native-like structures of RNA-protein complexes through the deep learning method
Evaluating native-like structures of RNA-protein complexes through the deep learning method
RNA-protein docking is a very challenging area. Here, the authors develop a deep-learning based method, DRPScore, to evaluate RNA-protein complexes. DRPScore is robust and consistently performs better than existing methods on representative testing sets.
- Chengwei Zeng
- , Yiren Jian
- & Yunjie Zhao
-
Article
| Open Access Protein complex prediction using Rosetta, AlphaFold, and mass spectrometry covalent labeling
Protein complex prediction using Rosetta, AlphaFold, and mass spectrometry covalent labeling
Covalent labeling (CL) from mass spectrometry experiments provides structural information of higher-order protein structure. Here, the authors develop an algorithm which integrates experimental CL data to predict protein complexes in the Rosetta molecular modeling suite using AlphaFold models.
- Zachary C. Drake
- , Justin T. Seffernick
- & Steffen Lindert
-
Article
| Open Access Proteome-wide 3D structure prediction provides insights into the ancestral metabolism of ancient archaea and bacteria
Proteome-wide 3D structure prediction provides insights into the ancestral metabolism of ancient archaea and bacteria
Previous studies have reconstructed ancestral metabolism using sequence-based approaches. This study uses a high-throughput version of AlphaFold2 to compare proteome-wide 3D structure predictions of two representative strains of ancient archaea and bacteria.
- Weishu Zhao
- , Bozitao Zhong
- & Xiang Xiao
-
Article
| Open Access Prediction of inter-chain distance maps of protein complexes with 2D attention-based deep neural networks
Prediction of inter-chain distance maps of protein complexes with 2D attention-based deep neural networks
Predicting inter-chain residue-residue distances of protein complexes is useful for constructing and evaluating quaternary structures of the protein complexes. Here, the authors develop a deep attention-based residual network method (CDPred) to predict inter-chain residue-residue distances of protein dimers.
- Zhiye Guo
- , Jian Liu
- & Jianlin Cheng
-
Article
| Open Access Predicting the structure of large protein complexes using AlphaFold and Monte Carlo tree search
Predicting the structure of large protein complexes using AlphaFold and Monte Carlo tree search
The accuracy of AlphaFold decreases with the number of protein chains and the available GPU memory limits the size of protein complexes that can be predicted. Here, the authors show that complexes with 10–30 chains can be assembled from predicted subcomponents using Monte Carlo tree search.
- Patrick Bryant
- , Gabriele Pozzati
- & Arne Elofsson
-
Article
| Open Access Predicting the structural basis of targeted protein degradation by integrating molecular dynamics simulations with structural mass spectrometry
Predicting the structural basis of targeted protein degradation by integrating molecular dynamics simulations with structural mass spectrometry
The formation of ternary degrader-protein complexes is a key step in the targeted degradation of proteins of interest. Here, the authors explore the structure and dynamics of such complexes applying high-performance computer simulations augmented with experimental data.
- Tom Dixon
- , Derek MacPherson
- & Jesus A. Izaguirre
-
Article
| Open Access Structure of the PAPP-ABP5 complex reveals mechanism of substrate recognition
Structure of the PAPP-ABP5 complex reveals mechanism of substrate recognition
PAPP-A substrate selectivity underlies the tight regulation of IGF signaling. Here, the authors report cryo-EM structures of dimeric PAPP-A in its substrate-free form and in complex with a peptide substrate, which combined with biochemical assays provide a mechanism for PAPP-A substrate binding and selectivity.
- Russell A. Judge
- , Janani Sridar
- & Qi Hao
-
Article
| Open Access Protein shape sampled by ion mobility mass spectrometry consistently improves protein structure prediction
Protein shape sampled by ion mobility mass spectrometry consistently improves protein structure prediction
Collision cross sections (CCS) from ion mobility mass spectrometry provide information about protein shape and size. Here, the authors develop an algorithm to predict CCS and integrate experimental ion mobility data into Rosetta-based molecular modelling to predict protein structures from sequence.
- SM Bargeen Alam Turzo
- , Justin T. Seffernick
- & Steffen Lindert
-
Article
| Open Access Many dissimilar NusG protein domains switch between α-helix and β-sheet folds
Many dissimilar NusG protein domains switch between α-helix and β-sheet folds
Folded proteins are composed of secondary structures, α-helices and β-sheets, that are generally assumed to be stable. Here, the authors combine computational prediction with experimental validation to show that many sequence-diverse NusG protein domains switch completely from α-helix to β-sheet folds.
- Lauren L. Porter
- , Allen K. Kim
- & Marie-Paule Strub
-
Article
| Open Access AF2Complex predicts direct physical interactions in multimeric proteins with deep learning
AF2Complex predicts direct physical interactions in multimeric proteins with deep learning
Accurate descriptions of protein-protein interactions are essential for understanding biological systems. Here the authors present AF2Complex and show that application to the E. coli cytochrome biogenesis system I yields confident computational models for three sought-after assemblies.
- Mu Gao
- , Davi Nakajima An
- & Jeffrey Skolnick
-
Article
| Open Access Improved prediction of protein-protein interactions using AlphaFold2
Improved prediction of protein-protein interactions using AlphaFold2
Predicting the structure of protein complexes is extremely difficult. Here, authors apply AlphaFold2 with optimized multiple sequence alignments to model complexes of interacting proteins, enabling prediction of both if and how proteins interact with state-of-art accuracy.
- Patrick Bryant
- , Gabriele Pozzati
- & Arne Elofsson
-
Article
| Open Access Harnessing protein folding neural networks for peptide–protein docking
Harnessing protein folding neural networks for peptide–protein docking
AlphaFold2 has originally been developed to provide highly accurate predictions of protein monomer structures. Here, the authors present a simple adaptation of AlphaFold2 that enables structural modeling of peptide–protein complexes, and explore the underlying mechanisms and limitations of this approach.
- Tomer Tsaban
- , Julia K. Varga
- & Ora Schueler-Furman
-
Article
| Open Access DeepRank: a deep learning framework for data mining 3D protein-protein interfaces
DeepRank: a deep learning framework for data mining 3D protein-protein interfaces
The authors present DeepRank, a deep learning framework for the data mining of large sets of 3D protein-protein interfaces (PPI). They use DeepRank to address two challenges in structural biology: distinguishing biological versus crystallographic PPIs in crystal structures, and secondly the ranking of docking models.
- Nicolas Renaud
- , Cunliang Geng
- & Li C. Xue
-
Article
| Open Access Ensuring scientific reproducibility in bio-macromolecular modeling via extensive, automated benchmarks
Ensuring scientific reproducibility in bio-macromolecular modeling via extensive, automated benchmarks
Computational methods are becoming an increasingly important part of biological research. Using the Rosetta framework as an example, the authors demonstrate how community-driven development of computational methods can be done in a reproducible and reliable fashion.
- Julia Koehler Leman
- , Sergey Lyskov
- & Richard Bonneau
-
Article
| Open Access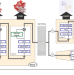 Mapping the glycosyltransferase fold landscape using interpretable deep learning
Mapping the glycosyltransferase fold landscape using interpretable deep learning
Glycosyltransferases (GT) are proteins that display extensive sequence and functional variation on a subset of 3D folds. Here, the authors use interpretable deep learning to predict 3D folds from sequence without the need for sequence alignment, which also enables the prediction of GTs with new folds.
- Rahil Taujale
- , Zhongliang Zhou
- & Natarajan Kannan
-
Article
| Open Access The antidepressant drug vilazodone is an allosteric inhibitor of the serotonin transporter
The antidepressant drug vilazodone is an allosteric inhibitor of the serotonin transporter
Vilazodone (VLZ) is a drug for the treatment of major depressive disorders that targets the serotonin transporter (SERT). Here, the authors combine pharmacology measurements and cryo-EM structural analysis to characterize VLZ binding to SERT and observe that VLZ exhibits non-competitive inhibition of serotonin transport and binds with nanomolar affinity to an allosteric site in SERT.
- Per Plenge
- , Dongxue Yang
- & Claus J. Loland
-
Article
| Open Access Improving fragment-based ab initio protein structure assembly using low-accuracy contact-map predictions
Improving fragment-based ab initio protein structure assembly using low-accuracy contact-map predictions
Predicting protein structure from sequence is still not possible for all proteins. Here, the authors introduce a method that integrates deep learning and information about protein co-evolution to guide the prediction of non-homologous protein structures with greater accuracy.
- S. M. Mortuza
- , Wei Zheng
- & Yang Zhang
-
Article
| Open Access flDPnn: Accurate intrinsic disorder prediction with putative propensities of disorder functions
flDPnn: Accurate intrinsic disorder prediction with putative propensities of disorder functions
The authors present flDPnn, a computational tool for disorder and disorder function predictions from protein sequences. flDPnn was assessed with the data from the “Critical Assessment of Protein Intrinsic Disorder Prediction” experiment and on an independent and low-similarity test dataset, which show that flDPnn offers accurate predictions of disorder, fully disordered proteins and four common disorder functions.
- Gang Hu
- , Akila Katuwawala
- & Lukasz Kurgan
-
Article
| Open Access Residue 6.43 defines receptor function in class F GPCRs
Residue 6.43 defines receptor function in class F GPCRs
The class Frizzled of G protein-coupled receptors (GPCRs) consist of ten Frizzled (FZD1-10) subtypes and Smoothened (SMO). Here the Schulte laboratory demonstrates that FZDs differ substantially from SMO in receptor activation-associated conformational changes, while SMO manifests a preference for a straight TM6, the TM6 of FZDs is kinked upon activation.
- Ainoleena Turku
- , Hannes Schihada
- & Gunnar Schulte
-
Article
| Open Access Structure-based protein function prediction using graph convolutional networks
Structure-based protein function prediction using graph convolutional networks
The rapid increase in the number of proteins in sequence databases and the diversity of their functions challenge computational approaches for automated function prediction. Here, the authors introduce DeepFRI, a Graph Convolutional Network for predicting protein functions by leveraging sequence features extracted from a protein language model and protein structures.
- Vladimir Gligorijević
- , P. Douglas Renfrew
- & Richard Bonneau
-
Article
| Open Access Pairing a high-resolution statistical potential with a nucleobase-centric sampling algorithm for improving RNA model refinement
Pairing a high-resolution statistical potential with a nucleobase-centric sampling algorithm for improving RNA model refinement
Predicting RNA structure from sequence is challenging due to the relative sparsity of experimentally-determined RNA 3D structures for model training. Here, the authors propose a way to incorporate knowledge on interactions at the atomic and base–base level to refine the prediction of RNA structures.
- Peng Xiong
- , Ruibo Wu
- & Yaoqi Zhou
-
Article
| Open Access CopulaNet: Learning residue co-evolution directly from multiple sequence alignment for protein structure prediction
CopulaNet: Learning residue co-evolution directly from multiple sequence alignment for protein structure prediction
Protein structure prediction is a challenge. A new deep learning framework, CopulaNet, is a major step forward toward end-to-end prediction of inter-residue distances and protein tertiary structures with improved accuracy and efficiency.
- Fusong Ju
- , Jianwei Zhu
- & Dongbo Bu
-
Article
| Open Access Structural and functional characterization of a putative de novo gene in Drosophila
Structural and functional characterization of a putative de novo gene in Drosophila
Previous work identified goddard as a putative de novo evolved gene in Drosophila melanogaster. Here, the authors characterize the structure and function of the Goddard protein in D. melanogaster, and they infer its ancestral and extant structures across the Drosophila genus.
- Andreas Lange
- , Prajal H. Patel
- & Erich Bornberg-Bauer
-
Article
| Open Access Large-scale discovery of protein interactions at residue resolution using co-evolution calculated from genomic sequences
Large-scale discovery of protein interactions at residue resolution using co-evolution calculated from genomic sequences
Our understanding of the residue-level details of protein interactions remains incomplete. Here, the authors show sequence coevolution can be used to infer interacting proteins with residue-level details, including predicting 467 interactions de novo in the Escherichia coli cell envelope proteome.
- Anna G. Green
- , Hadeer Elhabashy
- & Debora S. Marks
-
Article
| Open Access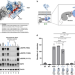 Structure–function analysis of oncogenic EGFR Kinase Domain Duplication reveals insights into activation and a potential approach for therapeutic targeting
Structure–function analysis of oncogenic EGFR Kinase Domain Duplication reveals insights into activation and a potential approach for therapeutic targeting
An EGFR mutant with kinase domain duplication (EGFR-KDD) was previously identified in an index patient, but the functional and therapeutic implications remain unclear. Here, the authors show that KDD occurs in other ErbB receptors in multiple cancers, and characterize the mechanism and inhibition of EGFR-KDD.
- Zhenfang Du
- , Benjamin P. Brown
- & Christine M. Lovly
-
Article
| Open Access Improved protein structure refinement guided by deep learning based accuracy estimation
Improved protein structure refinement guided by deep learning based accuracy estimation
Here the authors present DeepAccNet, a deep learning framework that estimates per-residue accuracy and residue-residue distance signed error in protein models, which are used to guide Rosetta protein structure refinement. Benchmarking suggests an improvement of accuracy prediction and refinement compared to other related state of the art methods.
- Naozumi Hiranuma
- , Hahnbeom Park
- & David Baker
-
Article
| Open Access Crystal structure of steroid reductase SRD5A reveals conserved steroid reduction mechanism
Crystal structure of steroid reductase SRD5A reveals conserved steroid reduction mechanism
Steroid 5α-reductase 2 (SRD5A2), a testosterone metabolism enzyme, is implicated in human disease. Structural and biochemical analyses of PbSRD5A, a bacterial homolog, reveal SRD5A2 substrate binding pocket and provide framework for the design of new drugs targeting this enzyme.
- Yufei Han
- , Qian Zhuang
- & Ruobing Ren
-
Article
| Open Access Heme-binding enables allosteric modulation in an ancient TIM-barrel glycosidase
Heme-binding enables allosteric modulation in an ancient TIM-barrel glycosidase
Family 1 glycosidases (GH1) are present in the three domains of life and share classical TIM-barrel fold. Structural and biochemical analyses of a resurrected ancestral GH1 enzyme reveal heme binding, not known in its modern descendants. Heme rigidifies the TIM-barrel and allosterically enhances catalysis.
- Gloria Gamiz-Arco
- , Luis I. Gutierrez-Rus
- & Jose M. Sanchez-Ruiz
-
Article
| Open Access Two distinct catalytic pathways for GH43 xylanolytic enzymes unveiled by X-ray and QM/MM simulations
Two distinct catalytic pathways for GH43 xylanolytic enzymes unveiled by X-ray and QM/MM simulations
Family 43 glycoside hydrolases (GH43) are involved in the breakdown of hemicellulose. Functional, structural and computational characterization of a GH43 enzyme, including a snapshot of an active Michaelis complex, reveal the hydrolysis mechanism and suggest two possible reaction pathways.
- Mariana A. B. Morais
- , Joan Coines
- & Mario T. Murakami
-
Article
| Open Access Integrative modeling of membrane-associated protein assemblies
Integrative modeling of membrane-associated protein assemblies
Most approaches for modeling the membrane protein complexes are not capable of incorporating the topological information provided by the membrane. Here authors present an integrative computational protocol for the modeling of membrane-associated protein assemblies, specifically complexes consisting of a membrane-embedded protein and a soluble partner.
- Jorge Roel-Touris
- , Brian Jiménez-García
- & Alexandre M. J. J. Bonvin
-
Article
| Open Access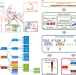 Inferring the molecular and phenotypic impact of amino acid variants with MutPred2
Inferring the molecular and phenotypic impact of amino acid variants with MutPred2
Identifying variants capable of causing genetic disease is challenging. The authors use semisupervised learning to predict pathogenic missense variants and their impacts on protein structure and function, enabling a molecular mechanism-driven approach to studying different types of human disease.
- Vikas Pejaver
- , Jorge Urresti
- & Predrag Radivojac
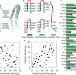
 Hairpin trimer transition state of amyloid fibril
Hairpin trimer transition state of amyloid fibril