Featured
-
-
Article
| Open Access Deep generative model embedding of single-cell RNA-Seq profiles on hyperspheres and hyperbolic spaces
Deep generative model embedding of single-cell RNA-Seq profiles on hyperspheres and hyperbolic spaces
Single-cell RNA-seq allows the study of tissues at cellular resolution. Here, the authors demonstrate how deep learning can be used to gain biological insight from such data by accounting for biological and technical variability. Data exploration is improved by accurately visualizing cells on an interactive 3D surface.
- Jiarui Ding
- & Aviv Regev
-
Article
| Open Access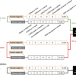 Learning a genome-wide score of human–mouse conservation at the functional genomics level
Learning a genome-wide score of human–mouse conservation at the functional genomics level
Understanding conserved functional genomic properties between human and mouse provides important context for mouse model studies. Here, the authors present a genome-wide conservation score integrating epigenomic, transcription factor binding, and transcriptomic data from mouse and human genomes.
- Soo Bin Kwon
- & Jason Ernst
-
Article
| Open Access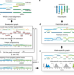 Integrative reconstruction of cancer genome karyotypes using InfoGenomeR
Integrative reconstruction of cancer genome karyotypes using InfoGenomeR
Karyotyping of cancer genomes at the base-level is technically challenging. Here, the authors introduce InfoGenomeR, an algorithm that can infer cancer genome karyotypes from whole-genome sequencing data, and test their model on breast, ovarian and brain cancer samples; and identify private and shared mutations between primary and metastatic cancer samples.
- Yeonghun Lee
- & Hyunju Lee
-
Article
| Open Access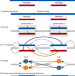 Extended haplotype-phasing of long-read de novo genome assemblies using Hi-C
Extended haplotype-phasing of long-read de novo genome assemblies using Hi-C
Methods to produce haplotype-resolved genome assemblies often rely on access to family trios. The authors present FALCON-Phase, a tool that combines ultra-long range Hi-C chromatin interaction data with a long read de novo assembly to extend haplotype phasing to the contig or scaffold level.
- Zev N. Kronenberg
- , Arang Rhie
- & Sarah B. Kingan
-
Article
| Open Access Genomic insights into the conservation status of the world’s last remaining Sumatran rhinoceros populations
Genomic insights into the conservation status of the world’s last remaining Sumatran rhinoceros populations
Highly endangered species like the Sumatran rhinoceros are at risk from inbreeding. Five historical and 16 modern genomes from across the species range show mutational load, but little evidence for local adaptation, suggesting that future inbreeding depression could be mitigated by assisted gene flow among populations.
- Johanna von Seth
- , Nicolas Dussex
- & Love Dalén
-
Article
| Open Access Protein design and variant prediction using autoregressive generative models
Protein design and variant prediction using autoregressive generative models
The ability to design functional sequences is central to protein engineering and biotherapeutics. Here the authors introduce a deep generative alignment-free model for sequence design applied to highly variable regions and design and test a diverse nanobody library with improved properties for selection experiments.
- Jung-Eun Shin
- , Adam J. Riesselman
- & Debora S. Marks
-
Article
| Open Access The VRNetzer platform enables interactive network analysis in Virtual Reality
The VRNetzer platform enables interactive network analysis in Virtual Reality
Data-rich networks can be difficult to interpret beyond a certain size. Here, the authors introduce a platform that uses virtual reality to allow the visual exploration of large networks, while interfacing with data repositories and other analytical methods to improve the interpretation of big data.
- Sebastian Pirch
- , Felix Müller
- & Jörg Menche
-
Article
| Open Access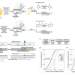 CRISPR-Cas9 cytidine and adenosine base editing of splice-sites mediates highly-efficient disruption of proteins in primary and immortalized cells
CRISPR-Cas9 cytidine and adenosine base editing of splice-sites mediates highly-efficient disruption of proteins in primary and immortalized cells
Base editors can inactivate splice sites or introduce stop codons into a gene sequence. Here the authors present SpliceR to design, rank, and test sgRNAs for efficient gene disruption in T cells.
- Mitchell G. Kluesner
- , Walker S. Lahr
- & Branden S. Moriarity
-
Article
| Open Access Leveraging community mortality indicators to infer COVID-19 mortality and transmission dynamics in Damascus, Syria
Leveraging community mortality indicators to infer COVID-19 mortality and transmission dynamics in Damascus, Syria
Reported COVID-19 mortality rates have been relatively low in Syria, but there has been concern about overwhelmed health systems. Here, the authors use community mortality indicators and estimate that <3% of COVID-19 deaths in Damascus were reported as of 2 September 2020.
- Oliver J. Watson
- , Mervat Alhaffar
- & Patrick Walker
-
Article
| Open Access Machine learning guided aptamer refinement and discovery
Machine learning guided aptamer refinement and discovery
Current aptamer discovery approaches are unable to probe the complete space of possible sequences. Here, the authors use machine learning to facilitate the development of DNA aptamers with improved binding affinities, and truncate them without significantly compromising binding affinity.
- Ali Bashir
- , Qin Yang
- & B. Scott Ferguson
-
Article
| Open Access An integrative analysis of the age-associated multi-omic landscape across cancers
An integrative analysis of the age-associated multi-omic landscape across cancers
Our understanding of the age-related molecular alterations in cancer is still limited. Here, the authors perform a pan-cancer analysis of age-associated genomic, transcriptomic, and epigenetic alterations, linking age-related gene expression changes to age-related DNA methylation alterations
- Kasit Chatsirisupachai
- , Tom Lesluyes
- & João Pedro de Magalhães
-
Article
| Open Access Genomic architecture and prediction of censored time-to-event phenotypes with a Bayesian genome-wide analysis
Genomic architecture and prediction of censored time-to-event phenotypes with a Bayesian genome-wide analysis
Few genome-wide association studies have explored the genetic architecture of age-of-onset for traits and diseases. Here, the authors develop a Bayesian approach to improve prediction in timing-related phenotypes and perform age-of-onset analyses across complex traits in the UK Biobank.
- Sven E. Ojavee
- , Athanasios Kousathanas
- & Matthew R. Robinson
-
Article
| Open Access Spatially interacting phosphorylation sites and mutations in cancer
Spatially interacting phosphorylation sites and mutations in cancer
Dysregulated phosphorylation is well-known in cancers, but it has largely been studied in isolation from mutations. Here the authors introduce HotPho, a tool that can discover spatial interactions between phosphosites and mutations, which are associated with activating mutation and genetic dependencies in cancer.
- Kuan-lin Huang
- , Adam D. Scott
- & Li Ding
-
Article
| Open Access Conserved long-range base pairings are associated with pre-mRNA processing of human genes
Conserved long-range base pairings are associated with pre-mRNA processing of human genes
Functional RNA secondary structure is important for the pre-mRNA processing including splicing, cleavage and polyadenylation, and RNA editing. Here the authors present a catalog of conserved long-range RNA structures in the human transcriptome by defining pairs of conserved complementary regions (PCCR) in pre-aligned evolutionarily conserved regions.
- Svetlana Kalmykova
- , Marina Kalinina
- & Dmitri Pervouchine
-
Article
| Open Access Detecting protein and DNA/RNA structures in cryo-EM maps of intermediate resolution using deep learning
Detecting protein and DNA/RNA structures in cryo-EM maps of intermediate resolution using deep learning
It is challenging to extract structural information from EM density maps at intermediate or low resolutions. Here, the authors present Emap2sec+, a program for detecting nucleotides and protein secondary structures in EM density maps at 5 to 10 Å resolution.
- Xiao Wang
- , Eman Alnabati
- & Daisuke Kihara
-
Article
| Open Access multiSLIDE is a web server for exploring connected elements of biological pathways in multi-omics data
multiSLIDE is a web server for exploring connected elements of biological pathways in multi-omics data
The integration and interpretation of different omics data types is an ongoing challenge for biologists. Here, the authors present a web-based, interactive tool called multiSLIDE for the visualization of protein, phosphoprotein, and RNA data presented as interlinked heatmaps.
- Soumita Ghosh
- , Abhik Datta
- & Hyungwon Choi
-
Article
| Open Access Design of multi-scale protein complexes by hierarchical building block fusion
Design of multi-scale protein complexes by hierarchical building block fusion
De novo design of self-assembling protein nanostructures and materials is of significant interest, however design of complex, multi-component assemblies is challenging. Here, the authors present a stepwise hierarchical approach to build such assemblies using helical repeat and helical bundle proteins as building blocks, and provide an in-depth structural characterization of the resulting assemblies.
- Yang Hsia
- , Rubul Mout
- & David Baker
-
Article
| Open Access An integrated multi-omics analysis identifies prognostic molecular subtypes of non-muscle-invasive bladder cancer
An integrated multi-omics analysis identifies prognostic molecular subtypes of non-muscle-invasive bladder cancer
Multiple molecular profiling methods are required to study urothelial non-muscle-invasive bladder cancer (NMIBC) due to its heterogeneity. Here the authors integrate multi-omics data of 834 NMIBC patients, identifying a molecular subgroup associated with multiple alterations and worse outcomes.
- Sia Viborg Lindskrog
- , Frederik Prip
- & Lars Dyrskjøt
-
Article
| Open Access Single cell regulatory landscape of the mouse kidney highlights cellular differentiation programs and disease targets
Single cell regulatory landscape of the mouse kidney highlights cellular differentiation programs and disease targets
Epigenetic and transcriptional dynamics are critical for both tissue homeostasis and injury response in the kidney. Leveraging a single cell multiomics atlas of the developing mouse kidney, the authors reveal key events in chromatin regulation and gene expression dynamics during postnatal development.
- Zhen Miao
- , Michael S. Balzer
- & Katalin Susztak
-
Article
| Open Access Democratising deep learning for microscopy with ZeroCostDL4Mic
Democratising deep learning for microscopy with ZeroCostDL4Mic
Deep learning methods show great promise for the analysis of microscopy images but there is currently an accessibility barrier to many users. Here the authors report a convenient entry-level deep learning platform that can be used at no cost: ZeroCostDL4Mic.
- Lucas von Chamier
- , Romain F. Laine
- & Ricardo Henriques
-
Article
| Open Access Defining super-enhancer landscape in triple-negative breast cancer by multiomic profiling
Defining super-enhancer landscape in triple-negative breast cancer by multiomic profiling
Triple-negative breast cancer (TNBC) is an aggressive breast cancer subtype with poor prognostic outcomes. Here the authors characterize super-enhancer heterogeneity and they identify genes that are specifically regulated by TNBC-specific super-enhancers, including FOXC1, MET and ANLN.
- Hao Huang
- , Jianyang Hu
- & Y. Rebecca Chin
-
Article
| Open Access Comprehensive omic characterization of breast cancer in Mexican-Hispanic women
Comprehensive omic characterization of breast cancer in Mexican-Hispanic women
Cancers in different populations have been shown to be genetically distinct. Here, the authors sequence breast cancers from Mexican-Hispanic patients and find that these patients have a higher percentage of Akt1 mutations compared to Caucasian and Asian populations, suggesting these are clinically actionable.
- Sandra L. Romero-Cordoba
- , Ivan Salido-Guadarrama
- & Alfredo Hidalgo-Miranda
-
Article
| Open Access Redundant and non-redundant cytokine-activated enhancers control Csn1s2b expression in the lactating mouse mammary gland
Redundant and non-redundant cytokine-activated enhancers control Csn1s2b expression in the lactating mouse mammary gland
Enhancers and promoters work together to actively regulate gene expression affecting several biological processes. Here, the authors provide molecular insights into the regulation of enhancers and super-enhancers in the Csn1s2b locus during lactation.
- Hye Kyung Lee
- , Michaela Willi
- & Lothar Hennighausen
-
Article
| Open Access Moss enables high sensitivity single-nucleotide variant calling from multiple bulk DNA tumor samples
Moss enables high sensitivity single-nucleotide variant calling from multiple bulk DNA tumor samples
The study of tumour heterogeneity can be improved by sequencing multiple samples, but currently available variant callers have not been tailored to integrate them. Here the authors present Moss, a tool that can leverage multiple samples to improve somatic variant calling in different cancers.
- Chuanyi Zhang
- , Mohammed El-Kebir
- & Idoia Ochoa
-
Article
| Open Access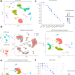 Single cell transcriptional and chromatin accessibility profiling redefine cellular heterogeneity in the adult human kidney
Single cell transcriptional and chromatin accessibility profiling redefine cellular heterogeneity in the adult human kidney
Single cell transcriptomic and epigenomic sequencing of human kidney highlight diverse cell types and states. These findings help characterize a novel population of injured proximal tubule cells and illustrate the power of multi-omic approaches to characterizing human tissue.
- Yoshiharu Muto
- , Parker C. Wilson
- & Benjamin D. Humphreys
-
Article
| Open Access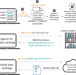 Go Get Data (GGD) is a framework that facilitates reproducible access to genomic data
Go Get Data (GGD) is a framework that facilitates reproducible access to genomic data
Modern biological research is complicated by the difficulty of collecting, transforming, annotating, and integrating datasets. Here, the authors present Go Get Data, a fast, reproducible approach to installing standardized data recipes, with an application to genomics data.
- Michael J. Cormier
- , Jonathan R. Belyeu
- & Aaron R. Quinlan
-
Article
| Open Access RA3 is a reference-guided approach for epigenetic characterization of single cells
RA3 is a reference-guided approach for epigenetic characterization of single cells
Methods for profiling differences between individual cells are constantly expanding. Here, the authors present a computational framework for the analysis of chromatin accessibility data at the single-cell level that takes into account previous knowledge and data-specific characteristics.
- Shengquan Chen
- , Guanao Yan
- & Zhixiang Lin
-
Article
| Open Access Uncovering transcriptional dark matter via gene annotation independent single-cell RNA sequencing analysis
Uncovering transcriptional dark matter via gene annotation independent single-cell RNA sequencing analysis
Conventional single-cell RNA sequencing analysis rely on genome annotations that may be incomplete or inaccurate especially for understudied organisms. Here the authors present a bioinformatic tool that leverages single-cell data to uncover biologically relevant transcripts beyond the best available genome annotation.
- Michael F. Z. Wang
- , Madhav Mantri
- & Iwijn De Vlaminck
-
Article
| Open Access Genetic evidence for the association between COVID-19 epidemic severity and timing of non-pharmaceutical interventions
Genetic evidence for the association between COVID-19 epidemic severity and timing of non-pharmaceutical interventions
Estimating the effects of non-pharmaceutical interventions for COVID-19 is challenging, partly due to variations in testing. Here, the authors use viral sequence data as an alternative means of inferring intervention effects, and show that delays in implementation resulted in more severe epidemics.
- Manon Ragonnet-Cronin
- , Olivia Boyd
- & Erik Volz
-
Article
| Open Access Learning cis-regulatory principles of ADAR-based RNA editing from CRISPR-mediated mutagenesis
Learning cis-regulatory principles of ADAR-based RNA editing from CRISPR-mediated mutagenesis
The RNA sequence and secondary structure regulate RNA editing by ADAR. Here the authors employ a CRISPR/Cas9-mediated saturation mutagenesis and machine learning to predict RNA editing efficiency of specific substrates.
- Xin Liu
- , Tao Sun
- & Jin Billy Li
-
Article
| Open Access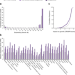 The landscape of molecular chaperones across human tissues reveals a layered architecture of core and variable chaperones
The landscape of molecular chaperones across human tissues reveals a layered architecture of core and variable chaperones
Tissue-specific differences in protein folding capacities are poorly understood. Here, the authors show that the human chaperone system consists of ubiquitous core chaperones and tissue-specific variable chaperones, perturbation of which leads to tissue-specific phenotypes.
- Netta Shemesh
- , Juman Jubran
- & Esti Yeger-Lotem
-
Article
| Open Access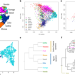 Genetic substructure and complex demographic history of South African Bantu speakers
Genetic substructure and complex demographic history of South African Bantu speakers
Despite linguistic and geographic diversity in South Eastern Bantu-speaking (SEB) groups of South Africa, genetic variation in these groups has not been investigated in depth. Here, the authors analyse genome-wide data from 5056 individuals, providing insights into demographic history across SEB groups.
- Dhriti Sengupta
- , Ananyo Choudhury
- & Michèle Ramsay
-
Article
| Open Access VESPER: global and local cryo-EM map alignment using local density vectors
VESPER: global and local cryo-EM map alignment using local density vectors
Here, the authors present VESPER, a program for EM density map search and alignment. Using benchmark datasets, they demonstrate that VESPER performs accurate global and local alignments and comparisons of EM maps.
- Xusi Han
- , Genki Terashi
- & Daisuke Kihara
-
Article
| Open Access Variable number tandem repeats mediate the expression of proximal genes
Variable number tandem repeats mediate the expression of proximal genes
Variable number tandem repeats (VNTRs) are implicated in human diseases yet have been difficult to analyse computationally. Here, the authors describe a neural network method, adVNTR-NN, that allows rapid and accurate genotyping of VNTRs from large whole genome sequencing datasets.
- Mehrdad Bakhtiari
- , Jonghun Park
- & Vineet Bafna
-
Article
| Open Access Identifying multiple sclerosis subtypes using unsupervised machine learning and MRI data
Identifying multiple sclerosis subtypes using unsupervised machine learning and MRI data
Multiple sclerosis is a heterogeneous progressive disease. Here, the authors use an unsupervised machine learning algorithm to determine multiple sclerosis subtypes, progression, and response to potential therapeutic treatments based on neuroimaging data.
- Arman Eshaghi
- , Alexandra L. Young
- & Olga Ciccarelli
-
Article
| Open Access Ontology-driven weak supervision for clinical entity classification in electronic health records
Ontology-driven weak supervision for clinical entity classification in electronic health records
In the electronic health record, using clinical notes to identify entities such as disorders and their temporality can inform many important analyses. Here, the authors present a framework for weakly supervised entity classification using medical ontologies and expert-generated rules.
- Jason A. Fries
- , Ethan Steinberg
- & Nigam H. Shah
-
Article
| Open Access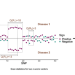 Detecting local genetic correlations with scan statistics
Detecting local genetic correlations with scan statistics
Genetic correlation analyses give insight on complex disease, yet are limited by oversimplification. Here, the authors present LOGODetect, a method using summary statistics from genome-wide association studies to identify genomic regions with correlation signals across multiple phenotypes.
- Hanmin Guo
- , James J. Li
- & Lin Hou
-
Article
| Open Access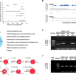 Presence of complete murine viral genome sequences in patient-derived xenografts
Presence of complete murine viral genome sequences in patient-derived xenografts
Patient-derived xenografts are widely used for drug development, but the impact of murine viral infection remains underexplored. Here, the authors demonstrate the extensive existence of murine viral sequences in patient-derived xenografts and significant expression change of crucial genes in samples with high virus load.
- Zihao Yuan
- , Xuejun Fan
- & W. Jim Zheng
-
Article
| Open Access Harnessing machine learning to guide phylogenetic-tree search algorithms
Harnessing machine learning to guide phylogenetic-tree search algorithms
Likelihood optimization in phylogenetic tree reconstruction is computationally intensive, especially as the number of sequences and taxa included increase. Here, Azouri et al. show how an artificial intelligence approach can reduce computational time without losing accuracy of tree inference.
- Dana Azouri
- , Shiran Abadi
- & Tal Pupko
-
Article
| Open Access Model-based assessment of replicability for genome-wide association meta-analysis
Model-based assessment of replicability for genome-wide association meta-analysis
In genome-wide association meta-analysis, it is often difficult to find an independent dataset of sufficient size to replicate associations. Here, the authors have developed MAMBA to calculate the probability of replicability based on consistency between datasets within the meta-analysis.
- Daniel McGuire
- , Yu Jiang
- & Dajiang J. Liu
-
Article
| Open Access The SARS-CoV-2 nucleocapsid protein is dynamic, disordered, and phase separates with RNA
The SARS-CoV-2 nucleocapsid protein is dynamic, disordered, and phase separates with RNA
SARS-CoV-2 nucleocapsid (N) protein is responsible for viral genome packaging. Here the authors employ single-molecule spectroscopy with all-atom simulations to provide the molecular details of N protein and show that it undergoes phase separation with RNA.
- Jasmine Cubuk
- , Jhullian J. Alston
- & Alex S. Holehouse
-
Article
| Open Access Implications of the school-household network structure on SARS-CoV-2 transmission under school reopening strategies in England
Implications of the school-household network structure on SARS-CoV-2 transmission under school reopening strategies in England
Many countries have closed schools as part of their COVID-19 response. Here, the authors model SARS-CoV-2 transmission on a network of schools and households in England, and find that risk of transmission between schools is lower if primary schools are open than if secondary schools are open.
- James D. Munday
- , Katharine Sherratt
- & Sebastian Funk
-
Article
| Open Access Hospital load and increased COVID-19 related mortality in Israel
Hospital load and increased COVID-19 related mortality in Israel
COVID-19 has caused many healthcare systems to become overwhelmed, potentially impacting patient care. Here, the authors show that COVID-19-related in-hospital mortality rates in Israel increased in periods of moderate or high hospital load, independent of patient characteristics.
- Hagai Rossman
- , Tomer Meir
- & Malka Gorfine
-
Article
| Open Access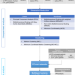 Nonlinear machine learning pattern recognition and bacteria-metabolite multilayer network analysis of perturbed gastric microbiome
Nonlinear machine learning pattern recognition and bacteria-metabolite multilayer network analysis of perturbed gastric microbiome
Drug use or bacterial infection can cause significant alterations of gastric microbiome. Here, the authors show how advanced pattern recognition by nonlinear machine intelligence can help disclose a bacteria-metabolite network which enlightens mechanisms behind such perturbations.
- Claudio Durán
- , Sara Ciucci
- & Carlo Vittorio Cannistraci
-
Article
| Open Access Model-based deep embedding for constrained clustering analysis of single cell RNA-seq data
Model-based deep embedding for constrained clustering analysis of single cell RNA-seq data
Clustering cells based on similarities in gene expression is the first step towards identifying cell types in scRNASeq data. Here the authors incorporate biological knowledge into the clustering step to facilitate the biological interpretability of clusters, and subsequent cell type identification.
- Tian Tian
- , Jie Zhang
- & Hakon Hakonarson
-
Article
| Open Access scGNN is a novel graph neural network framework for single-cell RNA-Seq analyses
scGNN is a novel graph neural network framework for single-cell RNA-Seq analyses
Single-cell RNA-Seq suffers from heterogeneity in sequencing sparsity and complex differential patterns in gene expression. Here, the authors introduce a graph neural network based on a hypothesis-free deep learning framework as an effective representation of gene expression and cell–cell relationships.
- Juexin Wang
- , Anjun Ma
- & Dong Xu
-
Article
| Open Access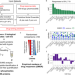 Drug ranking using machine learning systematically predicts the efficacy of anti-cancer drugs
Drug ranking using machine learning systematically predicts the efficacy of anti-cancer drugs
Artificial intelligence and machine learning promise to transform cancer therapies by accurately predicting the most appropriate drugs to treat individual patients. Here, the authors present an approach which uses omics data to produce ordered lists of drugs based on their effectiveness in decreasing cancer cell proliferation.
- Henry Gerdes
- , Pedro Casado
- & Pedro R. Cutillas
-
Article
| Open Access Predicting treatment response from longitudinal images using multi-task deep learning
Predicting treatment response from longitudinal images using multi-task deep learning
Radiographic imaging is routinely used to evaluate treatment response in solid tumors. Here, the authors present a multi-task deep learning approach that allows simultaneous tumor segmentation and response prediction from longitudinal images in a multi-center study on rectal cancer.
- Cheng Jin
- , Heng Yu
- & Ruijiang Li
-
Article
| Open Access A rationally engineered decoder of transient intracellular signals
A rationally engineered decoder of transient intracellular signals
Cells encode information by modulating signal dynamics. Here the authors developed a rapid prototyping tool, TopoDesign, to engineer a synthetic short-pulse decoder in yeast.
- Claude Lormeau
- , Fabian Rudolf
- & Jörg Stelling
Browse broader subjects
Browse narrower subjects
- Biochemical reaction networks
- Cellular signalling networks
- Classification and taxonomy
- Communication and replication
- Computational models
- Computational neuroscience
- Computational platforms and environments
- Data acquisition
- Data integration
- Data mining
- Data processing
- Data publication and archiving
- Databases
- Functional clustering
- Gene ontology
- Gene regulatory networks
- Genome informatics
- Hardware and infrastructure
- High-throughput screening
- Image processing
- Literature mining
- Machine learning
- Microarrays
- Network topology
- Phylogeny
- Power law
- Predictive medicine
- Probabilistic data networks
- Programming language
- Protein analysis
- Protein design
- Protein folding
- Protein function predictions
- Protein structure predictions
- Proteome informatics
- Quality control
- Scale invariance
- Sequence annotation
- Software
- Standards
- Statistical methods
- Virtual drug screening

 CopulaNet: Learning residue co-evolution directly from multiple sequence alignment for protein structure prediction
CopulaNet: Learning residue co-evolution directly from multiple sequence alignment for protein structure prediction