Article
|
Open Access
Featured
-
-
Article
| Open Access BASALT refines binning from metagenomic data and increases resolution of genome-resolved metagenomic analysis
BASALT refines binning from metagenomic data and increases resolution of genome-resolved metagenomic analysis
Binning is an essential step in genome-resolved metagenomic analysis in which assembled contigs originating from the same source population are clustered. However it is challenging, especially for low abundance microbial species. Here the authors introduce a toolkit that integrates multiple prominent binning tools and AI for efficient and high-resolution recovery of non-redundant bins from short- and long-read metagenomic sequencing datasets.
- Zhiguang Qiu
- , Li Yuan
- & Ke Yu
-
Article
| Open Access Enabling large-scale screening of Barrett’s esophagus using weakly supervised deep learning in histopathology
Enabling large-scale screening of Barrett’s esophagus using weakly supervised deep learning in histopathology
Diagnosis of Barrett’s esophagus depends on pathologist assessment of stained slides. Here, the authors utilise a deep learning approach to prioritize potential cases using diagnostic labels in two datasets, with the aim to improve Barrett’s screening capacity.
- Kenza Bouzid
- , Harshita Sharma
- & Javier Alvarez-Valle
-
Article
| Open Access AlphaPept: a modern and open framework for MS-based proteomics
AlphaPept: a modern and open framework for MS-based proteomics
Mass spectrometry-based proteomics faces the challenge of processing vast data amounts. Here, the authors introduce AlphaPept, an open-source, Python-based framework that offers high speed analysis and easy integration for large-scale proteome analysis.
- Maximilian T. Strauss
- , Isabell Bludau
- & Matthias Mann
-
Article
| Open Access Drivers and impact of the early silent invasion of SARS-CoV-2 Alpha
Drivers and impact of the early silent invasion of SARS-CoV-2 Alpha
The SARS-CoV-2 Alpha variant of concern emerged in the UK in late 2020 but spread internationally before it was detected. Here, the authors reconstruct the dynamics of dissemination of this variant out of the UK by combining extent of genomic sequencing, travel volume, and local epidemic dynamics in a Bayesian model.
- Benjamin Faucher
- , Chiara E. Sabbatini
- & Chiara Poletto
-
Article
| Open Access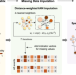 Interrogations of single-cell RNA splicing landscapes with SCASL define new cell identities with physiological relevance
Interrogations of single-cell RNA splicing landscapes with SCASL define new cell identities with physiological relevance
RNA splicing serves as a critical layer of gene expression regulation. Here, authors introduce SCASL for investigating the heterogeneity of RNA splicing landscapes at single-cell resolution, offering a novel scheme for classifying cell identities with physiological relevance.
- Xianke Xiang
- , Yao He
- & Xuerui Yang
-
Article
| Open Access The defensome of complex bacterial communities
The defensome of complex bacterial communities
Bacteria have evolved numerous innate and adaptive defence mechanisms. Here, Beavogui et al characterise the impact of biogeography, genetic mobility, and clustering in defense islands, on the defence systems of soil, marine, and human gut bacterial populations genomes.
- Angelina Beavogui
- , Auriane Lacroix
- & Pedro H. Oliveira
-
Article
| Open Access Viscosity-dependent control of protein synthesis and degradation
Viscosity-dependent control of protein synthesis and degradation
Xenopus egg extracts constitute a cell-like system for studying biochemical reactions. Here Chen and co-workers show that extract protein synthesis and degradation are differently affected by cytoplasmic concentration and viscosity.
- Yuping Chen
- , Jo-Hsi Huang
- & James E. Ferrell Jr.
-
Article
| Open Access Machine learning predictor PSPire screens for phase-separating proteins lacking intrinsically disordered regions
Machine learning predictor PSPire screens for phase-separating proteins lacking intrinsically disordered regions
Here the authors report a machine learning model, PSPire, which integrates both residue-level and structure-level features and outperforms tools in identifying phase-separating proteins lacking intrinsically disordered regions.
- Shuang Hou
- , Jiaojiao Hu
- & Yong Zhang
-
Article
| Open Access DeepETPicker: Fast and accurate 3D particle picking for cryo-electron tomography using weakly supervised deep learning
DeepETPicker: Fast and accurate 3D particle picking for cryo-electron tomography using weakly supervised deep learning
Picking particles of biological macromolecules is critical for solving their structures in situ using cryo-electron tomograms. Here, authors develop DeepETPicker, a deep learning-based tool for fast, accurate, and automated picking of three-dimensional particles.
- Guole Liu
- , Tongxin Niu
- & Ge Yang
-
Article
| Open Access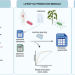 bacLIFE: a user-friendly computational workflow for genome analysis and prediction of lifestyle-associated genes in bacteria
bacLIFE: a user-friendly computational workflow for genome analysis and prediction of lifestyle-associated genes in bacteria
Many bacteria live in close association with eukaryotic hosts, exhibiting detrimental, neutral or beneficial effects on host growth and health. Here, the authors present a streamlined computational workflow for bacterial genome annotation, large-scale comparative genomics, and prediction of genes potentially involved in niche adaptation.
- Guillermo Guerrero-Egido
- , Adrian Pintado
- & Víctor J. Carrión
-
Article
| Open Access Biosensor and machine learning-aided engineering of an amaryllidaceae enzyme
Biosensor and machine learning-aided engineering of an amaryllidaceae enzyme
Amaryllidaceae alkaloids, such as the Alzheimer’s medication galantamine, are currently extracted from low-yielding daffodils. Here, authors pair biosensor-assisted screening with machine learning-guided protein design to rapidly engineer an improved Amaryllidaceae enzyme in a microbial host.
- Simon d’Oelsnitz
- , Daniel J. Diaz
- & Andrew D. Ellington
-
Article
| Open Access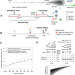 Evolving copy number gains promote tumor expansion and bolster mutational diversification
Evolving copy number gains promote tumor expansion and bolster mutational diversification
Understanding the timing and fitness of somatic copy number alterations (SCNAs) in cancer would shed light on cancer progression and evolution. Here, the authors develop Butte, a computational framework to estimate the timing of clonal SCNAs that encompass multiple gains, and apply it on whole-genome sequencing data from 184 samples.
- Zicheng Wang
- , Yunong Xia
- & Ruping Sun
-
Article
| Open Access Deep learning model for personalized prediction of positive MRSA culture using time-series electronic health records
Deep learning model for personalized prediction of positive MRSA culture using time-series electronic health records
Identification of patients at high risk of methicillin-resistant Staphylococcus aureus (MRSA) infection could improve treatment outcomes by optimising antimicrobial therapy. Here the authors develop a deep learning model that uses electronic health record data from the United States to predict MRSA culture positivity.
- Masayuki Nigo
- , Laila Rasmy
- & Degui Zhi
-
Article
| Open Access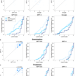 Systematic analysis of ChatGPT, Google search and Llama 2 for clinical decision support tasks
Systematic analysis of ChatGPT, Google search and Llama 2 for clinical decision support tasks
People will likely use ChatGPT to seek health advice. Here, the authors show promising performance of ChatGPT and open source models, but a lack of high accuracy considering medical question answering. Improvements are expected over time via domain-specific finetuning and integration of regulations.
- Sarah Sandmann
- , Sarah Riepenhausen
- & Julian Varghese
-
Article
| Open Access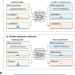 Riboformer: a deep learning framework for predicting context-dependent translation dynamics
Riboformer: a deep learning framework for predicting context-dependent translation dynamics
Riboformer is a deep learning-based framework that predicts changes in translation dynamics with codon-level precision. It corrects experimental artifacts in ribosome profiling data and identifies sequences causing ribosome stalling.
- Bin Shao
- , Jiawei Yan
- & Allen R. Buskirk
-
Article
| Open Access Machine learning-aided design and screening of an emergent protein function in synthetic cells
Machine learning-aided design and screening of an emergent protein function in synthetic cells
Here, the authors introduce a pipeline to screen machine learning generated variants of a protein that forms intracellular spatiotemporal patterns in E. coli, demonstrating the best variants can substitute the wildtype gene.
- Shunshi Kohyama
- , Béla P. Frohn
- & Petra Schwille
-
Article
| Open Access Enhancing the fairness of AI prediction models by Quasi-Pareto improvement among heterogeneous thyroid nodule population
Enhancing the fairness of AI prediction models by Quasi-Pareto improvement among heterogeneous thyroid nodule population
Artificial Intelligence (AI) models for medical diagnosis often face challenges of generalizability and fairness. Here, the authors show that the Quasi-Pareto Improvement approach is widely applicable to improving AI models among less-prevalent subgroups, promoting equitable healthcare outcomes.
- Siqiong Yao
- , Fang Dai
- & Hui Lu
-
Article
| Open Access A coarse-grained bacterial cell model for resource-aware analysis and design of synthetic gene circuits
A coarse-grained bacterial cell model for resource-aware analysis and design of synthetic gene circuits
Competition for the host cell’s resources influences synthetic biology circuit behavior. Here the authors present an E. coli cell model that combines insights into bacterial resource allocation with a simplified model of competition, facilitating resource-aware circuit design.
- Kirill Sechkar
- , Harrison Steel
- & Guy-Bart Stan
-
Article
| Open Access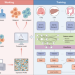 Domain generalization enables general cancer cell annotation in single-cell and spatial transcriptomics
Domain generalization enables general cancer cell annotation in single-cell and spatial transcriptomics
Efficient and accurate annotation of malignant cells is crucial for single-cell and spatial transcriptomics in cancer. Here, the authors develop Cancer-Finder, a deep-learning algorithm that can identify malignant cells in cancer single-cell and spatial transcriptomics data with speed and precision.
- Zhixing Zhong
- , Junchen Hou
- & Jia Song
-
Article
| Open Access Widespread stable noncanonical peptides identified by integrated analyses of ribosome profiling and ORF features
Widespread stable noncanonical peptides identified by integrated analyses of ribosome profiling and ORF features
By developing computational algorithms, the authors annotated translated open reading frames in five eukaryotes and found many stable peptides are encoded by putative ‘noncoding’ regions of genomes.
- Haiwang Yang
- , Qianru Li
- & Zhe Ji
-
Article
| Open Access Empirical data drift detection experiments on real-world medical imaging data
Empirical data drift detection experiments on real-world medical imaging data
Data drift is the systematic change in the underlying distribution of input features in prediction models, and can cause deterioration in model performance. Here, the authors highlight the importance of detecting data drift in clinical settings and evaluate methods for detecting drift in medical image data.
- Ali Kore
- , Elyar Abbasi Bavil
- & Mohamed Abdalla
-
Article
| Open Access Drug target prediction through deep learning functional representation of gene signatures
Drug target prediction through deep learning functional representation of gene signatures
Large-scale OMICs investigations of biological systems can be used to predict functional relationships between compounds, genes and proteins. Here, the authors develop a deep learning-based approach that significantly increases the number of high-quality compound-target predictions relative to existing methods.
- Hao Chen
- , Frederick J. King
- & Yingyao Zhou
-
Article
| Open Access A framework for evaluating clinical artificial intelligence systems without ground-truth annotations
A framework for evaluating clinical artificial intelligence systems without ground-truth annotations
Estimating the performance of clinical AI systems on data in the wild is complicated by distribution shift and the absence of ground-truth annotations. Here, we introduce SUDO, a framework for more reliably evaluating AI systems on data in the wild.
- Dani Kiyasseh
- , Aaron Cohen
- & Nicholas Altieri
-
Article
| Open Access Prediction of plasma ctDNA fraction and prognostic implications of liquid biopsy in advanced prostate cancer
Prediction of plasma ctDNA fraction and prognostic implications of liquid biopsy in advanced prostate cancer
Metastatic castration-resistant prostate cancer is a highly aggressive disease, with a variable response to treatment. Here, the authors validate ctDNA fraction as a poor prognostic factor and develop a model to predict whether patients harbor sufficient ctDNA for informative blood-based genotyping.
- Nicolette M. Fonseca
- , Corinne Maurice-Dror
- & Alexander W. Wyatt
-
Article
| Open Access Functional regulation of aquaporin dynamics by lipid bilayer composition
Functional regulation of aquaporin dynamics by lipid bilayer composition
Membrane proteins depend on their lipid environments. Using aquaporin as a model, the authors show that the choice of lipid bilayer fundamentally affects membrane protein structure, thermodynamics, kinetic, and function, even to the point of lipid-based inhibition.
- Anh T. P. Nguyen
- , Austin T. Weigle
- & Diwakar Shukla
-
Article
| Open Access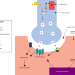 Rare disease research workflow using multilayer networks elucidates the molecular determinants of severity in Congenital Myasthenic Syndromes
Rare disease research workflow using multilayer networks elucidates the molecular determinants of severity in Congenital Myasthenic Syndromes
Congenital myasthenic syndromes are rare inherited neuromuscular disorders. Here, the authors attempt to explain diverse disease severity seen in 20 patients with shared CHRNE gene mutations with a multilayer network analysis that identifies individual-level impairments at the neuromuscular junction.
- Iker Núñez-Carpintero
- , Maria Rigau
- & Alfonso Valencia
-
Article
| Open Access Automatic data-driven design and 3D printing of custom ocular prostheses
Automatic data-driven design and 3D printing of custom ocular prostheses
Manual processes to produce ocular prostheses are time-consuming and yield varying quality. Here, authors present an automatic digital end-to-end process for custom ocular prostheses. It creates shape and appearance from image data of an OCT device and produces them using a full-colour 3D printer.
- Johann Reinhard
- , Philipp Urban
- & Mandeep S. Sagoo
-
Article
| Open Access A release of local subunit conformational heterogeneity underlies gating in a muscle nicotinic acetylcholine receptor
A release of local subunit conformational heterogeneity underlies gating in a muscle nicotinic acetylcholine receptor
Authors show that agonist binding to the muscle acetylcholine receptor releases local conformational heterogeneity transitioning all subunits into a symmetric open state. A release of conformational heterogeneity underlies allosteric communication.
- Mackenzie J. Thompson
- , Farid Mansoub Bekarkhanechi
- & John E. Baenziger
-
Article
| Open Access Statistical method scDEED for detecting dubious 2D single-cell embeddings and optimizing t-SNE and UMAP hyperparameters
Statistical method scDEED for detecting dubious 2D single-cell embeddings and optimizing t-SNE and UMAP hyperparameters
2D visualisation of single-cell data is highly impacted by the hyperparameter setting of the 2D embedding method, such as t-SNE and UMAP. Here, authors develop a statistical method scDEED to detect dubious cell embeddings and optimise the hyperparameter setting for trustworthy visualisation.
- Lucy Xia
- , Christy Lee
- & Jingyi Jessica Li
-
Article
| Open Access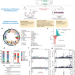 A distinct class of pan-cancer susceptibility genes revealed by an alternative polyadenylation transcriptome-wide association study
A distinct class of pan-cancer susceptibility genes revealed by an alternative polyadenylation transcriptome-wide association study
Alternative polyadenylation (APA) can play a key role in cancer initiation and progression. Here, the authors conducted a comprehensive pan-cancer APA TWAS analysis and discovered a distinct class of APA-mediated cancer susceptibility genes across 22 cancer types.
- Hui Chen
- , Zeyang Wang
- & Lei Li
-
Article
| Open Access SEMORE: SEgmentation and MORphological fingErprinting by machine learning automates super-resolution data analysis
SEMORE: SEgmentation and MORphological fingErprinting by machine learning automates super-resolution data analysis
There is a lack of universal tools to analyse protein assemblies and quantify underlying structures in single-molecule localization microscopy. Here, the authors present SEMORE, a semi-automatic machine learning framework for system- and input-dependent analysis of super-resolution data.
- Steen W. B. Bender
- , Marcus W. Dreisler
- & Nikos S. Hatzakis
-
Article
| Open Access Transfer learning with graph neural networks for improved molecular property prediction in the multi-fidelity setting
Transfer learning with graph neural networks for improved molecular property prediction in the multi-fidelity setting
Modern molecular discovery processes generate millions of measurements at different quality levels. Here, the authors develop a new deep learning method for transfer learning from low-cost and abundant data to enhance the efficiency of drug discovery.
- David Buterez
- , Jon Paul Janet
- & Pietro Lió
-
Article
| Open Access Rapid deep learning-assisted predictive diagnostics for point-of-care testing
Rapid deep learning-assisted predictive diagnostics for point-of-care testing
A key aim in the development of diagnostic assays is improving diagnostic speed while maintaining sensitivity. Here the authors report an approach for the rapid and accurate analysis of lateral flow tests, which integrates time-series deep learning and AI verification, achieving a diagnostic time of 1-2 minutes.
- Seungmin Lee
- , Jeong Soo Park
- & Jeong Hoon Lee
-
Article
| Open Access Metabolomic machine learning predictor for diagnosis and prognosis of gastric cancer
Metabolomic machine learning predictor for diagnosis and prognosis of gastric cancer
Gastric cancer detection by endoscopy is intrusive and time-consuming, and early detection is key to improving survival. Here, the authors propose a metabolite-based model to enable early detection.
- Yangzi Chen
- , Bohong Wang
- & Zeping Hu
-
Article
| Open Access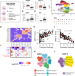 Complex regulatory networks influence pluripotent cell state transitions in human iPSCs
Complex regulatory networks influence pluripotent cell state transitions in human iPSCs
Stem cells exist in vitro in a spectrum of interconvertible pluripotent states. Here, authors show that pluripotency and self-renewal processes have a high level of regulatory complexity and suggest that genetic factors contribute to cell state transitions in human iPSC lines.
- Timothy D. Arthur
- , Jennifer P. Nguyen
- & Kelly A. Frazer
-
Article
| Open Access scCASE: accurate and interpretable enhancement for single-cell chromatin accessibility sequencing data
scCASE: accurate and interpretable enhancement for single-cell chromatin accessibility sequencing data
Single-cell chromatin accessibility sequencing (scCAS) data suffers from high sparsity and dimensionality. Here, authors propose an accurate and interpretable computational framework for enhancing scCAS data that considers cell-to-cell similarity.
- Songming Tang
- , Xuejian Cui
- & Shengquan Chen
-
Article
| Open Access Protein design using structure-based residue preferences
Protein design using structure-based residue preferences
Recent protein design methods rely on large neural networks, yet it is unclear which dependencies are critical for determining function. Here, authors show that learning the per residue mutation preferences, without considering interactions, enables design of functional and diverse protein variants.
- David Ding
- , Ada Y. Shaw
- & Debora S. Marks
-
Article
| Open Access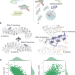 Design of target specific peptide inhibitors using generative deep learning and molecular dynamics simulations
Design of target specific peptide inhibitors using generative deep learning and molecular dynamics simulations
Here the authors report a computational approach which integrates deep learning and structural modelling to design target-specific peptides. They apply this to β-catenin and NF-κB essential modulator, resulting in improved binding, highlighting the efficacy of this strategy.
- Sijie Chen
- , Tong Lin
- & Xiaolin Cheng
-
Article
| Open Access Accurate global and local 3D alignment of cryo-EM density maps using local spatial structural features
Accurate global and local 3D alignment of cryo-EM density maps using local spatial structural features
Density map alignment is a fundamental step in Cryo-EM data postprocessing. Here, authors propose an accurate global and local density map alignment method using local density features.
- Bintao He
- , Fa Zhang
- & Renmin Han
-
Article
| Open Access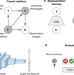 Learning representations for image-based profiling of perturbations
Learning representations for image-based profiling of perturbations
Assessing cell phenotypes in image-based assays requires solid computational methods for transforming images into quantitative data. Here, the authors present a strategy for learning representations of treatment effects from high-throughput imaging, following a causal interpretation.
- Nikita Moshkov
- , Michael Bornholdt
- & Juan C. Caicedo
-
Article
| Open Access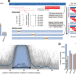 Efficient encoding of large antigenic spaces by epitope prioritization with Dolphyn
Efficient encoding of large antigenic spaces by epitope prioritization with Dolphyn
Profiling antibody responses to vast antigenic spaces has been challenging using programmable phage display (PhIP-Seq). Here, authors develop a methodology for compressing large proteomic spaces and have discovered human antibodies targeting gut bacteria-infecting phages.
- Anna-Maria Liebhoff
- , Thiagarajan Venkataraman
- & H. Benjamin Larman
-
Article
| Open Access Multi-omics analysis in human retina uncovers ultraconserved cis-regulatory elements at rare eye disease loci
Multi-omics analysis in human retina uncovers ultraconserved cis-regulatory elements at rare eye disease loci
Ultraconserved non-coding elements (UCNEs) can regulate developmental gene expression. Retinal multi-omics data integration revealed UCNEs to be candidate cis-regulatory elements during retinal development, which may be implicated in rare eye diseases.
- Victor Lopez Soriano
- , Alfredo Dueñas Rey
- & Elfride De Baere
-
Article
| Open Access Large language models streamline automated machine learning for clinical studies
Large language models streamline automated machine learning for clinical studies
A knowledge gap persists between machine learning developers and clinicians. Here, the authors show that the Advanced Data Analysis extension of ChatGPT could bridge this gap and simplify complex data analyses, making them more accessible to clinicians.
- Soroosh Tayebi Arasteh
- , Tianyu Han
- & Sven Nebelung
-
Article
| Open Access Iterative design of training data to control intricate enzymatic reaction networks
Iterative design of training data to control intricate enzymatic reaction networks
Kinetic modeling of in vitro enzymatic reaction networks (ERNs) is severely hampered by the lack of training data. Here, authors introduce a methodology that combines an active learning-like approach and flow chemistry to create optimized datasets for an intricate ERN.
- Bob van Sluijs
- , Tao Zhou
- & Wilhelm T. S. Huck
-
Article
| Open Access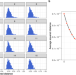 Imputation of plasma lipid species to facilitate integration of lipidomic datasets
Imputation of plasma lipid species to facilitate integration of lipidomic datasets
Advancements in plasma lipidomic profiling increase specificity of measurements but pose challenges in aligning datasets created at different times or platforms. Here the authors present a predictive framework for harmonising such datasets with different levels of granularity in their lipid measurements.
- Aleksandar Dakic
- , Jingqin Wu
- & Peter J. Meikle
-
Article
| Open Access Unsupervised classification of brain-wide axons reveals the presubiculum neuronal projection blueprint
Unsupervised classification of brain-wide axons reveals the presubiculum neuronal projection blueprint
The classification of different types of neurons has been a long-standing challenge in neuroscience. Here, the authors present a strategy to quantify all statistically distinct axonal patterns from a brain region based on their anatomical targeting, with this projection-driven neuron classification informing the functional architecture of the circuit.
- Diek W. Wheeler
- , Shaina Banduri
- & Giorgio A. Ascoli
-
Article
| Open Access An agricultural digital twin for mandarins demonstrates the potential for individualized agriculture
An agricultural digital twin for mandarins demonstrates the potential for individualized agriculture
A digital twin represents a real world object using available data. Here, the authors develop a digital twin for mandaring orchards in Jeju island showing the value of individualized agriculture to predict fruit quality at tree level.
- Steven Kim
- & Seong Heo
-
Article
| Open Access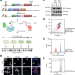 A mutational atlas for Parkin proteostasis
A mutational atlas for Parkin proteostasis
Gene variants can affect folding and stability of the encoded protein. Here, the authors apply deep mutational scanning to provide genotype-phenotype information for 99% of the possible PRKN variants and reveal mechanistic details on how some variants cause loss-of-function and Parkinsons disease.
- Lene Clausen
- , Vasileios Voutsinos
- & Rasmus Hartmann-Petersen
-
Article
| Open Access Recurrent evolutionary switches of mitochondrial cytochrome c maturation systems in Archaeplastida
Recurrent evolutionary switches of mitochondrial cytochrome c maturation systems in Archaeplastida
Cytochrome c maturation (CCM) is the process of covalent attachment of a heme group to the conserved cysteines to form the holocytochrome. Here, the authors report that the non-adaptive convergent evolution at the pathway level leads to mosaic distribution of CCM systems I and III among Archaeplastida species.
- Huang Li
- , Soujanya Akella
- & Jeffrey P. Mower
Browse broader subjects
Browse narrower subjects
- Biochemical reaction networks
- Cellular signalling networks
- Classification and taxonomy
- Communication and replication
- Computational models
- Computational neuroscience
- Computational platforms and environments
- Data acquisition
- Data integration
- Data mining
- Data processing
- Data publication and archiving
- Databases
- Functional clustering
- Gene ontology
- Gene regulatory networks
- Genome informatics
- Hardware and infrastructure
- High-throughput screening
- Image processing
- Literature mining
- Machine learning
- Microarrays
- Network topology
- Phylogeny
- Power law
- Predictive medicine
- Probabilistic data networks
- Programming language
- Protein analysis
- Protein design
- Protein folding
- Protein function predictions
- Protein structure predictions
- Proteome informatics
- Quality control
- Scale invariance
- Sequence annotation
- Software
- Standards
- Statistical methods
- Virtual drug screening

 Systematic review and meta-analysis for a Global Patient co-Owned Cloud (GPOC)
Systematic review and meta-analysis for a Global Patient co-Owned Cloud (GPOC)