Featured
-
-
Article
| Open Access Rare variant phasing and haplotypic expression from RNA sequencing with phASER
Rare variant phasing and haplotypic expression from RNA sequencing with phASER
Genome interpretation and analysis of allelic activity requires appropriate haplotype phasing. Here the authors present phASER, a fast and accurate method for variant phrasing from RNA-seq and genome sequencing data.
- Stephane E. Castel
- , Pejman Mohammadi
- & Tuuli Lappalainen
-
Article
| Open Access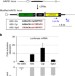 Sequences flanking the core-binding site modulate glucocorticoid receptor structure and activity
Sequences flanking the core-binding site modulate glucocorticoid receptor structure and activity
To modulate gene expression, the glucocorticoid receptor binds to response elements (RE) that vary in sequence. Here, the authors show that RE sequences can modulate glucocorticoid receptor structure and activity, which might provide regulatory specificity towards individual target genes.
- Stefanie Schöne
- , Marcel Jurk
- & Sebastiaan H. Meijsing
-
Article
| Open Access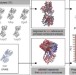 Prediction and validation of protein intermediate states from structurally rich ensembles and coarse-grained simulations
Prediction and validation of protein intermediate states from structurally rich ensembles and coarse-grained simulations
Protein conformational changes are key to a wide range of cellular functions but remain difficult to access experimentally. Here the authors describe eBDIMS, a novel approach to predict intermediates observed in structural transition pathways from experimental ensembles.
- Laura Orellana
- , Ozge Yoluk
- & Erik Lindahl
-
Article
| Open Access Prediction of allosteric sites and mediating interactions through bond-to-bond propensities
Prediction of allosteric sites and mediating interactions through bond-to-bond propensities
Allostery is a key molecular mechanism underpinning control and modulation in a variety of cellular processes. Here, the authors present a method that can be used to predict allosteric sites and the mediating interactions that connect them to the active site of the protein.
- B. R. C. Amor
- , M. T. Schaub
- & M. Barahona
-
Article
| Open Access Crowdsourced assessment of common genetic contribution to predicting anti-TNF treatment response in rheumatoid arthritis
Crowdsourced assessment of common genetic contribution to predicting anti-TNF treatment response in rheumatoid arthritis
Rheumatoid arthritis patients respond differently to anti-TNF treatment. Using community-based challenge, the authors show that currently available data does not reveal meaningful genetic predictors of response to anti-TNF therapy, thus confirming clinical observations.
- Solveig K. Sieberts
- , Fan Zhu
- & Lara M. Mangravite
-
Article
| Open Access Clonal haematopoiesis harbouring AML-associated mutations is ubiquitous in healthy adults
Clonal haematopoiesis harbouring AML-associated mutations is ubiquitous in healthy adults
Clonal haematopoiesis has been thought to occur in less than 10% of individuals younger than 70 years old. Here, the authors use an error corrected next-generation sequencing method to find clonal haematopoiesis in the peripheral blood of 19 of 20 healthy 50–70 year old individuals.
- Andrew L. Young
- , Grant A. Challen
- & Todd E. Druley
-
Article
| Open Access reChIP-seq reveals widespread bivalency of H3K4me3 and H3K27me3 in CD4+ memory T cells
reChIP-seq reveals widespread bivalency of H3K4me3 and H3K27me3 in CD4+ memory T cells
Co-localizing chromatin modifications and regulators can exert a combinatorial effect on chromatin structure and function. Here the authors describe reChIP-seq and normR to identify co-localizing proteins in an unbiased genome-wide manner.
- Sarah Kinkley
- , Johannes Helmuth
- & Ho-Ryun Chung
-
Article
| Open Access Extension of human lncRNA transcripts by RACE coupled with long-read high-throughput sequencing (RACE-Seq)
Extension of human lncRNA transcripts by RACE coupled with long-read high-throughput sequencing (RACE-Seq)
Long non-coding RNAs are increasingly recognised to be important factors in regulating cellular processes and comprise a large faction of the transcriptome, however most are uncharacterised. Here the authors present RACE-Seq, a tool to improve and extend the annotation of low-expression transcripts.
- Julien Lagarde
- , Barbara Uszczynska-Ratajczak
- & Jennifer Harrow
-
Article
| Open Access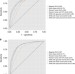 Predicting non-small cell lung cancer prognosis by fully automated microscopic pathology image features
Predicting non-small cell lung cancer prognosis by fully automated microscopic pathology image features
Diagnosis of lung cancer through manual histopathology evaluation is insufficient to predict patient survival. Here, the authors use computerized image processing to identify diagnostically relevant image features and use these features to distinguish lung cancer patients with different prognoses.
- Kun-Hsing Yu
- , Ce Zhang
- & Michael Snyder
-
Article
| Open Access Environmental change makes robust ecological networks fragile
Environmental change makes robust ecological networks fragile
Despite their complexity, ecological networks appear robust to species loss. Here, Strona and Lafferty use artificial life simulations and real-world data to show that such robustness applies to stable conditions, but can collapse when the environment changes.
- Giovanni Strona
- & Kevin D. Lafferty
-
Article
| Open Access Self-organized centripetal movement of corneal epithelium in the absence of external cues
Self-organized centripetal movement of corneal epithelium in the absence of external cues
The cornea is formed of cells that originate from the outer circle of stem cells and that move towards its centre. Here, the authors show that the movement pattern is self-organised, requiring no cues, and that stem cell leakage may account for the presence of stem cells at the centre of the cornea.
- Erwin P. Lobo
- , Naomi C. Delic
- & J. Guy Lyons
-
Article
| Open Access High-resolution imaging and computational analysis of haematopoietic cell dynamics in vivo
High-resolution imaging and computational analysis of haematopoietic cell dynamics in vivo
It is difficult to image haematopoietic stem cells (HSC) in their niche. Here, the authors present a new high-throughput computational approach to visualise HSCs in vivoat a high spatial and temporal resolution and also use a Msi2-reporter to label endogenous HSCs and progenitors, enabling cell tracking
- Claire S. Koechlein
- , Jeffrey R. Harris
- & Tannishtha Reya
-
Article
| Open Access Analysis of chromosomal aberrations and recombination by allelic bias in RNA-Seq
Analysis of chromosomal aberrations and recombination by allelic bias in RNA-Seq
Chromosomal aberrations can be detected by global gene expression analysis. Here, the authors report eSNP-Karyotyping, a new method that can detect chromosomal aberrations by measuring the ratio of expression between the two alleles without comparison to a matched diploid sample.
- Uri Weissbein
- , Maya Schachter
- & Nissim Benvenisty
-
Article
| Open Access Mpath maps multi-branching single-cell trajectories revealing progenitor cell progression during development
Mpath maps multi-branching single-cell trajectories revealing progenitor cell progression during development
The tools for constructing cell lineages from single cell data are limited. Here, the authors present Mpath: an algorithm to derive multi-branching trajectories using neighbourhood-based cell state transitions, and use this to predict developmental trajectories in dendritic and muscle cells.
- Jinmiao Chen
- , Andreas Schlitzer
- & Michael Poidinger
-
Article
| Open Access Information recovery from low coverage whole-genome bisulfite sequencing
Information recovery from low coverage whole-genome bisulfite sequencing
Here, Libertini and colleagues devise a computation tool that can analyze whole-genome bisulfite sequencing (WGBS) data to recover of ∼30% of the lost differential methylation position information. They use COMETgazer and COMETvintage to analyze 13 diffferent methylome data to demonstrate their performance.
- Emanuele Libertini
- , Simon C. Heath
- & Stephan Beck
-
Article
| Open Access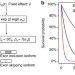 SURVIV for survival analysis of mRNA isoform variation
SURVIV for survival analysis of mRNA isoform variation
Clinical RNA-seq datasets can predict clinical outcomes. Here, Shen et al. report a statistical method for survival analysis of mRNA isoform variation using clinical RNA-seq datasets, and the identified isoform based survival predictors outperform gene expression based survival predictors using TCGA data on six cancer types.
- Shihao Shen
- , Yuanyuan Wang
- & Yi Xing
-
Article
| Open Access Synthetic mixed-signal computation in living cells
Synthetic mixed-signal computation in living cells
Digital and analogue gene circuits each have distinct advantages in natural and engineered cells. Here, Rubens et al. engineer synthetic gene circuits that implement mixed-signal digital and analogue computations in living cells.
- Jacob R. Rubens
- , Gianluca Selvaggio
- & Timothy K. Lu
-
Article
| Open Access Improving GENCODE reference gene annotation using a high-stringency proteogenomics workflow
Improving GENCODE reference gene annotation using a high-stringency proteogenomics workflow
Identifying and annotating functional elements in the human genome remains a challenging but important task. Here the authors propose a priority annotation score to rank identifications and suggest how proteogenomics evidence can be interpreted and what additional information substantiates protein-coding potential for annotation.
- James C. Wright
- , Jonathan Mudge
- & Jennifer Harrow
-
Article
| Open Access Mining 3D genome structure populations identifies major factors governing the stability of regulatory communities
Mining 3D genome structure populations identifies major factors governing the stability of regulatory communities
3D genome structures are plastic and vary from cell to cell even in an isogenic sample. Here, the authors present an approach to identify frequent 3D chromatin clusters across a population of genome structures, either deconvoluted from ensemble-averaged Hi-C data or from a collection of single-cell Hi-C data.
- Chao Dai
- , Wenyuan Li
- & Xianghong Jasmine Zhou
-
Article
| Open Access Cox process representation and inference for stochastic reaction–diffusion processes
Cox process representation and inference for stochastic reaction–diffusion processes
Stochastic reaction-diffusion systems are used for modelling spatial dynamics in many disciplines, but parameter inference and model selection remain challenging. Here the authors offer a solution enabled by a connection between reaction-diffusion and the well-studied spatio-temporal Cox processes.
- David Schnoerr
- , Ramon Grima
- & Guido Sanguinetti
-
Article
| Open Access Conservation of uORF repressiveness and sequence features in mouse, human and zebrafish
Conservation of uORF repressiveness and sequence features in mouse, human and zebrafish
Upstream open reading frames (uORFs) can repress gene expression. Here, Guo-Liang Chew and colleagues use bioinformatics approaches to show that conservation of uORF-mediated translational repression is mediated by sequence features in human, mouse and zebrafish genomes.
- Guo-Liang Chew
- , Andrea Pauli
- & Alexander F. Schier
-
Article
| Open Access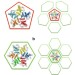 Coarse-grained simulation reveals key features of HIV-1 capsid self-assembly
Coarse-grained simulation reveals key features of HIV-1 capsid self-assembly
Significant morphological changes occur during the conversion of the immature HIV virion into a mature infectious form. Here the authors use coarse-grained molecular dynamics simulations to model HIV-1 capsid self-assembly and disassembly events that suggests several metastable capsid intermediates sensitive to local conditions.
- John M. A. Grime
- , James F. Dama
- & Gregory A. Voth
-
Article
| Open Access A combined computational and structural model of the full-length human prolactin receptor
A combined computational and structural model of the full-length human prolactin receptor
The prolactin receptor consists of a folded extracellular domain, a transmembrane domain and an intracellular intrinsically disordered domain. Here the authors use a combined experimental and computational approach to obtain a structure of a class I cytokine receptor, the human prolactin receptor.
- Katrine Bugge
- , Elena Papaleo
- & Birthe B. Kragelund
-
Article
| Open Access Arrhythmia risk stratification of patients after myocardial infarction using personalized heart models
Arrhythmia risk stratification of patients after myocardial infarction using personalized heart models
Sudden arrhythmic death is a leading cause of mortality, however approaches to identify at-risk patients are of low sensitivity and specificity. Here, the authors develop a personalized approach to assess arrhythmia risk in post-infarction patients based on cardiac imaging and computational modelling that significantly outperforms existing clinical metrics.
- Hermenegild J. Arevalo
- , Fijoy Vadakkumpadan
- & Natalia A. Trayanova
-
Article
| Open Access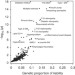 Identifying genetically driven clinical phenotypes using linear mixed models
Identifying genetically driven clinical phenotypes using linear mixed models
Use of general linear mixed models (GLMMs) in genetic variance analysis can quantify the relative contribution of additive effects from genetic variation on a given trait. Here, Jonathan Mosley and colleagues apply GLMM in a phenome-wide analysis and show that genetic variations in the HLA region are associated with 44 phenotypes, 5 phenotypes which were not previously reported in GWASes.
- Jonathan D. Mosley
- , John S. Witte
- & Joshua C. Denny
-
Article
| Open Access Fast and sensitive taxonomic classification for metagenomics with Kaiju
Fast and sensitive taxonomic classification for metagenomics with Kaiju
Here, Anders Krogh and colleagues describe Kaiju, a metagenome taxonomic classification program that uses maximum (in-)exact matches on the protein-level to account for evolutionary divergence. The authors show that Kaiju performs faster and is more sensitive compared with existing algorithms and can be used on a standard computer.
- Peter Menzel
- , Kim Lee Ng
- & Anders Krogh
-
Article
| Open Access Tor forms a dimer through an N-terminal helical solenoid with a complex topology
Tor forms a dimer through an N-terminal helical solenoid with a complex topology
The target of rapamycin (Tor) is a Ser/Thr protein kinase that regulates a wide range of anabolic and catabolic processes. Here the authors describe a sub-nanometer cryo-EM structure of a yeast Tor–Lst8 complex and propose an overall topology that differs from that previously suggested for mTORC1.
- Domagoj Baretić
- , Alex Berndt
- & Roger L. Williams
-
Article
| Open Access High-dimensional genomic data bias correction and data integration using MANCIE
High-dimensional genomic data bias correction and data integration using MANCIE
Analyses of data from high-throughput genomic technologies are challenging given large data dimensionality. Here, Liu and colleagues describe a method called MANCIE (Matrix Analysis and Normalization by Concordant Information Enhancement) that can conduct genomic data normalization and bias correction to detect biologically relevant information.
- Chongzhi Zang
- , Tao Wang
- & X. Shirley Liu
-
Article
| Open Access Genome-wide assessment of differential translations with ribosome profiling data
Genome-wide assessment of differential translations with ribosome profiling data
The global measurement of ribosome occupancy on mRNAs is commonly used as a proxy in estimating rates of protein synthesis. Here the authors describe Xtail, a computational approach that facilitates the extraction of accurate quantitative insight from ribosome profiling data (Ribo-Seq).
- Zhengtao Xiao
- , Qin Zou
- & Xuerui Yang
-
Article
| Open Access Rationally reduced libraries for combinatorial pathway optimization minimizing experimental effort
Rationally reduced libraries for combinatorial pathway optimization minimizing experimental effort
Rational design in metabolic engineering is often difficult and limited to small screens, favouring construction of compressed smart libraries. Here the authors introduce RedLibs, an algorithm to design combinatorial RBS libraries to allow pathway optimization with minimal experimental resources.
- Markus Jeschek
- , Daniel Gerngross
- & Sven Panke
-
Article
| Open Access Constructing 3D interaction maps from 1D epigenomes
Constructing 3D interaction maps from 1D epigenomes
The human genome is highly organized, with one-dimensional chromatin states packaged into higher level three-dimensional architecture. Here, the authors present EpiTensor that can identify 3D spatial associations from 1D epigenetic information.
- Yun Zhu
- , Zhao Chen
- & Wei Wang
-
Article
| Open Access Data publication with the structural biology data grid supports live analysis
Data publication with the structural biology data grid supports live analysis
The validation and analysis of X-ray crystallographic data is essential for reproducibility and the development of crystallographic methods. Here, the authors describe a repository for crystallographic datasets and demonstrate some of the ways it could serve the crystallographic community.
- Peter A. Meyer
- , Stephanie Socias
- & Piotr Sliz
-
Article
| Open Access A workflow to process 3D+time microscopy images of developing organisms and reconstruct their cell lineage
A workflow to process 3D+time microscopy images of developing organisms and reconstruct their cell lineage
Quantitative analysis of embryonic cell dynamics from large data sets remains a major challenge in the field of developmental biology. Here the authors develop software and a workflow to reconstruct cell lineage trees from 3D time lapse imaging data sets from several developing organisms including zebrafish, tunicates and sea urchins.
- Emmanuel Faure
- , Thierry Savy
- & Paul Bourgine
-
Article
| Open Access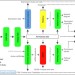 An interactive web-based application for Comprehensive Analysis of RNAi-screen Data
An interactive web-based application for Comprehensive Analysis of RNAi-screen Data
Analysis of RNAi screens is a multi-step process requiring the sequential use of several unrelated resources. Here the authors generate an online resource integrating RNAi analytic tools and filters into a seamless workflow, which improves the specificity, selectivity and reproducibility of the results.
- Bhaskar Dutta
- , Alaleh Azhir
- & Iain D. C. Fraser
-
Article
| Open Access Accurate prediction of cellular co-translational folding indicates proteins can switch from post- to co-translational folding
Accurate prediction of cellular co-translational folding indicates proteins can switch from post- to co-translational folding
The folding of protein domains can occur concomitant with their synthesis, and the rates at which individual codons are translated by the ribosome can affect the folding process. Here the authors present a kinetic model that accurately predicts the probability that a nascent protein domain will co-translationally fold in vivo.
- Daniel A. Nissley
- , Ajeet K. Sharma
- & Edward P. O’Brien
-
Article
| Open Access Affinity and competition for TBP are molecular determinants of gene expression noise
Affinity and competition for TBP are molecular determinants of gene expression noise
TATA boxes in gene promoters are associated with high level of cell-to-cell variation in gene expression. Through integration of multiple data sets, the authors now provide insights into how the interactions of TBP with DNA and other proteins can lead to noisy expression.
- Charles N. J. Ravarani
- , Guilhem Chalancon
- & M. Madan Babu
-
Article
| Open Access Using food-web theory to conserve ecosystems
Using food-web theory to conserve ecosystems
The influence of species conservation on food webs is less well understood than the effects of species loss. Here, the authors test several indices against optimal food web management and find no current metrics are reliably effective at identifying species conservation priorities.
- E. McDonald-Madden
- , R. Sabbadin
- & H. P. Possingham
-
Article
| Open Access Rapid antibiotic-resistance predictions from genome sequence data for Staphylococcus aureus and Mycobacterium tuberculosis
Rapid antibiotic-resistance predictions from genome sequence data for Staphylococcus aureus and Mycobacterium tuberculosis
The clinical application of new sequencing techniques is expected to accelerate pathogen identification. Here, Bradley et al. present a clinician-friendly software package that uses sequencing data for quick and accurate prediction of antibiotic resistance profiles for S. aureus and M. tuberculosis.
- Phelim Bradley
- , N. Claire Gordon
- & Zamin Iqbal
-
Article
| Open Access A new tool called DISSECT for analysing large genomic data sets using a Big Data approach
A new tool called DISSECT for analysing large genomic data sets using a Big Data approach
Availability of computing power can limit computational analysis of large genetic and genomic datasets. Here, Canela-Xandri, et al. describe a software called DISSECT that is capable of analyzing large-scale genetic data by distributing the work across thousands of networked computers.
- Oriol Canela-Xandri
- , Andy Law
- & Albert Tenesa
-
Article
| Open Access A comprehensive assessment of somatic mutation detection in cancer using whole-genome sequencing
A comprehensive assessment of somatic mutation detection in cancer using whole-genome sequencing
Cancer genetics has benefited from the advent of next generation sequencing, yet a comparison of sequencing and analysis techniques is lacking. Here, the authors sequence a normal-tumour pair and perform data analysis at multiple institutes and highlight some of the pitfalls associated with the different methods.
- Tyler S. Alioto
- , Ivo Buchhalter
- & Ivo G. Gut
-
Article
| Open Access Temporal and spatial dynamics of scaling-specific features of a gene regulatory network in Drosophila
Temporal and spatial dynamics of scaling-specific features of a gene regulatory network in Drosophila
How pattern formation is regulated relative to the size of an organism is unclear. Here, Wu et al.take data from gap gene expression in flies of different sizes together with simulations, identifying how scaling emerges dynamically and that local patterning influences global gene regulatory networks.
- Honggang Wu
- , Manu
- & Jun Ma
-
Article
| Open Access miRNA–target chimeras reveal miRNA 3′-end pairing as a major determinant of Argonaute target specificity
miRNA–target chimeras reveal miRNA 3′-end pairing as a major determinant of Argonaute target specificity
microRNAs (miRNAs) act as sequence-specific guides for Argonaute (AGO) proteins. By using a modified AGO HITS-CLIP strategy that enriches miRNAs ligated to their endogenous mRNA targets, here the authors show that miRNA 3' end pairing is a general determinant of AGO binding specificity.
- Michael J. Moore
- , Troels K. H. Scheel
- & Robert B. Darnell
-
Article
| Open Access Characterizing noise structure in single-cell RNA-seq distinguishes genuine from technical stochastic allelic expression
Characterizing noise structure in single-cell RNA-seq distinguishes genuine from technical stochastic allelic expression
Single-cell RNA-sequencing (scRNA-seq) can be applied to dissect the kinetics of gene expression and patterns of allele-specific expression. Here, Kim et al.report a generative statistical model that can separate biological variability from technical noise by quantifying technical noise using external RNA spike-ins.
- Jong Kyoung Kim
- , Aleksandra A. Kolodziejczyk
- & John C. Marioni
-
Article
| Open Access ChEC-seq kinetics discriminates transcription factor binding sites by DNA sequence and shape in vivo
ChEC-seq kinetics discriminates transcription factor binding sites by DNA sequence and shape in vivo
In chromatin endogenous cleavage (ChEC), micrococcal nuclease (MNase) is fused to a protein of interest and its cleavage is thus targeted to specific genomic loci in vivo. Here, the authors show that time-resolved ChEC-seq (high-throughput sequencing after ChEC) can detect DNA shape patterns regardless of motif strength.
- Gabriel E. Zentner
- , Sivakanthan Kasinathan
- & Steven Henikoff
-
Article
| Open Access Bayesian integration of genetics and epigenetics detects causal regulatory SNPs underlying expression variability
Bayesian integration of genetics and epigenetics detects causal regulatory SNPs underlying expression variability
Das et al. present a novel Bayesian approach called expression Quantitative Trait enhancer Loci (eQTeL), which effectively integrates genetic and epigenetic information to identify combination of regulatory genomic variants underlying expression variance. Using various functional data, the authors show the variants identified by eQTeL are likely to be causal.
- Avinash Das
- , Michael Morley
- & Sridhar Hannenhalli
-
Article
| Open Access Systematic analysis of somatic mutations impacting gene expression in 12 tumour types
Systematic analysis of somatic mutations impacting gene expression in 12 tumour types
Assessing functional impact of mutations in cancer on gene expression can improve our understanding of cancer biology and may identify potential therapeutic targets. Here, Ding et al. describe a novel statistical model named xseq for a systematic survey of how mutations impact transcriptome landscapes across 12 different tumour types.
- Jiarui Ding
- , Melissa K. McConechy
- & Sohrab P. Shah
-
Article
| Open Access Lower glycolysis carries a higher flux than any biochemically possible alternative
Lower glycolysis carries a higher flux than any biochemically possible alternative
The biochemical pathways of central carbon metabolism are highly conserved across all domains of life. Here, Courtet al. use a computational approach to test all possible pathways of glycolysis and gluconeogenesis and find that the existing trunk pathways may represent a maximal flux solution selected for during evolution.
- Steven J. Court
- , Bartlomiej Waclaw
- & Rosalind J. Allen
-
Article
| Open Access Global priorities for an effective information basis of biodiversity distributions
Global priorities for an effective information basis of biodiversity distributions
Comprehensive digital information on species distributions is crucial for research in ecology, evolution and conservation. Here, Meyer et al.find large gaps and biases in global vertebrate point records, especially in emerging economies, and identify key factors currently limiting information.
- Carsten Meyer
- , Holger Kreft
- & Walter Jetz
-
Article
| Open Access Systematic chromatin state comparison of epigenomes associated with diverse properties including sex and tissue type
Systematic chromatin state comparison of epigenomes associated with diverse properties including sex and tissue type
In contrast to genetic information, epigenetic state varies greatly under different conditions. Here, the authors develop ChromDiff and apply it to ENCODE and Roadmap Epigenomics datasets to find chromatin state differences associated with sex, tissue, and developmental age.
- Angela Yen
- & Manolis Kellis
Browse broader subjects
Browse narrower subjects
- Biochemical reaction networks
- Cellular signalling networks
- Classification and taxonomy
- Communication and replication
- Computational models
- Computational neuroscience
- Computational platforms and environments
- Data acquisition
- Data integration
- Data mining
- Data processing
- Data publication and archiving
- Databases
- Functional clustering
- Gene ontology
- Gene regulatory networks
- Genome informatics
- Hardware and infrastructure
- High-throughput screening
- Image processing
- Literature mining
- Machine learning
- Microarrays
- Network topology
- Phylogeny
- Power law
- Predictive medicine
- Probabilistic data networks
- Programming language
- Protein analysis
- Protein design
- Protein folding
- Protein function predictions
- Protein structure predictions
- Proteome informatics
- Quality control
- Scale invariance
- Sequence annotation
- Software
- Standards
- Statistical methods
- Virtual drug screening

 Sequence element enrichment analysis to determine the genetic basis of bacterial phenotypes
Sequence element enrichment analysis to determine the genetic basis of bacterial phenotypes