Featured
-
-
Article
| Open Access Visual analysis of mass cytometry data by hierarchical stochastic neighbour embedding reveals rare cell types
Visual analysis of mass cytometry data by hierarchical stochastic neighbour embedding reveals rare cell types
Single cell profiling yields high dimensional data of very large numbers of cells, posing challenges of visualization and analysis. Here the authors introduce a method for analysis of mass cytometry data that can handle very large datasets and allows their intuitive and hierarchical exploration.
- Vincent van Unen
- , Thomas Höllt
- & Boudewijn P. F. Lelieveldt
-
Article
| Open Access Genome-wide identification and differential analysis of translational initiation
Genome-wide identification and differential analysis of translational initiation
Translation initiation sequencing (TI-seq) has revealed unexpected diversity in protein isoforms. Here, Zhang et al. present Ribo-TISH, a computational toolkit that can detect and compare TIs across conditions and improve open reading frame prediction from different types of ribosome profiling data.
- Peng Zhang
- , Dandan He
- & Yiwen Chen
-
Article
| Open Access An extracellular matrix-related prognostic and predictive indicator for early-stage non-small cell lung cancer
An extracellular matrix-related prognostic and predictive indicator for early-stage non-small cell lung cancer
Prognosis and prediction of adjuvant chemotherapy response in non-small cell lung cancer can have significant clinical impact. Here, the authors show that differential expression of a 29 extracellular matrix gene indicator, EPPI, can predict patient outcome.
- Su Bin LIM
- , Swee Jin TAN
- & Chwee Teck LIM
-
Article
| Open Access Exocytosis-coordinated mechanisms for tip growth underlie pollen tube growth guidance
Exocytosis-coordinated mechanisms for tip growth underlie pollen tube growth guidance
Tip-growing cells can find their growing path toward the source of attractive signals. Here, using experimental data and mathematical modeling, Luo et al. demonstrate that tip-localized exocytosis can integrate guidance cues with Rho GTPase signaling to control cell wall mechanics and direct tip growth in Arabidopsis pollen tubes.
- Nan Luo
- , An Yan
- & Zhenbiao Yang
-
Article
| Open Access Deconvoluting interrelationships between concentrations and chemical shifts in urine provides a powerful analysis tool
Deconvoluting interrelationships between concentrations and chemical shifts in urine provides a powerful analysis tool
The NMR chemical shifts of a substance in urine strongly depend on the composition of the mixture itself, and this makes automatic assignment for quantification very difficult. Here the authors show the chemical shifts of signals and the concentration of NMR-invisible inorganic ions in urine, are predictable.
- Panteleimon G. Takis
- , Hartmut Schäfer
- & Claudio Luchinat
-
Article
| Open Access Decoding critical long non-coding RNA in ovarian cancer epithelial-to-mesenchymal transition
Decoding critical long non-coding RNA in ovarian cancer epithelial-to-mesenchymal transition
The role of lncRNA is unclear with respect to epithelial-to-mesenchymal transition (EMT), which is linked to ovarian cancer metastasis. Here, the authors show lncRNA DNM3OS expression contributes to ovarian cancer EMT, cell migration/invasion, and correlates with worse overall patient survival.
- Ramkrishna Mitra
- , Xi Chen
- & Christine M. Eischen
-
Article
| Open Access Shaping bacterial population behavior through computer-interfaced control of individual cells
Shaping bacterial population behavior through computer-interfaced control of individual cells
Individual bacteria interact with each other and their environment to produce population-level patterns of gene expression. Here the authors use an automated platform combined with optogenetic feedback to manipulate population behaviors through dynamic control of individual cells.
- Remy Chait
- , Jakob Ruess
- & Călin C. Guet
-
Article
| Open Access Comparative genomic analysis of esophageal squamous cell carcinoma between Asian and Caucasian patient populations
Comparative genomic analysis of esophageal squamous cell carcinoma between Asian and Caucasian patient populations
Esophageal squamous cell carcinoma (ESCC) exhibits differences in incidence and survival patterns among races. Here, analysis of Chinese and TCGA ESCC patients reveals that Asian patients exhibit higher TP53, EP300 and NFE2L2 mutational frequencies, and mutated CSMD3 associates with better prognosis.
- Jiaying Deng
- , Hu Chen
- & Kuaile Zhao
-
Article
| Open Access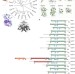 A comprehensive structural, biochemical and biological profiling of the human NUDIX hydrolase family
A comprehensive structural, biochemical and biological profiling of the human NUDIX hydrolase family
The NUDIX hydrolases are known to be involved in several cellular processes and diseases, such as cancer, but remain poorly characterized as a family. Here, the authors provide a comprehensive analysis of the structural, biochemical, and expression properties of 18 human NUDIX proteins, and begin to address their functional inter-relationships.
- Jordi Carreras-Puigvert
- , Marinka Zitnik
- & Thomas Helleday
-
Article
| Open Access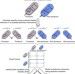 Genome-driven evolutionary game theory helps understand the rise of metabolic interdependencies in microbial communities
Genome-driven evolutionary game theory helps understand the rise of metabolic interdependencies in microbial communities
The rise of metabolic interdependencies among microbes is still poorly understood. Here, taking the underlying biochemical networks into consideration, Zomorrodi and Segrè integrate genome-scale metabolic models with evolutionary game theory to study the rise of cross-feeding in microbial communities.
- Ali R. Zomorrodi
- & Daniel Segrè
-
Article
| Open Access Significance estimation for large scale metabolomics annotations by spectral matching
Significance estimation for large scale metabolomics annotations by spectral matching
Matching fragment spectra to reference library spectra is an important procedure for annotating small molecules in untargeted mass spectrometry based metabolomics studies. Here, the authors develop strategies to estimate false discovery rates (FDR) by empirical Bayes and target-decoy based methods which enable a user to define the scoring criteria for spectral matching.
- Kerstin Scheubert
- , Franziska Hufsky
- & Sebastian Böcker
-
Article
| Open Access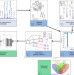 Network inference from glycoproteomics data reveals new reactions in the IgG glycosylation pathway
Network inference from glycoproteomics data reveals new reactions in the IgG glycosylation pathway
IgG glycosylation is an important factor in immune function, yet the molecular details of protein glycosylation remain poorly understood. The data-driven approach presented here uses large-scale plasma IgG mass spectrometry measurements to infer new biochemical reactions in the glycosylation pathway.
- Elisa Benedetti
- , Maja Pučić-Baković
- & Jan Krumsiek
-
Article
| Open Access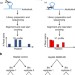 RNA editing by ADAR1 leads to context-dependent transcriptome-wide changes in RNA secondary structure
RNA editing by ADAR1 leads to context-dependent transcriptome-wide changes in RNA secondary structure
Adenosine deaminase acting on RNA 1 (ADAR1) edits adenosine to inosine. Here the authors, using parallel analysis of RNA secondary structure sequencing, provide evidence that ADAR1 induces sequence-context-dependent RNA secondary structures changes, often leading to stabilization of the RNA duplex.
- Oz Solomon
- , Ayelet Di Segni
- & Gidi Rechavi
-
Article
| Open Access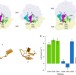 Origin of the omnipotence of eukaryotic release factor 1
Origin of the omnipotence of eukaryotic release factor 1
The eukaryotic release factor eRF1 is able to recognize the three stop codons UAA, UAG and UGA with high accuracy, while discriminating against near-cognate codons. Here the authors use molecular dynamic simulation to provide insight into the molecular basis behind the remarkable codon specificity of eRF1.
- Christoffer Lind
- , Ana Oliveira
- & Johan Åqvist
-
Article
| Open Access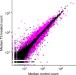 The North American bullfrog draft genome provides insight into hormonal regulation of long noncoding RNA
The North American bullfrog draft genome provides insight into hormonal regulation of long noncoding RNA
The globally-distributed Ranidae (true frogs) are the largest frog family. Here, Hammond et al. present a draft genome of the North American bullfrog, Rana (Lithobates) catesbeiana, as a foundation for future understanding of true frog genetics as amphibian species face difficult environmental challenges.
- S. Austin Hammond
- , René L. Warren
- & Inanc Birol
-
Article
| Open Access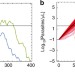 Assessing the impact of imperfect adherence to artemether-lumefantrine on malaria treatment outcomes using within-host modelling
Assessing the impact of imperfect adherence to artemether-lumefantrine on malaria treatment outcomes using within-host modelling
Artemether lumefantrine is widely used for the treatment of uncomplicated malaria. The impact of imperfect patient adherence to the six-dose regimen is hard to assess. Using adherence data for unsupervised patients, the authors model how suboptimal adherence affects treatment outcomes.
- Joseph D. Challenger
- , Katia Bruxvoort
- & Lucy C. Okell
-
Article
| Open Access Paradoxes in leaky microbial trade
Paradoxes in leaky microbial trade
Microbes live in communities and exchange metabolites, but the resulting dynamics are poorly understood. Here, the authors study the interplay between metabolite production strategies and population dynamics, and find that complex and unexpected dynamics emerge even in simple microbial economies.
- Yoav Kallus
- , John H. Miller
- & Eric Libby
-
Article
| Open Access The challenge of mapping the human connectome based on diffusion tractography
The challenge of mapping the human connectome based on diffusion tractography
Though tractography is widely used, it has not been systematically validated. Here, authors report results from 20 groups showing that many tractography algorithms produce both valid and invalid bundles.
- Klaus H. Maier-Hein
- , Peter F. Neher
- & Maxime Descoteaux
-
Article
| Open Access Mutational signatures reveal the dynamic interplay of risk factors and cellular processes during liver tumorigenesis
Mutational signatures reveal the dynamic interplay of risk factors and cellular processes during liver tumorigenesis
Tumorigenesis is a complex process driven by numerous risk factors. Here, genomic analysis of liver cancer reveals the evolution of mutational signatures during tumor development, highlighting mutational and structural signatures linked to environmental exposures and endogenous cellular processes.
- Eric Letouzé
- , Jayendra Shinde
- & Jessica Zucman-Rossi
-
Article
| Open Access Dense and accurate whole-chromosome haplotyping of individual genomes
Dense and accurate whole-chromosome haplotyping of individual genomes
Haplotype information is important in investigating many biological phenomena. Here, Porubsky et al. combine Strand-seq with long-read or linked-read sequencing to obtain complete and genome-wide haplotypes of a single individual genome at manageable costs.
- David Porubsky
- , Shilpa Garg
- & Tobias Marschall
-
Article
| Open Access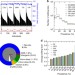 Structure-mediated modulation of mRNA abundance by A-to-I editing
Structure-mediated modulation of mRNA abundance by A-to-I editing
RNA A-to-I editing introduces single nucleotide changes to RNA, but its role in cells remains unclear. Here, the authors analyse A-to-I editomes in human samples and find that A-to-I editing stabilizes RNA secondary structures and reduces the accessibility of AGO2-miRNA to target sites in mRNAs.
- Anneke Brümmer
- , Yun Yang
- & Xinshu Xiao
-
Article
| Open Access Percolation transition of cooperative mutational effects in colorectal tumorigenesis
Percolation transition of cooperative mutational effects in colorectal tumorigenesis
Cancer is caused by accumulating genetic mutations. Here, the authors investigate the cooperative effect of these mutations in colorectal cancer patients and identify a giant cluster of mutation-propagating modules that undergoes percolation transition during tumorigenesis.
- Dongkwan Shin
- , Jonghoon Lee
- & Kwang-Hyun Cho
-
Article
| Open Access Identifying host regulators and inhibitors of liver stage malaria infection using kinase activity profiles
Identifying host regulators and inhibitors of liver stage malaria infection using kinase activity profiles
Host kinases facilitate Plasmodium liver stage (LS) infection, but systematic accounting of important players is lacking. Here, the authors use a computational approach and kinase activity profiles to identify host kinase regulators of LS infection and drugs that could eliminate parasite burden.
- Nadia Arang
- , Heather S. Kain
- & Alexis Kaushansky
-
Article
| Open Access Pan-cancer analysis of homozygous deletions in primary tumours uncovers rare tumour suppressors
Pan-cancer analysis of homozygous deletions in primary tumours uncovers rare tumour suppressors
Homozygous deletions are rare in cancers and often target tumour suppressor genes. Here, the authors conduct pan-cancer analyses and apply statistical modelling to identify 27 candidate tumour suppressors, including MAFTRR, KIAA1551, and IGF2BP2.
- Jiqiu Cheng
- , Jonas Demeulemeester
- & Peter Van Loo
-
Article
| Open Access Algorithm for post-clustering curation of DNA amplicon data yields reliable biodiversity estimates
Algorithm for post-clustering curation of DNA amplicon data yields reliable biodiversity estimates
A central problem in biodiversity estimation from genetic markers is the ability of algorithms to retain ‘true’ species while discarding artefacts. Here, the authors present a new post-clusturing curation algorithm using OTU co-occurrences to estimate plant biodiversity from soil samples.
- Tobias Guldberg Frøslev
- , Rasmus Kjøller
- & Anders Johannes Hansen
-
Article
| Open Access PLATO software provides analytic framework for investigating complexity beyond genome-wide association studies
PLATO software provides analytic framework for investigating complexity beyond genome-wide association studies
Centralized infrastructure to support analyses involving complexity beyond genome-wide association studies is broadly needed. Here, Ritchie and colleagues develop PLATO, a software tool to process and integrate various methods for this task.
- Molly A. Hall
- , John Wallace
- & Marylyn D. Ritchie
-
Article
| Open Access Functional reconstruction of a eukaryotic-like E1/E2/(RING) E3 ubiquitylation cascade from an uncultured archaeon
Functional reconstruction of a eukaryotic-like E1/E2/(RING) E3 ubiquitylation cascade from an uncultured archaeon
In eukaryotic cells, the ubiquitylation system regulates several cellular processes central to protein homoeostasis. Here the authors demonstrate the existence of an eukaryotic-like ubiquitylation cascade requiring E1, E2 and E3-like enzymes in the archaeon C. subterraneum, shedding light on the evolution of the ubiquitin-proteasome system.
- Rory Hennell James
- , Eva F. Caceres
- & Nicholas P. Robinson
-
Article
| Open Access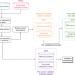 Gene isoforms as expression-based biomarkers predictive of drug response in vitro
Gene isoforms as expression-based biomarkers predictive of drug response in vitro
Altered mRNA splicing features in many cancers, but it has not been linked to drug response. Here, with their meta-analytic framework, the authors analyse pharmacogenomic data to identify isoform-based biomarkers predictive of in vitro drug response, and show them to frequently be strong predictors.
- Zhaleh Safikhani
- , Petr Smirnov
- & Benjamin Haibe-Kains
-
Article
| Open Access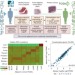 Comprehensive analysis of normal adjacent to tumor transcriptomes
Comprehensive analysis of normal adjacent to tumor transcriptomes
Normal tissue adjacent to the tumour (NAT) is often used as a control in cancer studies. Here, the authors analyse across cancer types the transcriptomes of healthy, NAT, and tumour tissues, and find that NAT presents a unique state, potentially due to inflammatory response of the NAT to the tumour tissue.
- Dvir Aran
- , Roman Camarda
- & Atul J. Butte
-
Article
| Open Access An integrative method to decode regulatory logics in gene transcription
An integrative method to decode regulatory logics in gene transcription
Existing transcriptional regulatory networks models fall short of deciphering the cooperation between multiple transcription factors on dynamic gene expression. Here the authors develop an integrative method that combines gene expression and transcription factor-DNA binding data to decode transcription regulatory logics.
- Bin Yan
- , Daogang Guan
- & Hailong Zhu
-
Article
| Open Access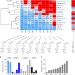 The β20–β21 of gp120 is a regulatory switch for HIV-1 Env conformational transitions
The β20–β21 of gp120 is a regulatory switch for HIV-1 Env conformational transitions
Binding of viral envelope glycoproteins (Env) to the host cell CD4 receptor mediates HIV-1 entry. Here, the authors develop compounds that inhibit the CD4-induced conformational changes in Env and show that the gp120 β20-β21 element is a key regulator for Env transitions.
- Alon Herschhorn
- , Christopher Gu
- & Joseph G. Sodroski
-
Article
| Open Access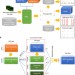 Genome-wide prediction of DNase I hypersensitivity using gene expression
Genome-wide prediction of DNase I hypersensitivity using gene expression
A map of the activities of all genomic regulatory elements across cell types and conditions would be a tremendous resource. The computational method introduced here predicts genome-wide accessible sites from gene expression data and allows the authors to build a database of regulatory element activities using publicly available transcriptome data.
- Weiqiang Zhou
- , Ben Sherwood
- & Hongkai Ji
-
Article
| Open Access Wind loads and competition for light sculpt trees into self-similar structures
Wind loads and competition for light sculpt trees into self-similar structures
Tree branches follow allometric scalings between length, thickness and dry mass. Here, Eloy and colleagues develop a functional-structural model that shows how such allometries in tree architecture can emerge through evolution as a result of competition for light, wind biomechanics, and wind sensing.
- Christophe Eloy
- , Meriem Fournier
- & Bruno Moulia
-
Article
| Open Access Inference of RNA decay rate from transcriptional profiling highlights the regulatory programs of Alzheimer’s disease
Inference of RNA decay rate from transcriptional profiling highlights the regulatory programs of Alzheimer’s disease
“mRNA abundance is determined by the rates of transcription and decay. Here, the authors propose a method for estimating the rate of differential mRNA decay from RNA-seq data and model mRNA stability in the brain, suggesting a link between mRNA stability and Alzheimer’s disease.”
- Rached Alkallas
- , Lisa Fish
- & Hamed S. Najafabadi
-
Article
| Open Access Counteracting structural errors in ensemble forecast of influenza outbreaks
Counteracting structural errors in ensemble forecast of influenza outbreaks
Inaccuracy of influenza forecasts based on dynamical models is partly due to nonlinear error growth. Here the authors address the error structure of a compartmental influenza model, and develop a new improved forecast approach combining dynamical error correction and statistical filtering techniques.
- Sen Pei
- & Jeffrey Shaman
-
Article
| Open Access Pan-cancer analysis of bi-allelic alterations in homologous recombination DNA repair genes
Pan-cancer analysis of bi-allelic alterations in homologous recombination DNA repair genes
Germline mutations in homologous recombination (HR) DNA repair genes are linked to breast and ovarian cancer. Here, the authors show that mutually exclusive bi-allelic inactivation of HR genes are present in other cancer types and associated with genomic features of HR deficiency, expanding the potential use of HR-directed therapies.
- Nadeem Riaz
- , Pedro Blecua
- & Jorge S. Reis-Filho
-
Article
| Open Access Integrative analysis of genomic and epigenomic regulation of the transcriptome in liver cancer
Integrative analysis of genomic and epigenomic regulation of the transcriptome in liver cancer
Hepatocellular carcinoma is known to harbour numerous genomic and epigenomic aberrations, driving transcriptomic deregulation. Here, the authors integrate genomic, epigenomic, and expression data to reveal three prognostic subtypes, providing insight to the pathobiology of hepatocellular carcinoma.
- Hyun Goo Woo
- , Ji-Hye Choi
- & Yoon Jun Kim
-
Article
| Open Access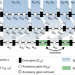 The chromosomal organization of horizontal gene transfer in bacteria
The chromosomal organization of horizontal gene transfer in bacteria
Horizontal gene transfer (HGT) is an important mechanism for genome evolution and adaptation in bacteria. Here, Oliveira and colleagues find HGT hotspots comprising ~ 1% of the chromosomal regions in 80 bacterial species.
- Pedro H. Oliveira
- , Marie Touchon
- & Eduardo P. C. Rocha
-
Article
| Open Access CNV-association meta-analysis in 191,161 European adults reveals new loci associated with anthropometric traits
CNV-association meta-analysis in 191,161 European adults reveals new loci associated with anthropometric traits
Individual SNPs have small effects on anthropometric traits, yet the impact of CNVs has remained largely unknown. Here, Kutalik and co-workers perform a large-scale genome-wide meta-analysis of structural variation and find rare CNVs associated with height, weight and BMI with large effect sizes.
- Aurélien Macé
- , Marcus A. Tuke
- & Zoltán Kutalik
-
Article
| Open Access Transcriptional regulation of endothelial cell behavior during sprouting angiogenesis
Transcriptional regulation of endothelial cell behavior during sprouting angiogenesis
Angiogenesis is a complex process that requires coordinated changes in endothelial cell behavior. Here the authors use Ribo-tag and RNA-Seq to determine temporal profiles of transcriptional activity during postnatal retinal angiogenesis, identifying transcriptional regulators of the process.
- Hyun-Woo Jeong
- , Benjamín Hernández-Rodríguez
- & Ralf H. Adams
-
Article
| Open Access Predicting metabolic adaptation from networks of mutational paths
Predicting metabolic adaptation from networks of mutational paths
The structure and dynamics of microbial communities reflect trade-offs in the ability to use different resources. Here, Josephides and Swain incorporate metabolic trade-offs into an eco-evolutionary model to predict networks of mutational paths and the evolutionary outcomes for microbial communities.
- Christos Josephides
- & Peter S. Swain
-
Article
| Open Access Aromatic side-chain conformational switch on the surface of the RNA Recognition Motif enables RNA discrimination
Aromatic side-chain conformational switch on the surface of the RNA Recognition Motif enables RNA discrimination
The RNA Recognition Motif (RRM) is the most ubiquitous RNA binding domain. Here the authors combined NMR and molecular dynamics simulations and show that the RRM RNA binding surface exists in different states and that a conformational switch of aromatic side-chains fine-tunes sequence specific binding affinities.
- Nana Diarra dit Konté
- , Miroslav Krepl
- & Frédéric H.-T. Allain
-
Article
| Open Access Stochastic priming and spatial cues orchestrate heterogeneous clonal contribution to mouse pancreas organogenesis
Stochastic priming and spatial cues orchestrate heterogeneous clonal contribution to mouse pancreas organogenesis
The pancreas arises from a small population of cells but how individual cells contribute to organ formation is unclear. Here, the authors deconstruct pancreas organogenesis into clonal units, showing that single progenitors give rise to heterogeneous multi-lineage and endocrinogenic single-lineage clones.
- Hjalte List Larsen
- , Laura Martín-Coll
- & Anne Grapin-Botton
-
Article
| Open Access A network integration approach for drug-target interaction prediction and computational drug repositioning from heterogeneous information
A network integration approach for drug-target interaction prediction and computational drug repositioning from heterogeneous information
Network-based data integration for drug–target prediction is a promising avenue for drug repositioning, but performance is wanting. Here, the authors introduce DTINet, whose performance is enhanced in the face of noisy, incomplete and high-dimensional biological data by learning low-dimensional vector representations.
- Yunan Luo
- , Xinbin Zhao
- & Jianyang Zeng
-
Article
| Open Access The interdependent network of gene regulation and metabolism is robust where it needs to be
The interdependent network of gene regulation and metabolism is robust where it needs to be
Although networks of interacting genes and metabolic reactions are interdependent, they have largely been treated as separate systems. Here the authors apply a statistical framework for interdependent networks to E. coli, and show that it is sensitive to gene and protein perturbations but robust against metabolic changes.
- David F. Klosik
- , Anne Grimbs
- & Marc-Thorsten Hütt
-
Article
| Open Access Accurate immune repertoire sequencing reveals malaria infection driven antibody lineage diversification in young children
Accurate immune repertoire sequencing reveals malaria infection driven antibody lineage diversification in young children
Somatic hypermutation of antibodies can occur in infants but are difficult to track. Here the authors present a new method called MIDCIRS for deep quantitative repertoire sequencing with few cells, and show infants as young as 3 months can expand antibody lineage complexity in response to malaria infection.
- Ben S. Wendel
- , Chenfeng He
- & Ning Jiang
-
Article
| Open Access Identifying topologically associating domains and subdomains by Gaussian Mixture model And Proportion test
Identifying topologically associating domains and subdomains by Gaussian Mixture model And Proportion test
Spatial organization of the genome plays a crucial role in regulating gene expression. Here the authors introduce GMAP, the Gaussian Mixture model And Proportion test, to identify topologically associating domains and subdomains in Hi-C data.
- Wenbao Yu
- , Bing He
- & Kai Tan
-
Article
| Open Access Discovery of a proteolytic flagellin family in diverse bacterial phyla that assembles enzymatically active flagella
Discovery of a proteolytic flagellin family in diverse bacterial phyla that assembles enzymatically active flagella
So far no enzymatic activity has been attributed to flagellin, the major component of bacterial flagella. Here the authors use bioinformatic analysis and identify a metallopeptidase insertion in flagellins from 74 bacterial species and show that recombinant flagellin and flagellar filaments have proteolytic activity.
- Ulrich Eckhard
- , Hina Bandukwala
- & Andrew C. Doxey
-
Article
| Open Access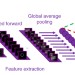 Reconstructing cell cycle and disease progression using deep learning
Reconstructing cell cycle and disease progression using deep learning
The interpretation of information-rich, high-throughput single-cell data is a challenge requiring sophisticated computational tools. Here the authors demonstrate a deep convolutional neural network that can classify cell cycle status on-the-fly.
- Philipp Eulenberg
- , Niklas Köhler
- & F. Alexander Wolf
Browse broader subjects
Browse narrower subjects
- Biochemical reaction networks
- Cellular signalling networks
- Classification and taxonomy
- Communication and replication
- Computational models
- Computational neuroscience
- Computational platforms and environments
- Data acquisition
- Data integration
- Data mining
- Data processing
- Data publication and archiving
- Databases
- Functional clustering
- Gene ontology
- Gene regulatory networks
- Genome informatics
- Hardware and infrastructure
- High-throughput screening
- Image processing
- Literature mining
- Machine learning
- Microarrays
- Network topology
- Phylogeny
- Power law
- Predictive medicine
- Probabilistic data networks
- Programming language
- Protein analysis
- Protein design
- Protein folding
- Protein function predictions
- Protein structure predictions
- Proteome informatics
- Quality control
- Scale invariance
- Sequence annotation
- Software
- Standards
- Statistical methods
- Virtual drug screening

 Single-cell absolute contact probability detection reveals chromosomes are organized by multiple low-frequency yet specific interactions
Single-cell absolute contact probability detection reveals chromosomes are organized by multiple low-frequency yet specific interactions